Abstract
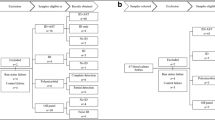
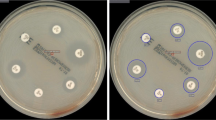
Delaying effective antibiotic therapy is a major cause of sepsis-associated mortality. The EUCAST rapid antibiotic susceptibility test (RAST) is performed from positive blood cultures to provide rapid results. Disc diffusion tests inoculated with positive blood culture broth are read at 4, 6, and 8 h and interpreted against species and time-specific criteria. Potential problems are the possibility of missing specific reading times for tests and slower growth in incubators that are frequently opened. The current study aimed to assess if digital visualization by the BD Kiestra™ total laboratory automation system is suitable for reading RASTs by capturing images at the correct times and retaining them for review. Utilizing the Kiestra™ InoqulA, 100 μl of positive blood culture broth was lawn-inoculated onto Mueller-Hinton agar and incubated at 35 °C for automated digital zone measurement at 4, 6, and 8 h. Aliquots from 135 positive blood cultures were tested against EUCAST-recommended and other drugs and assessed for readability of digital images. Microdilution MICs were determined in parallel to RASTs. All isolates except 7/10 enterococci yielded images of suitable quality for zone measurement. Of the 641 digitally read tests for other organisms, 207 (32.3%) were readable in 4 h, 555 (86.6%) in 6 h, and 641 (100%) in 8 h. For tests included in EUCAST criteria, 92.1% provided categorical agreement with microdilution MICs. Digital image reading of RASTs is a potentially viable, inexpensive tool for providing rapid susceptibility results which can help reduce sepsis-associated mortality.


Similar content being viewed by others
References
Reinhart K, Daniels R, Kissoon N, Machado FR, Schachter RD, Finfer S (2017) Recognizing sepsis as a global health priority - a WHO resolution. N Engl J Med 377(5):414–417. https://doi.org/10.1056/NEJMp1707170
Singer M, Deutschman CS, Seymour CW, Shankar-Hari M, Annane D, Bauer M, Bellomo R, Bernard GR, Chiche JD, Coopersmith CM, Hotchkiss RS, Levy MM, Marshall JC, Martin GS, Opal SM, Rubenfeld GD, van der Poll T, Vincent JL, Angus DC (2016) The third international consensus definitions for sepsis and septic shock (Sepsis-3). JAMA 315(8):801–810. https://doi.org/10.1001/jama.2016.0287
Rhee C, Dantes R, Epstein L, Murphy DJ, Seymour CW, Iwashyna TJ, Kadri SS, Angus DC, Danner RL, Fiore AE, Jernigan JA, Martin GS, Septimus E, Warren DK, Karcz A, Chan C, Menchaca JT, Wang R, Gruber S, Klompas M, Program CDCPE (2017) Incidence and trends of sepsis in US hospitals using clinical vs claims data, 2009-2014. JAMA 318(13):1241–1249. https://doi.org/10.1001/jama.2017.13836
Dantes RB, Epstein L (2018) Combatting sepsis: a public health perspective. Clin Infect Dis 67(8):1300–1302. https://doi.org/10.1093/cid/ciy342
Rudd KE, Johnson SC, Agesa KM, Shackelford KA, Tsoi D, Kievlan DR, Colombara DV, Ikuta KS, Kissoon N, Finfer S, Fleischmann-Struzek C, Machado FR, Reinhart KK, Rowan K, Seymour CW, Watson RS, West TE, Marinho F, Hay SI, Lozano R, Lopez AD, Angus DC, Murray CJL, Naghavi M (2020) Global, regional, and national sepsis incidence and mortality, 1990-2017: analysis for the Global Burden of Disease Study. Lancet 395(10219):200–211. https://doi.org/10.1016/S0140-6736(19)32989-7
Evans IVR, Phillips GS, Alpern ER, Angus DC, Friedrich ME, Kissoon N, Lemeshow S, Levy MM, Parker MM, Terry KM, Watson RS, Weiss SL, Zimmerman J, Seymour CW (2018) Association between the New York sepsis care mandate and in-hospital mortality for pediatric sepsis. JAMA 320(4):358–367. https://doi.org/10.1001/jama.2018.9071
Rhodes A, Evans LE, Alhazzani W, Levy MM, Antonelli M, Ferrer R, Kumar A, Sevransky JE, Sprung CL, Nunnally ME, Rochwerg B, Rubenfeld GD, Angus DC, Annane D, Beale RJ, Bellinghan GJ, Bernard GR, Chiche JD, Coopersmith C, De Backer DP, French CJ, Fujishima S, Gerlach H, Hidalgo JL, Hollenberg SM, Jones AE, Karnad DR, Kleinpell RM, Koh Y, Lisboa TC, Machado FR, Marini JJ, Marshall JC, Mazuski JE, McIntyre LA, McLean AS, Mehta S, Moreno RP, Myburgh J, Navalesi P, Nishida O, Osborn TM, Perner A, Plunkett CM, Ranieri M, Schorr CA, Seckel MA, Seymour CW, Shieh L, Shukri KA, Simpson SQ, Singer M, Thompson BT, Townsend SR, Van der Poll T, Vincent JL, Wiersinga WJ, Zimmerman JL, Dellinger RP (2017) Surviving sepsis campaign: international guidelines for management of sepsis and septic shock: 2016. Crit Care Med 45(3):486–552. https://doi.org/10.1097/CCM.0000000000002255
Weinberger J, Rhee C, Klompas M (2020) A critical analysis of the literature on time-to-antibiotics in suspected sepsis. J Infect Dis 222(Supplement_2):S110–S1S8. https://doi.org/10.1093/infdis/jiaa146
Minejima E, Mai N, Bui N, Mert M, Mack WJ, She RC, Nieberg P, Spellberg B, Wong-Beringer A (2020) Defining the breakpoint duration of Staphylococcus aureus bacteremia predictive of poor outcomes. Clin Infect Dis 70(4):566–573. https://doi.org/10.1093/cid/ciz257
Martinez-Nadal G, Puerta-Alcalde P, Gudiol C, Cardozo C, Albasanz-Puig A, Marco F, Laporte-Amargos J, Moreno-Garcia E, Domingo-Domenech E, Chumbita M, Martinez JA, Soriano A, Carratala J, Garcia-Vidal C (2020) Inappropriate empirical antibiotic treatment in high-risk neutropenic patients with bacteremia in the era of multidrug resistance. Clin Infect Dis 70(6):1068–1074. https://doi.org/10.1093/cid/ciz319
Spanik S, Kukuckova E, Pichna P, Grausova S, Krupova I, Rusnakova V, Kralovicova K, Krchnakova A, Mrazova M, Lacka J, Koren P, Stopkova K, Nogova J, Demitrovicova A, Helpianska L, Krcmery V Jr (1997) Analysis of 553 episodes of monomicrobial bacteraemia in cancer patients: any association between risk factors and outcome to particular pathogen? Support Care Cancer 5(4):330–333. https://doi.org/10.1007/s005200050083
Rodriguez-Bano J, Perez-Moreno MA, Penalva G, Garnacho-Montero J, Pinto C, Salcedo I, Fernandez-Urrusuno R, Neth O, Gil-Navarro MV, Perez-Milena A, Sierra R, Estella A, Lupion C, Irastorza A, Marquez JL, Pascual A, Rojo-Martin MD, Perez-Lozano MJ, Valencia-Martin R, Cisneros JM, Group PP (2020) Outcomes of the PIRASOA programme, an antimicrobial stewardship programme implemented in hospitals of the Public Health System of Andalusia, Spain: an ecologic study of time-trend analysis. Clin Microbiol Infect 26(3):358–365. https://doi.org/10.1016/j.cmi.2019.07.009
Eubank TA, Long SW, Perez KK (2020) Role of rapid diagnostics in diagnosis and management of patients with sepsis. J Infect Dis 222(Supplement_2):S103–S1S9. https://doi.org/10.1093/infdis/jiaa263
Mangioni D, Peri AM, Rossolini GM, Viaggi B, Perno CF, Gori A, Bandera A (2020) Toward rapid sepsis diagnosis and patient stratification: what’s new from microbiology and omics science. J Infect Dis 221(7):1039–1047. https://doi.org/10.1093/infdis/jiz585
Grumaz C, Hoffmann A, Vainshtein Y, Kopp M, Grumaz S, Stevens P, Decker SO, Weigand MA, Hofer S, Brenner T, Sohn K (2020) Rapid next-generation sequencing-based diagnostics of bacteremia in septic patients. J Mol Diagn 22(3):405–418. https://doi.org/10.1016/j.jmoldx.2019.12.006
Strich JR, Heil EL, Masur H (2020) Considerations for empiric antimicrobial therapy in sepsis and septic shock in an era of antimicrobial resistance. J Infect Dis 222(Supplement_2):S119–SS31. https://doi.org/10.1093/infdis/jiaa221
Ehren K, Meissner A, Jazmati N, Wille J, Jung N, Vehreschild JJ, Hellmich M, Seifert H (2020) Clinical impact of rapid species identification from positive blood cultures with same-day phenotypic antimicrobial susceptibility testing on the management and outcome of bloodstream infections. Clin Infect Dis 70(7):1285–1293. https://doi.org/10.1093/cid/ciz406
Posteraro P, Gasparoli C, Errico FM, Massaria G, Sanguinetti M, Posteraro B (2019) Off-site versus on-site clinical microbiology laboratory: a 2-year comparison study of blood culture result reporting. Clin Microbiol Infect 25(11):1441–1442. https://doi.org/10.1016/j.cmi.2019.06.021
Opota O, Croxatto A, Prod’hom G, Greub G (2015) Blood culture-based diagnosis of bacteraemia: state of the art. Clin Microbiol Infect 21(4):313–322. https://doi.org/10.1016/j.cmi.2015.01.003
Jonasson E, Matuschek E, Kahlmeter G (2020) The EUCAST rapid disc diffusion method for antimicrobial susceptibility testing directly from positive blood culture bottles. J Antimicrob Chemother 75(4):968–978. https://doi.org/10.1093/jac/dkz548
Jasuja JK, Zimmermann S, Burckhardt I (2020) Evaluation of EUCAST rapid antimicrobial susceptibility testing (RAST) for positive blood cultures in clinical practice using a total lab automation. Eur J Clin Microbiol Infect Dis 39(7):1305–1313. https://doi.org/10.1007/s10096-020-03846-3
Bourbeau PP, Ledeboer NA (2013) Automation in clinical microbiology. J Clin Microbiol 51(6):1658–1665. https://doi.org/10.1128/JCM.00301-13
Croxatto A, Marcelpoil R, Orny C, Morel D, Prod’hom G, Greub G (2017) Towards automated detection, semi-quantification and identification of microbial growth in clinical bacteriology: a proof of concept. Biom J 40(6):317–328. https://doi.org/10.1016/j.bj.2017.09.001
Bailey AL, Ledeboer N, Burnham CD (2019) Clinical microbiology is growing up: the total laboratory automation revolution. Clin Chem 65(5):634–643. https://doi.org/10.1373/clinchem.2017.274522
Clinical and Laboratory Standards Institute (2020) CLSI M100-ED30:2020 performance standards for antimicrobial susceptibility testing, 30th edn. In. Clinical and Laboratory Standards Institute, Wayne, PA
Osih RB, McGregor JC, Rich SE, Moore AC, Furuno JP, Perencevich EN, Harris AD (2007) Impact of empiric antibiotic therapy on outcomes in patients with Pseudomonas aeruginosa bacteremia. Antimicrob Agents Chemother 51(3):839–844
Lodise TP Jr, Patel N, Kwa A, Graves J, Furuno JP, Graffunder E, Lomaestro B, McGregor JC (2007) Predictors of 30-day mortality among patients with Pseudomonas aeruginosa bloodstream infections: impact of delayed appropriate antibiotic selection. Antimicrob Agents Chemother 51(10):3510–3515
Morata L, Cobos-Trigueros N, Martinez JA, Soriano A, Almela M, Marco F, Sterzik H, Nunez R, Hernandez C, Mensa J (2012) Influence of multidrug resistance and appropriate empirical therapy on the 30-day mortality rate of Pseudomonas aeruginosa bacteremia. Antimicrob Agents Chemother 56(9):4833–4837. https://doi.org/10.1128/AAC.00750-12
Tofas P, Samarkos M, Piperaki ET, Kosmidis C, Triantafyllopoulou ID, Kotsopoulou M, Pantazatou A, Perlorentzou S, Poulli A, Vagia M, Daikos GL (2017) Pseudomonas aeruginosa bacteraemia in patients with hematologic malignancies: risk factors, treatment and outcome. Diagn Microbiol Infect Dis 88(4):335–341. https://doi.org/10.1016/j.diagmicrobio.2017.05.003
Tacconelli E, Cataldo MA, Mutters NT, Carrara E, Bartoloni A, Raglio A, Cauda R, Mantengoli E, Luzzaro F, Pan A, Beccara LA, Pecile P, Tinelli M, Rossolini GM (2019) Role of place of acquisition and inappropriate empirical antibiotic therapy on the outcome of extended-spectrum beta-lactamase-producing Enterobacteriaceae infections. Int J Antimicrob Agents 54(1):49–54. https://doi.org/10.1016/j.ijantimicag.2019.04.007
Acknowledgements
We thank our colleagues at the Microbiology Department of the University of Louisville Hospital, Louisville, KY, USA, for the assistance with the conduct of this study.
Availability of data
The datasets generated and/or analyzed during the current study are available from KST (kenneth.thomson20@gmail.com) on reasonable request.
Author information
Authors and Affiliations
Contributions
The contributions from the respective authors were as follows: Thomson, G. and Jamros, K, design and technical oversight of the study, data acquisition, analysis and interpretation of data, drafting and revising the manuscript for intellectual content, and final approval for submission; Thomson, K., analysis and interpretation of data, drafting and revising the manuscript for intellectual content, and final approval for submission; Snyder, J., analysis and interpretation of data, drafting and revising the manuscript for intellectual content, and final approval for submission.
Corresponding author
Ethics declarations
Ethical approval
This study does not contain any studies with human participants performed by any of the authors.
Consent to participate
Not applicable. This study does not contain any studies with human participants.
Consent to publish
Not applicable. The authors have no constraints to declare concerning the publication of this manuscript.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Thomson, G.K., Jamros, K., Snyder, J.W. et al. Digital imaging for reading of direct rapid antibiotic susceptibility tests from positive blood cultures. Eur J Clin Microbiol Infect Dis 40, 2105–2112 (2021). https://doi.org/10.1007/s10096-021-04249-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-021-04249-8