Abstract
Haloarchaeal alcohol dehydrogenases are exciting biocatalysts with potential industrial applications. In this study, two alcohol dehydrogenase enzymes from the extremely halophilic archaeon Haloferax volcanii (HvADH1 and HvADH2) were homologously expressed and subsequently purified by immobilized metal-affinity chromatography. The proteins appeared to copurify with endogenous alcohol dehydrogenases, and a double Δadh2 Δadh1 gene deletion strain was constructed to prevent this occurrence. Purified HvADH1 and HvADH2 were compared in terms of stability and enzymatic activity over a range of pH values, salt concentrations, and temperatures. Both enzymes were haloalkaliphilic and thermoactive for the oxidative reaction and catalyzed the reductive reaction at a slightly acidic pH. While the NAD+-dependent HvADH1 showed a preference for short-chain alcohols and was inherently unstable, HvADH2 exhibited dual cofactor specificity, accepted a broad range of substrates, and, with respect to HvADH1, was remarkably stable. Furthermore, HvADH2 exhibited tolerance to organic solvents. HvADH2 therefore displays much greater potential as an industrially useful biocatalyst than HvADH1.




Similar content being viewed by others
References
Adams MWW, Perler FB, Kelly RM (1995) Extremozymes: expanding the limits of biocatalysis. Nat Biotechnol 13:662–668
Allers T, Ngo HP, Mevarech M, Lloyd RG (2004) Development of additional selectable markers for the halophilic archaeon Haloferax volcanii based on the leuB and trpA genes. Appl Environ Microbiol 70:943–953
Allers T, Barak S, Liddell S, Wardell K, Mevarech M (2010) Improved strains and plasmid vectors for conditional overexpression of his-tagged proteins in Haloferax volcanii. Appl Environ Microbiol 76:1759–1769
Baneyx F (1999) Recombinant protein expression in Escherichia coli. Curr Opin Biotechnol 10:411–421
Baneyx F, Mujacic M (2004) Recombinant protein folding and misfolding in Escherichia coli. Nat Biotechnol 22:1399–1408
Bitan-Banin G, Ortenberg R, Mevarech M (2003) Development of a gene knockout system for the halophilic archaeon Haloferax volcanii by use of the pyrE gene. J Bacteriol 185:772–778
Camacho M, Rodriguez-Arnedo A, Bonete M (2002) NADP-dependent isocitrate dehydrogenase from the halophilic archaeon Haloferax volcanii: cloning, sequence determination and overexpression in Escherichia coli. FEMS Microbiol Lett 209:155–160
Cao Y, Liao L, Xu X, Oren A, Wang C, Zhu X, Wu M (2008a) Characterization of alcohol dehydrogenase from the haloalkaliphilic archaeon Natronomonas pharaonis. Extremophiles 12:471–476
Cao Y, Liao L, Xu X, Oren A, Wu M (2008b) Aldehyde dehydrogenase of the haloalkaliphilic archaeon Natronomonas pharaonis and its function in ethanol metabolism. Extremophiles 12:849–854
Cendrin F, Chroboczek J, Zaccai G, Eisenberg H, Mevarech M (1993) Cloning, sequencing, and expression in Escherichia coli of the gene coding for malate dehydrogenase of the extremely halophilic archaebacterium Haloarcula marismortui. Biochemistry 32:4308–4313
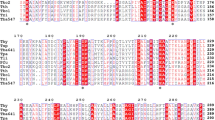
Chenna R, Sugawara H, Koike T, Lopez R, Gibson TJ, Higgins DG, Thompson JD (2003) Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res 31:3497–3500
Connaris H, Chaudhuri JB, Danson MJ, Hough DW (1999) Expression, reactivation, and purification of enzymes from Haloferax volcanii in Escherichia coli. Biotechnol Bioeng 64:38–45
Coquelle N, Talon R, Juers DH, Girard E, Kahn R, Madern D (2010) Gradual adaptive changes of a protein facing high salt concentrations. J Mol Biol 404:493–505
Danson MJ, Hough DW (1998) Structure, function and stability of enzymes from the Archaea. Trends Microbiol Rev 6:307–314
Dym O, Mevarech M, Sussman JL (1995) Structural features that stabilize halophilic malate dehydrogenase from an archaebacterium. Science 267:1344–1346
Eichler J (2001) Biotechnological uses of archaeal extremozymes. Biotechnol Adv 19:261–278
Flam F (1994) The chemistry of life at the margins. Science 265:471–472
Friest JA, Maezato Y, Broussy S, Blum P, Berkowitz DB (2010) Use of a robust dehydrogenase from an archael hyperthermophile in asymmetric catalysis-dynamic reductive kinetic resolution entry into (S)-profens. J Am Chem Soc Commun 132:5930–5931
Giacomini D, Galletti P, Quintavalla A, Gucciardo G, Paradisi F (2007) Highly efficient asymmetric reduction of arylpropionic aldehydes by horse liver alcohol dehydrogenase through dynamic kinetic resolution. Chem Commun 21:4038–4040
Goldberg K, Schroer K, Lütz S, Liese A (2007) Biocatalytic ketone reduction—a powerful tool for the production of chiral alcohols—part I: processes with isolated enzymes. Appl Microbiol Biotechnol 76:237–248
Gray CJ (1988) Additives and enzyme stability. Biocatal Biotrans 1:187–196
Guy CP, Haldenby S, Brindley A, Walsh DA, Briggs GS, Warren MJ, Allers T, Bolt EL (2006) Interactions of RadB, a DNA repair protein in archaea, with DNA and ATP. J Mol Biol 358:46–56
Hartman AL, Norais C, Badger JH, Delmas S, Haldenby S, Madupu R, Robinson J, Khouri H, Ren Q, Lowe TM, Maupin-Furlow J, Pohlschroder M, Daniels C, Pfeiffer F, Allers T, Eisen JA (2010) The complete genome sequence of Haloferax volcanii DS2, a model archaeon. PLoS One 5:e9605
Hough DW, Danson MJ (1999) Extremozymes. Curr Opin Chem Biol 3:39–46
Huisman GW, Liang J, Krebber A (2010) Practical chiral alcohol manufacture using ketoreductases. Curr Opin Chem Biol 14:122–129
Lesk AM (1995) NAD-binding domains of dehydrogenases. Curr Opin Struct Biol 5:775–783
McKinley-McKee J (1964) The mechanism of action and the active centre of the alcohol dehydrogenases. Progr Biophys Mol Biol 14:225–262
Norais C, Hawkins M, Hartman AL, Eisen JA, Myllykallio H, Allers T (2007) Genetic and physical mapping of DNA replication origins in Haloferax volcanii. PLoS Genet 3:e77
Oren A (2002) Diversity of halophilic microorganisms: environments, phylogeny, physiology, and applications. J Ind Microbiol Biotechnol 28:56–63
Parkot J, Gröger H, Hummel W (2010) Purification, cloning, and overexpression of an alcohol dehydrogenase from Nocardia globerula reducing aliphatic ketones and bulky ketoesters. Appl Microbiol Biotechnol 86:1813–1820
Pire C, Esclapez J, Diaz S, Perez-Pomares F, Ferrer J, Bonete MJ (2009) Alteration of coenzyme specificity in halophilic NAD(P)+ glucose dehydrogenase by site-directed mutagenesis. J Mol Catal B: Enzym 59:261–265
Schmid RD (1979) Stabilized soluble enzymes. Adv Biochem Eng Biotechnol 12:41–118
Sellek GA, Chaudhuri JB (1999) Biocatalysis in organic media using enzymes from extremophiles. Enzyme Microb Technol 25:471–482
Singh SM, Panda AK (2005) Solubilization and refolding of bacterial inclusion body proteins. J Biosci Bioeng 99:303–310
Timpson LM, Alsafadi D, Mac Donnchadha C, Liddell S, Sharkey MA, Paradisi F (2012) Characterization of alcohol dehydrogenase (ADH12) from Haloarcula marismortui, an extreme halophile from the Dead Sea. Extremophiles 16:57–66
Vallee BL, Hoch FL (1957) Zinc in horse liver alcohol dehydrogenase. J Biol Chem 225:185–195
Wang W (2000) Lyophilization and development of solid protein pharmaceuticals. Int J Pharm 203:1–60
Wierenga RK, De Maeyer MCH, Hol WGJ (1985) Interaction of pyrophosphate moieties with alpha-helixes in dinucleotide-binding proteins. Biochemistry 24:1346–1357
Wilkinson GN (1961) Statistical estimations in enzyme kinetics. Biochem J 80:324–332
Yakushi T, Matsushita K (2010) Alcohol dehydrogenase of acetic acid bacteria: structure, mode of action, and applications in biotechnology. Appl Microbiol Biotechnol 86:1257–1265
Acknowledgments
This work was supported by a research grant awarded to F.P. by Science Foundation Ireland (SFI), a University Research Fellowship awarded to T.A. by the Royal Society, and funding provided by the Islamic Development Bank (IsDB).
Author information
Authors and Affiliations
Corresponding authors
Additional information
L. M. Timpson and A.-K. Liliensiek contributed equally to this work.
Rights and permissions
About this article
Cite this article
Timpson, L.M., Liliensiek, AK., Alsafadi, D. et al. A comparison of two novel alcohol dehydrogenase enzymes (ADH1 and ADH2) from the extreme halophile Haloferax volcanii . Appl Microbiol Biotechnol 97, 195–203 (2013). https://doi.org/10.1007/s00253-012-4074-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-012-4074-4