Abstract
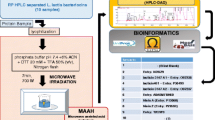
An analysis strategy that can simultaneously determine the amino acid site information and configuration for polymyxin (PM) components is reported in this study. The main components of PMB1, PMB2, and PMB1-I (the Leucine at position 7 changed to be Isoleucine) were purified from the mixture of PMB. The PMB component was hydrolyzed by hydrochloric acid to be a mixture of amino acids, which were derivatized with Marfey’s reagent of Nα-(5-fluoro-2,4-dinitrophenyl)-l-alanine amide (l-FDAA), and then, the configurations of amino acids were easily determined by HPLC–MS. However, the aforementioned method could not offer the information of amino acid at each site, a specific enzymatic hydrolysis method combined Edman degradation was developed. The fatty acyl-diaminobutyric acid (position 1) of PMB component was removed by ficin, and the PMB components were transformed to be PMB nonapeptide. The amino acids at each site of the nonapeptide were determined by N-terminal sequencing. Combining the results of the above methods, the amino acid information and configuration of each site in PM components can be determined, especially the l-Leucine and l-Isoleucine in PMB1 and PMB1-I. The workflow was useful for other PM analogues to identify the sequence and configuration for quality control.








Similar content being viewed by others
References
World Health Organization (2017) Global priority list of antibiotic-resistant bacteria guide to guide research, discovery, and development of new antibiotics. World Health Organization, Geneva
Nang SC, Azad MAK, Velkov T, Zhou Q, Li J (2021) Rescuing the last-line polymyxins: achievements and challenges. Pharmacol Rev 73:679–728
European Pharmacopoeia (2020) Specific monograph: directorate for the quality of medicines and healthcare of the Council of Europe. Strasbourg 10:3587–3588
Bossche LVD, Schepdael AV, Chopra S, Hoogmartens J, Adams E (2011) Identification of impurities in polymyxin B and colistin bulk sample using liquid chromatography coupled to mass spectrometry. Talanta 83:1521–1529
Brown P, Dawson MJ (2017) Development of new polymyxin derivatives for multi-drug-resistant Gram-negative infections. J Antibiot 70:386–394
Cui AL, Hu XX, Gao Y, Jin J, Yi H, Wang XK, Nie TY, Chen Y, He QY, Guo HF, Jiang JD, You XF, Li ZR (2018) Synthesis and bioactivity investigation of the individual components of cyclic lipopeptides antibiotics. J Med Chem 61:1845–1857
Tang HQ, Zhang Y, Ma J, Dong YZ, Gao Q, Feng J (2020) Design, synthesis and antimicrobial studies of some polymyxin analogues. J Antibiot 73:158–166
Lepak A, Wang W, Andes DR (2021) Pharmacodynamic evaluation of MRX-8, a novel polymyxin, in the neutropenic mouse thigh and lung infection models against Gram-negative pathogens. Antimicrob Agents Chemother. https://doi.org/10.1128/AAC.01517-20
Velkov T, Gallardo-Godoy A, Swarbrick JD, Blaskovich MAT, Elliott AG, Han ML, Thompson PE, Roberts KD, Huang JX, Becker B, Butler MS, Lash LH, Henriques ST, Nation RL, Sivanesan S, Sani MA, Separovic F, Mertens H, Bulach D, Seemann T, Owen J, Li J, Cooper MA (2018) Structure, function, and biosynthetic origin of octapeptin antibiotics active against extensively drug-resistant Gram-negative bacteria. Cell Chem Biol 25:380–391
Shaheen M, Li JR, Ross AC, Vederas JC, Jensen SE (2011) Paenibacillus polymyxa PKB1 produces variants of polymyxin B-type antibiotics. Chem Biol 18:1640–1648
Zhang HZ, Qin F, Liu H (2018) Identification of unknown impurities in polymyxin B sulfate by HPLC–MS/MS. Chin Pharm J 53:918–924
Zhang HZ, Sun N, Qin F, Luo WY, Zhao JD, Wen HL, Qiu Y, Liu H, Li M (2021) Structure analysis of components in polymyxin E sulfate by high performance liquid chromatography—quadrupole/time of flight mass spectrometry. Chin Pharm J 56:54–62
Hausmann W, Craig LC (1954) Polymyxin B1, fractionation, molecular weight determination, amino acid and fatty acid composition. J Am Chem Soc 76:4892–4896
Hausmann W (1956) The amino acid sequence of polymyxin B1. J Am Chem Soc 78:3663–3667
Wu Y, Zhou MY, Xue CJ, Feng J (2018) Purification and structure elucidation of colistin E analogues. Chin J Antibiot 43:59–63
Lobas AA, Verenchikov AN, Goloborodko AA, Levitsky LI, Gorshkov MV (2012) Combination of Edman degradation of peptides with liquid chromatography/mass spectrometry workflow for peptide identification in bottom-up proteomics. Rapid Commun Mass Spectrom 27:391–400
Chihara S, Tobita T, Yahata M, Ito A, Koyama Y (1973) Enzymatic degradation of colistin isolation and identification of α-N-acyl α, γ-diaminobutyric acid and colistin nonapeptide. Arg Biol Chem 37:2455–2463
Kimura Y, Matsunaga H, Yasuda N, Tatsuki T, Suzuki T (1987) Polymyxin acylase: a new enzyme for preparing starting materials for semisynthetic polymyxin antibiotics. Arg Biol Chem 51:1617–1623
Acknowledgements
This work was financially supported by the Science and Technology Commission of Shanghai Municipality (No. 20142202200) and Shanghai Institute for Food and Drug Control (No. 2021-YKT-17).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing financial interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, H., Li, F., Dun, J. et al. Combination of Derivatization–HPLC–MS and Enzymatic Hydrolysis–Edman Degradation for Amino Acid Sequence and Configuration of Polymyxin B Components. Chromatographia 84, 1057–1064 (2021). https://doi.org/10.1007/s10337-021-04091-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10337-021-04091-2