Abstract
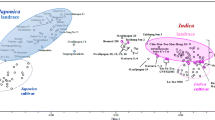
Information on genotype diversity and interrelationships is essential for germplasm management, crop improvement and seed production. In this study the diversity of a collection of 36 rice accessions comprising of Oryza sativa L., O. glaberrima Steud. and interspecific rice (O. sativa × O. glaberrima) was assessed using 24 agro-morphological and 27 microsatellite markers. The collection was composed mainly of improved cultivars and a few landraces which are adapted to the upland ecology in West Africa. The average Shannon–Weaver diversity index value (H′) was 0.23. The highest diversity was found in O. sativa group Indica (H′ = 0.32), and the lowest (H′ = 0.18) in O. glaberrima. Interspecific rice had intermediate diversity. There were 321 alleles with an average of 11.9 alleles per locus and an average major allele frequency of 0.29 per locus. The average gene diversity value was 0.81 and polymorphism information content was 0.80 per locus. Diversity indices for interspecific rice were intermediate between O. sativa and O. glaberrima, but closer to O. sativa group Japonica. Accessions comprised of two populations which corresponded to O. sativa group Indica and O. sativa group Japonica. Interspecific rice accessions were dispersed between the two groups.



Similar content being viewed by others
References
Babaei A, Nematzadeh GA, Hashemi H (2011) An evaluation of genetic differentiation in rice mutants using semi-random markers and morphological characteristics. Aust J Crop Sci 5:1715–1722
Baek HJ, Beharav A, Nevo E (2003) Ecological-genomic diversity of microsatellites in wild barley, Hordeum spontaneum, populations in Jordan. Theor Appl Genet 106:397–410
Barry MB, Pham JL, Noyer JL, Billot C, Courtois B, Ahmadi N (2007) Genetic diversity of the two cultivated rice species (O. sativa and O. glaberrima) in Maritime Guinea. Evidence for interspecific recombination. Euphytica 154:127–137
Bioversity International, IRRI and WARDA (2007) Descriptors for wild and cultivated rice (Oryza spp.). Bioversity International, Rome, Italy; International Rice Research Institute, Los Baños, Philippines; WARDA, Africa Rice Center, Cotonou, Benin
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Chang TT (1984) Conservation of rice genetic resources: luxury or necessity? Science 224:251–256
Dawe D, Pandey S, Nelson A (2010) Emerging trends and spatial patterns of rice production. In: Pandey S, Byerlee D, Dawe D, Doberman A, Mohanta S, Rozelle S, Hardy B (eds) Rice in the global economy: strategic research and policy issues for food security. International Rice Research Institute (IRRI), Philippines, pp 15–35
Dong CF, Ding CL, Xu NX, Cheng YH, Shen YX, Gu HR (2013) Double-purpose rice (Oryza sativa L.) variety selection and their morphological traits. Field Crops Res 149:276–282
Edwards K, Johnstone C, Thompson C (1991) A simple and rapid method for the preparation of plant genomic DNA for PCR analysis. Nucleic Acids Res 19:1349
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
FAOSTAT (2013) http://faostat3.fao.org/faostat-gateway/go/to/download/C/CC/E
Garris AJ, Tai TH, Coburn J, Kresovich S, McCouch S (2005) Genetic structure and diversity in Oryza sativa L. Genetics 169:1631–1638
GRiSP (2010) CGIAR Thematic Area 3: susceptible crop productivity increase for global food security. A CGIAR Research Program on Rice-Based Production Systems. Cited Nov 2014
Hubisz MJ, Falush D, Stephens M, Pritchard JK (2009) Inferring weak population structure with the assistance of sample group information. Mol Ecol Res 9:1322–1332
Jones MP, Dingkuhn M, Aluko GK, Semon M (1997) Interspecific Oryza sativa L. × O. glaberrima Steud. Progenies in upland rice improvement. Euphytica 92:237–246
Khush GS (1997) Origin, dispersal, cultivation and variation of rice. Plant Mol Biol 35:25–34
Kumar R, Singh AK (2012) Evaluation of genetic diversity in rice using simple sequence repeats (SSR) markers. Afr J Biotechnol 11:14956–14995
Li YC, Fahima T, Korol AB, Peng J, Roder MS, Kirzhner V, Beiles A, Nevo E (2000) Microsatellite diversity correlated with ecological-edaphic and genetic factors in three microsites of wild emmer wheat in north Israel. Mol Biol Evol 17:851–862
Liu K, Muse SV (2005) PowerMarker: integrated analysis environment for genetic marker data. Bioinformatics 21:2128–2129
Monna L, Kitazawa N, Yoshino R, Suzuki J, Masuda H, Maehara Y, Tanji M, Sato M, Nasu S, Minobe Y (2002) Positional cloning of rice semidwarfing gene, sd-1: Rice “green revolution gene” encodes a mutant enzyme involved in gibberellin synthesis. DNA Res 9:11–17
Perrier X, Jacquemoud-Collet JP (2006) DARwin software http://darwin.cirad.fr/darwin
Perrier X, Flori A, Bonnot F (2003) Data analysis methods. In: Hamon P, Seguin M, Perrier X, Glaszmann JC (eds) Genetic diversity of cultivated tropical plants. Enfield, Science Publishers, Montpellier, pp 43–76
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Qi Y, Zhang H, Zhang D, Wang M, Sun J, Ding L, Wang F, Li Z (2009) Assessing indica–japonica differentiation of improved rice varieties using microsatellite markers. J Genet Genom 36:305–312
Semagn K, Ndjiondjopd MN, Cissoko M (2006) Microsatellites and agronomic traits for assessing genetic relationships among 18 New Rice for Africa (NERICA) varieties. Afr J Biotechnol 5:800–810
Semagn K, Ndjiondjop MN, Lorieux M, Cissoko M, Jones M, McCouch S (2007) Molecular profiling of an interspecific rice population derived from a cross between WAB 56-104 (Oryza sativa) and CG 14 (Oryza glaberrima). Afr J Biotechnol 6:2014–2022
Semon M, Nielsen R, Jones MP, McCouch SR (2005) The population structure of African cultivated rice Oryza glaberrima (Steud.): Evidence for elevated levels of linkage disequilibrium caused by admixture with O. sativa and ecological adaptation. Genetics 169:1639–1647
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Urbana
Sokal RR, Michener CD (1958) A statistical method for evaluating systematic relationships. Univ Kans Sci Bull 38:1409–1438
Sow M, Sido A, Laing M, Ndjiondjop MN (2014) Agro-morphological variability of rice species collected from Niger. Plant Genet Res 12:22–34
Sweeney M, McCouch S (2007) The complex history of the domestication of rice. Ann Bot 100:951–957
Vaughan DA, Lu BR, Tomooka N (2008) The evolving story of rice evolution. Plant Sci 174:394–408
Weir BS (1996) Genetic data analysis II: methods for discrete population genetic data. Sinaur Associates, Sunderland
Willocquet L, Noel M, Sackville HR, Savary S (2012) Susceptibility of rice to sheath blight: an assessment of the diversity of rice germplasm according to genetic groups and morphological traits. Euphytica 183:227–241
Acknowledgement
This research was funded by Africa Rice.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bah, S., Labuschagne, M. & van der Merwe, R. Genetic diversity of improved varieties of intraspecific (O. sativa and O. glaberrima) and interspecific (O. sativa × O. glaberrima) rice. Genet Resour Crop Evol 65, 797–809 (2018). https://doi.org/10.1007/s10722-017-0573-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-017-0573-6