Abstract
Background
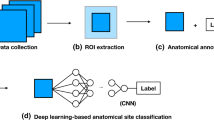
Esophagogastroduodenoscopy (EGD) is generally a safe procedure, but adverse events often occur. This highlights the necessity of the quality control of EGD. Complete visualization and photo documentation of upper gastrointestinal (UGI) tracts are important measures in quality control of EGD. To evaluate these measures in large scale, we developed an AI-driven quality control system for EGD through convolutional neural networks (CNNs) using archived endoscopic images.
Methods
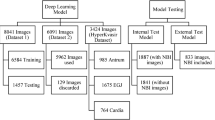
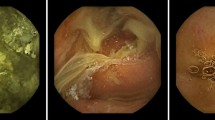
We retrospectively collected and labeled images from 250 EGD procedures, a total of 2599 images from eight locations of the UGI tract, using the European Society of Gastrointestinal Endoscopy (ESGE) photo documentation methods. The label confirmed by five experts was considered the gold standard. We developed a CNN model for multi-class classification of EGD images to one of the eight locations and binary classification of each EGD procedure based on its completeness.
Results
Our CNN model successfully classified the EGD images into one of the eight regions of UGI tracts with 97.58% accuracy, 97.42% sensitivity, 99.66% specificity, 97.50% positive predictive value (PPV), and 99.66% negative predictive value (NPV). Our model classified the completeness of EGD with 89.20% accuracy, 89.20% sensitivity, 100.00% specificity, 100.00% PPV, and 64.94% NPV. We analyzed the credibility of our model using a probability heatmap.
Conclusions
We constructed a CNN model that could be used in the quality control of photo documentation in EGD. Our model needs further validation with a large dataset, and we expect our model to help both endoscopists and patients by improving the quality of EGD procedures.






Similar content being viewed by others
References
Hamashima C, Systematic Review Group and Guideline Development Group for Gastric Cancer Screening Guidelines (2018) Update version of the Japanese guidelines for gastric cancer screening. Jpn J Clin Oncol 48:673–683
Jun JK, Choi KS, Lee HY, Suh M, Park B, Song SH, Jung KW, Lee CW, Choi IJ, Park EC, Lee D (2017) Effectiveness of the Korean National cancer screening program in reducing gastric cancer mortality. Gastroenterology 152:1319–1328
Choi KS, Jun JK, Park EC, Park S, Jung KW, Han MA, Choi IJ, Lee HY (2012) Performance of different gastric cancer screening methods in Korea: a population-based study. PLoS ONE 7:e50041
Choi KS, Suh M (2014) Screening for gastric cancer: the usefulness of endoscopy. Clin Endosc 47:490–496
ASGE Standards of Practice Committee, Ben-Menachem T, Decker GA, Early DS, Evans J, Fanelli RD, Fisher DA, Fisher L, Fukami N, Hwang JH, Ikenberry SO, Jain R, Jue TL, Khan KM, Krinsky ML, Malpas PM, Maple JT, Sharaf RN, Dominitz JA, Cash BD (2012) Adverse events of upper GI endoscopy. Gastrointest Endosc 76:707–718
Mema SC, Yang H, Vaska M, Elnitsky S, Jiang Z (2016) Integrated cancer screening performance indicators: a systematic review. PLoS ONE 11:e0161187
Rey JF, Lambert R, ESGE Quality Assurance Committee (2001) ESGE recommendations for quality control in gastrointestinal endoscopy: guidelines for image documentation in upper and lower GI endoscopy. Endoscopy 33:901–903
Marques S, Bispo M, Pimentel-Nunes P, Chagas C, Dinis-Ribeiro M (2017) Image documentation in gastrointestinal endoscopy: review of recommendations. GE Port J Gastroenterol 24:269–274
Bisschops R, Areia M, Coron E, Dobru D, Kaskas B, Kuvaev R, Pech O, Ragunath K, Weusten B, Familiari P, Domagk D, Valori R, Kaminski MF, Spada C, Bretthauer M, Bennett C, Senore C, Dinis-Ribeiro M, Rutter MD (2016) Performance measures for upper gastrointestinal endoscopy: a European Society of gastrointestinal endoscopy (ESGE) quality improvement initiative. Endoscopy 48:843–864
Buch VH, Ahmed I, Maruthappu M (2018) Artificial intelligence in medicine: current trends and future possibilities. Br J Gen Pract 68:143–144
Urban G, Tripathi P, Alkayali T, Mittal M, Jalali F, Karnes W, Baldi P (2018) Deep learning localizes and identifies polyps in real time with 96% accuracy in screening colonoscopy. Gastroenterology 155:1069–1078
Kominami Y, Yoshida S, Tanaka S, Sanomura Y, Hirakawa T, Raytchev B, Tamaki T, Koide T, Kaneda K, Chayama K (2016) Computer-aided diagnosis of colorectal polyp histology by using a real-time image recognition system and narrow-band imaging magnifying colonoscopy. Gastrointest Endosc 83:643–649
Chen PJ, Lin MC, Lai MJ, Lin JC, Lu HH, Tseng VS (2018) Accurate classification of diminutive colorectal polyps using computer-aided analysis. Gastroenterology 154:568–575
Nakashima H, Kawahira H, Kawachi H, Sakaki N (2018) Artificial intelligence diagnosis of Helicobacter pylori infection using blue laser imaging-bright and linked color imaging: a single-center prospective study. Ann Gastroenterol 31:462–468
Kanesaka T, Lee TC, Uedo N, Lin KP, Chen HZ, Lee JY, Wang HP, Chang HT (2018) Computer-aided diagnosis for identifying and delineating early gastric cancers in magnifying narrow-band imaging. Gastrointest Endosc 87:1339–1344
de Souza LA, Palm C, Mendel R, Hook C, Ebigbo A, Probst A, Messmann H, Weber S, Papa JP (2018) A survey on Barrett’s esophagus analysis using machine learning. Comput Biol Med 96:203–213
Trindade AJ, McKinley MJ, Fan C, Leggett CL, Kahn A, Pleskow DK (2019) Endoscopic surveillance of Barrett’s esophagus using volumetric laser endomicroscopy with artificial intelligence image enhancement. Gastroenterology 157:303–305
Paszke A, Gross S, Massa F, Lerer A, Bradbury J, Chanan G, Killeen T, Lin Z, Gimelshein N, Antiga L (2019) PyTorch: An imperative style, high-performance deep learning library. In: Advances in neural information processing systems, pp 8024–8035
Krizhevsky A, Sutskever I, Hinton G (2012) ImageNet classification with deep convolutional neural networks. In: Neural information processing systems, p 25
Hu J, Shen L, Albanie S, Sun G, Wu E (2019) Squeeze-and-excitation networks. In: IEEE transition pattern analytical machine intelligence
Gulshan V, Peng L, Coram M, Stumpe MC, Wu D, Narayanaswamy A, Venugopalan S, Widner K, Madams T, Cuadros J, Kim R, Raman R, Nelson PC, Mega JL, Webster DR (2016) Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. JAMA 316:2402–2410
Yosinski J, Clune J, Bengio Y, Lipson H (2014) How transferable are features in deep neural networks? Adv Neural Inf Process Syst 27:3320–3328
Kingma D, Ba J (2014) Adam: a method for stochastic optimization. In: International conference on learning representations
Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D (2017) Grad-CAM: visual explanations from deep networks via gradient-based localization. In: 2017 IEEE international conference on computer vision (ICCV), pp 618–626
Takiyama H, Ozawa T, Ishihara S, Fujishiro M, Shichijo S, Nomura S, Miura M, Tada T (2018) Automatic anatomical classification of esophagogastroduodenoscopy images using deep convolutional neural networks. Sci Rep 8:7497
Wu L, Zhang J, Zhou W, An P, Shen L, Liu J, Jiang X, Huang X, Mu G, Wan X, Lv X, Gao J, Cui N, Hu S, Chen Y, Hu X, Li J, Chen D, Gong D, He X, Ding Q, Zhu X, Li S, Wei X, Li X, Wang X, Zhou J, Zhang M, Yu HG (2019) Randomised controlled trial of WISENSE, a real-time quality improving system for monitoring blind spots during esophagogastroduodenoscopy. Gut 68:2161–2169
Chen D, Wu L, Li Y, Zhang J, Liu J, Huang L, Jiang X, Huang X, Mu G, Hu S, Hu X, Gong D, He X, Yu H (2020) Comparing blind spots of unsedated ultrafine, sedated, and unsedated conventional gastroscopy with and without artificial intelligence: a prospective, single-blind, 3-parallel-group, randomized, single-center trial. Gastrointest Endosc 91:332–339
Peery AF, Crockett SD, Murphy CC, Lund JL, Dellon ES, Williams JL, Jensen ET, Shaheen NJ, Barritt AS, Lieber SR, Kochar B, Barnes EL, Fan YC, Pate V, Galanko J, Baron TH, Sandler RS (2019) Burden and cost of gastrointestinal, liver, and pancreatic diseases in the United States: update 2018. Gastroenterology 156:254–272
Acknowledgements
This work was supported by National IT Industry Promotion Agency (A0105-20-1007) and Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2018R1D1A1B07048202).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Disclosures
Seong Ji Choi, Mohammad Azam Khan, Hyuk Soon Choi, Jaegul Choo, Jae Min Lee, Soonwook Kwon, Bora Keum, and Hoon Jai Chun have no conflicts of interest or financial ties to disclose.
Ethical approval
This study was conducted in accordance with the Helsinki Declaration, and the Ethics Committee of Korea University Anam Hospital approved this study (2019AN0253).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Choi, S.J., Khan, M.A., Choi, H.S. et al. Development of artificial intelligence system for quality control of photo documentation in esophagogastroduodenoscopy. Surg Endosc 36, 57–65 (2022). https://doi.org/10.1007/s00464-020-08236-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00464-020-08236-6