Abstract
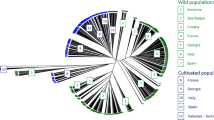
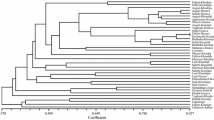
In the second half of the nineteenth century, intensive renovation of vineyards took place due to the losses caused by phylloxera and local varieties were mostly replaced by several worldwide cultivars. Shift in genotypic structure in favor of modern cultivars resulted in the decrease or even disappearance of regionally typical local varieties. A total of sixty five Turkish grape genotypes, including 5 references (four cultivars and one rootstock), were genotyped with 16 SSR and 15 SRAP markers. Sixteen SSR primers generated a total of 60 SSR amplicons in which 43 were polymorphic with 73.4% average polymorphism percentage. A total of 111 well-resolved clear DNA bands were obtained from 15 SRAP primers. Of these bands, 53 were highly polymorphic with an average of 47.74%. Cluster analysis based on pooled marker data generated a well resolved grouping pattern. The analyzed genotypes grouped basing on their geographical belongings. There were many cultivar pairs on the dendrogram most of which occurred between 0.75 and 0.90 levels. SSR and SRAP data revealed a wide genetic variability as well as certain synonyms among the historical grape varieties cultivated for decades in local vineyards lengthwise the mountainous regions of Konya, Karaman and Mersin provinces. All the genotypes have been maintained in a grapevine germplasm glasshouse. Preservation and use of these endangered genotypes will be helpful to avoid genetic erosion and diversity loss in this part of Turkey. Also, the molecular data generated in this study could be of great use in determining the optimal breeding strategies to allow continued progress in grapevine breeding.

Similar content being viewed by others
References
Aliscioni SS, Giussani LM, Zuloaga FO, Kellogg EA (2003) A molecular phylogeny of Panicum (Poaceae: Paniceae): tests of monophyly and phylogenetic placement within the Panicoideae. Am J Bot 90:796–821
Alleweldt G, Possingham JV (1988) Progress in grapevine breeding. Theor Appl Genet 75:669–673
Alleweldt G, Spiegel-Roy P, Reisch B (1990) Grapes (Vitis). In: Moore JN, Ballington JR Jr (eds.) Genetic resources of temperate fruit and nut crops, vol I. ISHS, Leuven, pp 291–327
Almadanim MC, Baleiras-Couto MM, Pereira HS, Carneiro LC, Fevereiro P, Eiras-Dias JE, Morais-Cecilio L, Viegas W, Veloso MM (2007) Genetic diversity for the grapevine (Vitis vinifera L.) cultivars most utilized for wine production in Portugal. Vitis 46:116–119
Amar MH, Biswas MK, Zhang Z, Guo W (2011) Exploitation of SSR, SRAP and CAPS-SNP markers for genetic diversity of Citrus germplasm collection. Sci Hortic 128:220–227
Aradhya MK, Dangl GS, Prins BH, Boursiquot JM, Walker MA, Meredith CP, Simon CJ (2003) Genetic structure and differentiation in cultivated grape, Vitis vinifera L. Genet Res 81:179–192
Arroyo-Garcia R, Lefort F, Andres MTD, Ibanez J, Borrego J, Jouve N, Vabello F, Martinez-Zapater JM (2002) Chloroplast microsatellite polymorphisms in Vitis species. Genome 45:1142–1149
Bacu A, Bajraktari Y, Papa S, Thomaj F, Michailidou S, Argyriou A (2015) Characterization of main grapevine varieties of Albania and Kosovo based on molecular data. Vitis 54:91–92
Bowers JE, Dangl GS, Vignani R, Meredith CP (1996) Isolation and characterization of new polymorphic simple sequence repeat loci in grape (Vitis vinifera L). Genome 39:628–633
Bowers JE, Dangl GS, Meredith CP (1999) Development and characterization of additional microsatellite DNA markers for grape. Am J Enol Vitic 50:243–246
Chen W, Zhang Y, Liu X, Chen B, Tu J, Tingdong F (2007) Detection of QTL for six yield-related traits in oilseed rape (Brassica napus) using DH and immortalized F(2) populations. Theor Appl Genet 115:849–858
Cipriani G, Spadotto A, Jurman I, di Gaspero G, Crespan M, Meneghetti S, Frare E, Vignani R, Cresti M, Morgante M, Pezzotti M, Pe E, Policriti A, Testolin A (2010) The SSR-based molecular profile of 1005 grapevine (Vitis vinifera L.) accessions uncovers new synonymy and parentages, and reveals a large admixture amongst varieties of different geographic origin. Theor Appl Genet 12:1569–1585
Crespan M, Botta R, Milani N (1999) Molecular characterization of twenty seeded and seedless table grape cultivars grape (Vitis vinifera L.). Vitis 38:87–92
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus (Madison) 12:13–15
Fan X, Jiang J, Zhang Y, Sun H, Jiao J, Liu C (2015) Genetic diversity assessment of Vitis ficifolia Bge. populations from Henan province of China by SRAP markers. Biotechnol Biotechnol Equip 29:15–20
Guo D, Zhang J, Liu C, Zhang G, Li M, Zhang Q (2012) Genetic variability and relationships between and within grape cultivated varieties and wild species based on SRAP markers. Tree Genet Genomes 8:789–800
Huang LK, Bughrara SS, Zhang XQ, Bales-Arcelo CJ, Bin X (2011) Genetic diversity of switchgrass and its relative species in Panicum genus using molecular markers. Biochem Syst Ecol 39:685–693
Ibanĕz JM, Andrés T, Molino A, Borrego J (2003) Genetic study of key Spanish grapevine varieties using microsatellites analysis. Am J Enol Vitic 54:22–29
Kayesh E, Zhang YY, Liu GS, Bilkish N, Sun X, Leng XP, Fang JG (2013) Development of highly polymorphic EST-SSR markers and segregation in F1 hybrid population of Vitis vinifera L. Genet Mol Res 12:3871–3878
Laiadi Z, Bentchikou MM, Bravo G, Cabello F, Martinez-Zapater JM (2009) Molecular identification and genetic relationship of Algerian grapevine cultivars maintained at the germplasm collection of Skikda (Algeria). Vitis 48:25–32
Levi A, Thomas CE (2007) DNA markers from different linkage regions of watermelon genome useful in differentiating among closely related watermelon genotypes. HortScience 42:210–214
Li G, Quiros CF (2001) Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: Its application to mapping and gene tagging in Brassica. Theor Appl Genet 103:455–461
Li Y, Liu Z, Wang Y, Yang N, Xin X, Yang S, Feng H (2012) Identification of quantitative trait loci for yellow inner leaves in Chinese cabbage (Brassica rapa L. ssp. pekinensis) based on SSR and SRAP markers. Sci Hortic 133:10–17
Lou Y, Lin X, He Q, Guo X, Huang L, Fang W (2010) Analysis on genetic relationship of Puji-bamboo species by AFLP and SRAP. Mol Plant Breed 8:83–88
Lópes MS, Sefc KM, Dias EE, Steinkellner H, Laimer da Cámara Machado M, da Cámara Machado A (1999) The use of microsatellites for germplasm management in a Portuguese grapevine collection. Theor Appl Genet 99:733–739
Martinez LE, Cavagnaro PF, Masuelli RW, Zuniga M (2006) SSR-based assessment of genetic diversity in South American Vitis vinifera varieties. Plant Sci 170:1036–1044
Merkouropoulos G, Michailidou S, Alifragkis A, Zioziou E, Koundouras S, Argiriou A, Nikolaou N (2015) A combined approach involving ampelographic description, berry oenological traits and molecular analysis to study native grapevine varieties of Greece. Vitis 54:99–103
Rohlf FJ (1993) NTSYS-PC, numerical taxonomy and multivariate analysis system. Version 1.8. Exeter Software, Setauket
Sabir A (2008) Ampelographic and molecular characterization of some grape cultivars and rootstocks. Doctorate Dissertation, Cukurova Univeristy
Sabir A, Kafkas S, Tangolar S, Buyukalaca S (2008) Genetic relationship of grape cultivars by ISSR (Inter-Simple Sequence Repeats) markers. Eur J Hortic Sci 73:84–88
Sabir A, Tangolar S, Buyukalaca S, Kafkas S (2009) Ampelographic and molecular diversity among grapevine (Vitis spp.) cultivars. Czech J Genet Plant Breed 45:160–168
Sefc KM, López MS, Lefort F, Botta R, Roubelakis-Angelakis KA, Ibanĕz J, Pejic I, Wagner HW, Glóssl J, Steinkellner H (2000) Microsatellites variability in grapevine cultivars from different European regions and evaluation of assignment testing to assess the geographic origin of cultivars. Theor Appl Genet 100:498–505
Štajner N, Tomić L, Ivanišević D, Korać N, Cvetković-Jovanović T, Beleski K, Angelova E, Maraš V, Javornik B (2014) Microsatellite inferred genetic diversity and structure of Western Balkan grapevines (Vitis vinifera L.). Tree Genet Genom 10:127–140
This P, Jung A, Boccacci P, Borrego J, Botta R, Costantini L, Crespan M, Dangl GS, Eisenheld C, FereriraMonterio F, Grando S, Ibanez J, Lacombe T, Laucou V, Magalhaes R, Meredith CP, Milani N, Peterlunger E, Regner F, Zulini L, Maul E (2004) Development of a standard set of microsatellite reference alleles for identification of grape cultivars. Theor Appl Genet 109:1448–1458
This P, Lacombe T, Thomas MR (2006) Historical origins and genetic diversity of wine grapes. Trends Genet 22:511–519
Uzun A, Yesiloglu T, Aka-Kacar Y, Tuzcu O, Gulsen O (2009) Genetic diversity and relationships within Citrus and related genera based on sequence related amplified polymorphism markers (SRAPs). Sci Hortic 121:306–312
Vezzulli S, Michela T, Giuseppina C, Angelica J, Dustin C, Andrey Z (2008) A reference integrated map for cultivated grapevine (Vitis vinifera L.) from three crosses based on 283 SSR and 501 SNP-based markers. Theor Appl Genet 117:499–511
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
A. Sabir, H. Ikten, N. Mutlu and D. Sari declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Sabir, A., Ikten, H., Mutlu, N. et al. Genetic Identification and Conservation of Local Turkish Grapevine (Vitis vinifera L.) Genotypes on the Edge of Extinction. Erwerbs-Obstbau 60, 31–38 (2018). https://doi.org/10.1007/s10341-017-0335-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10341-017-0335-9