Abstract
Low-abundance proteins (LAPs) play a very important role in interaction, regulation, and metabolism of plant biological processes. A combinatorial peptide ligand library (CPLL) can solve the problem of high-abundance proteins (HAPs) masking LAPs and enlarging the dynamic range of protein concentrations perfectly and be considered as one of the most advanced approaches for plant proteomics research. In this paper, a proper CPLL method to rice leaf proteins was established for the first time and 1056 proteins were identified in rice leaf extracts, and 624 (59.1%) LAPs were newly detected after CPLL. Based on this technology, we detected the response of rice to Cd stress and analyzed the differential LAPs and the biological significance of misexpressed proteins before and after Cd stress by bioinformatics analysis. An important contribution has also been made to a better understanding of the complex mechanisms by which rice adapts to Cd stress.

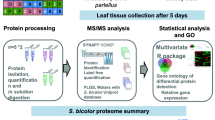
Graphical abstract







Similar content being viewed by others
References
Zhao YL, Huang X, Liu LW, Wang PY, Long QS, Tao QQ, et al. Identification of racemic and chiral carbazole derivatives containing an isopropanolamine linker as prospective surrogates against plant pathogenic bacteria: in vitro and in vivo assays and quantitative proteomics. J Agric Food Chem. 2019;67(26):7512–25.
Phizicky E, Bastiaens PIH, Zhu H, Snyder M, Fields S. Protein analysis on a proteomic scale. Nature. 2003;422:208–15.
Yan SP, Zhang QY, Tang ZC, Su WA, Sun WN. Comparative proteomic analysis provides new insights into chilling stress responses in rice. Mol Cell Proteomics. 2006;5:484–96.
Cremer JE, Bean SR, Tilley MM, Ioerger BP, Ohm JB, Kaufman RC, et al. Grain sorghum proteomics: integrated approach toward characterization of endosperm storage proteins in kafirin allelic variants. J Agric Food Chem. 2014;62(40):9819–31.
Herman EM, Helm RM, Jung R, Kinney AJ. Genetic modification removes an immunodominant allergen from soybean. Plant Physiol. 2003;132:36–43.
Pier GR, Annalisa C, Paolo A. Prefractionation techniques in proteome analysis: the mining tools of the third millennium. Electrophoresis. 2005;26:297–319.
Espagne C, Martinez A, Valot B, Meinnel T, Giglione C. Alternative and effective proteomic analysis in Arabidopsis. Proteomics. 2007;7:3788–99.
Cho JH, Hwang H, Cho MH, Kwon YK, Jeon JS, Bhoo SH, et al. The effect of DTT in protein preparations for proteomic analysis: removal of a highly abundant plant enzyme, ribulose bisphosphate carboxylase/oxygenase. Plant Biol. 2008;51:297–301.
Boschetti E, Righetti PG. The ProteoMiner in the proteomic arena: a non-depleting tool for discovering low-abundance species. J Proteome. 2008;71:255–64.
Boschetti E, Righetti PG. The art of observing rare protein species in proteomes with peptide ligand libraries. Proteomics. 2009;9:1492–510.
Righetti PG, Boschetti E. Low-abundance plant protein enrichment with peptide libraries to enlarge proteome coverage and related applications. Plant Sci. 2020;290:110302. https://doi.org/10.1016/j.plantsci.2019.110302.
Fasoli E, D'Amato A, Kravchuk AV, Boschetti E, Bachi A, Righetti PG. Popeye strikes again: the deep proteome of spinach leaves. J Proteome. 2011;74:127–36.
Castagna A, Cecconi D, Sennels L, Rappsilber J, Guerrier L, Fortis F, et al. Exploring the hidden human urinary proteome via ligand library beads. J Proteome Res. 2005;4:1917–30.
Thulasiraman V, Lin S, Gheorghiu L, Lathrop J, Lomas L, Hammond D, et al. Reduction of the concentration difference of proteins in biological liquids using a library of combinatorial ligands. Electrophoresis. 2005;26:3561–71.
Guerrier L, Claverol S, Finzi L, Paye F, Fortis F, Boschetti E, et al. Contribution of solid-phase hexapeptide ligand libraries to the repertoire of human bile proteins. J Chromatogr A. 2007;1176:192–205.
D’Ambrosio C, Arena S, Scaloni A, Guerrier L, Boschetti E, Mendieta ME, et al. Exploring the chicken egg white proteome with combinatorial peptide ligand libraries. J Proteome Res. 2008;7:3461–74.
Roux-Dalvai F, Gonzalez de Peredo A, Simó C, Guerrier L, Bouyssié D, Zanella A, et al. Extensive analysis of the cytoplasmic proteome of human erythrocytes using the peptide ligand library technology and advanced mass spectrometry. Mol Cell Proteomics. 2008;7:2254–69.
D’Amato A, Bachi A, Fasoli E, Boschetti E, Peltre G, Sénéchal H, et al. In-depth exploration of cow’s whey proteome via combina-torial peptide ligand libraries. J Proteome Res. 2009;8:3925–36.
Farinazzo A, Restuccia U, Bachi A, Guerrier L, Fortis F, Boschetti E, et al. Chicken egg yolk cytoplasmic proteome, mined via combinatorial peptide ligand libraries. J Chromatogr A. 2009;1216:1241–52.
Shahali Y, Sutra JP, Fasoli E, D'Amato A, Righetti PG, Futamura N, et al. Allergomic study of cypress pollen via combinatorial peptide ligand libraries. J Proteome. 2012;77:101–10.
D’Amato A, Esteve C, Fasoli E, Citterio A, Righetti PG. Proteomic analysis of Lycium barbarum (Goji) fruit via combinatorial peptide ligand libraries. Electrophoresis. 2013;34:1729–36.
Saez V, Fasoli E, D’Amato A, Simó-Alfonso E, Righetti PG. Artichoke and Cynar liqueur: two (not quite) entangled proteomes. Biochim Biophys Acta. 1834;2013:119–26.
Lee PY, Osman J, Low TY, Jamal R. Plasma/serum proteomics: depletion strategies for reducing high-abundance proteins for biomarker discovery. Bioanalysis. 2019;6:4290–303.
Girolamo DF, Boschetti E, Chung MC, Jamal R. “Proteomineering” or not? The debate on biomarker discovery in sera continues. J Proteome. 2011;74:589–94.
Kim ST, Cho KS, Jang YS, Kang KY. Two-dimensional electrophoretic analysis of rice proteins by polyethylene glycol fractionation for protein arrays. Electrophoresis. 2001;22:2103–9.
Mouton-Barbosa E, Roux-Dalvai F, Bouyssié D, Berger F, Schmidt E, Righetti PG, et al. In-depth exploration of cerebrospinal fluid by combining peptide ligand library treatment and label-free protein quantification. Mol Cell Proteomics. 2010;9:1006–21.
Righetti PG, Boschetti E. Global proteome analysis in plants by means of peptide libraries and applications. J Proteome. 2016;143:3–14.
Kilambi HV, Manda K, Sanivarapu H, MauryaVK SR, Sreelakshmi Y. Shotgun proteomics of tomato fruits: evaluation, optimization and validation of sample preparation methods and mass spectrometric parameters. Front Plant Sci. 2016;7:1–14.
Fröhlich A, Gaupels F, Sarioglu H, Holzmeister C, Spannagl M, Durner J, et al. Looking deep inside: detection of low-abundance proteins in leaf extracts of arabidopsis and phloem exudates of pumpkin. Plant Physiol. 2012;59:902–14.
Guerrier L, Righetti PG, Boschetti E. Reduction of dynamic protein concentration range of biological extracts for the discovery of low-abundance proteins by means of hexapeptide ligand library. Nat Protoc. 2008;3:883–90.
Pennington ME, Lam KS, Cress AE. The use of a combinatorial library method to isolate human tumor cell adhesion peptides. Mol Divers. 1996;2:19–28.
Hu ZT, Tang X, Xiao Z, Fu M, Hu X. Electrophoresis separation and MS analysis of proteins in fermentation broth of Poria cocos. Chin Tradit Herb Drug. 2016;47(13):2269–76.
Chai SS, Ma YN, Gao HH, Qin ML, Yang H, Zhang HT, et al. Two-dimensional liquid chromatography separation and high resolution mass spectrometry analysis for proteome of rice leaves based on different extraction methods. Chin J Chromatogr. 2018;36(2):107–13.
Simo C, Bachi A, Cattaneo A, Guerrier L, Fortis F, Boschetti E, et al. Performance of combinatorial peptide libraries in capturing the low-abundance proteome of red blood cells. 1. behavior of mono- to hexapeptides. Anal Chem. 2008;80:3547–56.
Boschetti E, Bindschedler LV, Tang C, Elisa F, Giorgio RP. Combinatorial peptide ligand libraries and plant proteomics: a winning strategy at a price. J Chromatogr A. 2009;1216:1215–22.
Tu CJ, Li J, Young R, Page BJ, Engler F, Halfon MS, et al. A combinatorial peptide ligand libraries treatment followed by a dual-enzyme, dual-activation approach on a nano-flow LC/Orbitrap/ETD for comprehensive analysis of swine plasma proteome. Anal Chem. 2011;83(12):4802–13.
Lei AP, Chen H, Li SF, Hu ZL. Function, application, cloning and expression of glutathione s-transferase. Environ Sci Technol. 2009;32:85–91.
Yang YN, Jiang YS. Effect of calmodulin and its antagonist on heavy metal poisoning. Health Res. 1995;S1:22–5.
Luque-Garcia JL, Cabezas-Sanchez P, Camara C. Proteomics as a tool for examining the toxicity of heavy metals. Trends Anal Chem. 2011;30:703–16.
Sergeant K, Renaut J, Hausman JF. Proteomics as a toolbox to study the metabolic adjustment of trees during exposure to metal trace elements. Metal Toxicity in Plants: Perception, Signaling and Remediation. In: Gupta DK, editor. Springer-Verlag; 2012. p. 143–164.
Silke L, Martin B, Lena B. Shared sulfur mobilization routes for tRNA thiolation and molybdenum cofactor biosynthesis in prokaryotes and eukaryotes. Biomole. 2017;7(1):5–24.
Noma A, Sakaguchi Y, Suzuki T. Mechanistic characterization of the sulfur-relay system for eukaryotic 2-thiouridine biogenesis at tRNA wobble positions. Nucleic Acids Res. 2009;37(4):1337–52.
Liao XY, Chen TB, Xie H, Liu YR. Soil as contamination and its risk assessment in areas near the industrial districts of Chenzhou City, Southern China. Environ Int. 2005;31:791–8.
Ahsan N, Renaut J, Komatsu S. Recent developments in the application of proteomics to the analysis of plant responses to heavy metals. Proteomics. 2009;9:2602–21.
Cvjetko P, Zovko M, Balen B. Proteomics of heavy metal toxicity in plants. Arh Hig Rada Toksikol. 2014;65:1–18.
Funding
This research was supported by the National Natural Science Foundation of China (No. 31701408).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lin, X., Chai, S., Huang, S. et al. Insight of low-abundance proteins in rice leaves under Cd stress using combinatorial peptide ligand library technology. Anal Bioanal Chem 412, 5435–5446 (2020). https://doi.org/10.1007/s00216-020-02760-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-020-02760-z