Abstract
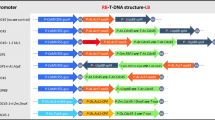
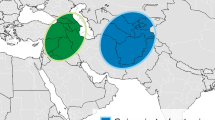
Recently, the less Shattering1 gene (SvLes1), an MYB transcription factor on chromosome V, in Setaria viridis, was reported to control the degree of seed shattering within S. viridis. SvLes1-1 and SvLes1-2 are the wild type (high shattering) allele and the reduced shattering allele, respectively. In addition to these two alleles, the loss-of-function allele through a transposable-element (TE) insertion in exon 2 was found in foxtail millet, a domesticated type of S. viridis, and was designated as SiLes1-TE. This gene is considered to be a domestication gene in foxtail millet. We screened 131 accessions of foxtail millet and found that 16 landraces (12.2%) of foxtail millet do not have the TE, despite expression of the non-shattering phenotype. We sequenced the SvLes1 gene of these 16 accessions and classified them into three alleles, SvLes1-1, SvLes1-2, and a new allele, SvLes1-3. The geographical distribution of these three alleles was different, suggesting that foxtail millet domestication and differentiation are more complex than expected.




Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Bennetzen JL, Schmutz J, Wang H, Percifield R, Hawkins J, Pontaroli AC, Estep M, Feng L, Vaughn JN, Grimwood J, Jenkins J, Barry K, Lindquist E, Hellsten U, Deshpande S, Wang X, Wu X, Mitros T, Triplett J, Yang X, Ye CY, Mauro-Herrera M, Wang L, Li P, Sharma M, Sharma R, Ronald PC, Panaud O, Kellogg EA, Brutnell TP, Doust AN, Tuskan GA, Rokhsar D, Devos KM (2012) Reference genome sequence of the model plant Setaria. Nat Biotechnol 30:555–561
Chen Y, Nie F, Xie S, Zheng Y, Bray T, Dai Q, Wang Y, Xing J, Huang Z, Wang D, He L, Luo F, Wang J, Liu Y, Xiao C (2020) Fast and accurate assembly of Nanopore reads via progressive error correction and adaptive read selection. bioRxiv https://doi.org/10.1101/2020.02.01.930107
Doust AN, Kellogg EA, Devos KM, Bennetzen JL (2009) Foxtail millet: a sequence-driven grass model system. Plant Physiol 149:137–141
Doust AN, Lukens L, Olsen KM, Mauro-Herrera M, Meyer A, Rogers K (2014) Beyond the single gene - do epistasis and gene-by-environment effects influence crop domestication? Proc Natl Acad Sci 111:6178–6183
Fukunaga K (2017) Genetic differentiation and crop evolution of foxtail millet. In: Doust A, Diao X (eds) Genetics and genomics of Setaria (pp 115–131). Springer International Publishing AG
Fukunaga K, Kawase M, Kato K (2002) Structural variation in the Waxy gene and differentiation in foxtail millet [Setaria italica (L.) P. Beauv.]: implications for multiple origins of the waxy phenotype. Mol Genet Genomics 268:214–222
Fukunaga K, Nur MZ, Inoue T, Taketa S, Ichitani K (2020) Phylogenetic analysis of the Si7PPO gene in foxtail millet, Setaria italica, provides further evidence for multiple origins of negative phenol color reaction phenotype. Genes Genet Syst 95:191–198
Hachiken T, Sato K, Hasegawa T, Ichitani K, Kawase M, Fukunaga K (2013) Geographical distribution of the Waxy gene SNPs and Indels in foxtail millet, Setaria italica (L.) P. Beauv. Genet Resour Crop Evol 60:1559–1570
Hirano R, Naito K, Fukunaga K, Watanabe KN, Ohsawa R, Kawase M (2011) Genetic structure of landraces in foxtail millet (Setaria italica (L.) P. Beauv.) revealed with transposon display and interpretation to crop evolution of foxtail millet. Genome 54:498–506
Huang X, Kurata N, Wei X, Wang ZX, Wang A, Zhao Q, Zhao Y, Liu K, Lu H, Li W, Guo Y, Lu Y, Zhou C, Fan D, Weng Q, Zhu C, Huang T, Zhang L, Wang Y, Feng L, Furuumi H, Kubo T, Miyabayashi T, Yuan X, Xu Q, Dong G, Zhan Q, Li C, Fujiyama A, Toyoda A, Lu T, Feng Q, Qian Q, Li J, Han B (2012) A map of rice genome variation reveals the origin of cultivated rice. Nature 490:497–501
Inoue T, Yuo T, Ohta T, Hitomi E, Ichitani K, Kawase M, Taketa S, Fukunaga K (2015) Multiple origins of the phenol reaction negative phenotype in foxtail millet, Setaria italica (L.) P. Beauv., were caused by independent loss-of-function mutations of the polyphenol oxidase (Si7PPO) gene during domestication. Mol Genet Genomics 290:1563–1574
Jia G, Huang XZ, H, Zhao Y, Zhao Q, Li W, Chai Y, Yang L, Liu K, Lu H, Zhu C, Lu Y, Zhou C, Fan D, Weng Q, Guo Y, Huang T, Zhang L, Lu T, Feng Q, Hao H, Liu H, Lu P, Zhang N, Li Y, Guo E, Wang S, Wang S, Liu J, Zhang W, Chen G, Zhang B, Li W, Wang Y, Li H, Zhao B, Li J, Diao X, Han B, (2013) A haplotype map of genomic variations and genome-wide association studies of agronomic traits in foxtail millet (Setaria italica). Nat Genet 45:957–961
Kawase M, Fukunaga K, Kato K (2005) Diverse origins of waxy foxtail millet crops in East and Southeast Asia mediated by multiple transposable element insertions. Mol Genet Genomics 274:131–140
Kihara H, Kishimoto E (1942) Bastarde zwischen Setaria italica und S. viridis. Bot Mag 20:63–67
Konishi S, Izawa T, Lin SY, Ebana K, Fukuta Y, Sasaki T, Yano M (2006) An SNP caused loss of seed shattering during rice domestication. Science 312:1392–1396
Kundu R, Casey J, Sung W (2019) HyPo: super fast & accurate polisher for long read genome assemblies. bioRxiv. https://doi.org/10.1101/2019.12.19.882506
Li P, Brutnell TP (2011) Setaria viridis and Setaria italica, model genetic systems for the Panicoid grasses. J Exp Bot 62:3031–3037
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760
Li HW, Li CH, Pao WK (1945) Cytological and genetical studies of the interspecific cross of the cultivated foxtail millet, Setaria italica (L.) Beauv., and the green foxtail millet S. viridis. J Am Soc Agron 37:32–54
Li C, Zhou A, Sang T (2006) Rice domestication by reducing shattering. Science 311:1936–1939
Lin Z, Li X, Shannon LM, Yeh CT, Wang ML, Bai G, Peng Z, Li J, Trick HN, Clemente TE, Doebley J, Schnable PS, Tuinstra MR, Tesso TT, White F, Yu J (2012) Parallel domestication of the Shattering1 genes in cereals. Nat Genet 44:720–724
Mamidi S, Healey A, Huang P, Grimwood J, Jenkins J, Barry K, Sreedasyam A, Shu S, Lovell JT, Feldman M, Wu J, Yu Y, Chen C, Johnson J, Sakakibara H, Kiba T, Sakurai T, Tavares R, Nusinow DA, Baxter I, Schmutz J, Brutnell TP, Kellogg EA (2020) A genome resource for green millet Setaria viridis enables discovery of agronomically valuable loci. Nat Biotechnol 38:1203–1210
Odonkor S, Choi S, Chakraborty D, Martinez-Bello L, Wang X, Bahri BA, Tenaillon MI, Panaud O, Devos K (2018) QTL mapping combined with comparative analyses identified candidate genesfor reduced shattering in Setaria italica. Front Plant Sci 19:918
Onishi K, Takagi K, Kontani M, Tanaka T, Sano Y (2007) Different patterns of genealogical relationships found in the two major QTLs causing reduction of seed shattering during rice domestication. Genome 50:757–766
Pourkheirandish M, Hensel G, Kilian B, Senthil N, Chen G, Sameri M, Azhaguvel P, Sakuma S, Dhanagond S, Sharma R, Mascher M, Himmelbach A, Gottwald S, Nair SK, Tagiri A, Yukuhiro F, Nagamura Y, Kanamori H, Matsumoto T, Willcox G, Middleton CP, Wicker T, Walther A, Waugh R, Fincher GB, Stein N, Kumlehn J, Sato K, Komatsuda T (2015) Evolution of the grain dispersal system in barley. Cell 162:527–539
Thielen PM, Pendleton AL, Player RA, Bowden KV, Lawton TJ, Wisecaver JH (2020) Reference genome for the highly transformable Setaria viridis ME034V. G3 10:3467–3478
van’t Hof AE, Campagne P, Rigden DJ, Yung CJ, Lingley J, Quail MA, Hall N, Darby AC, Saccheri IJ, (2016) The industrial melanism mutation in British peppered moths is a transposable element. Nature 534:102–105
Vaser R, Sović I, Nagarajan N, Šikić M (2017) Fast and accurate de novo genome assembly from long uncorrected reads. Genome Res 27:737–746
Zhang G, Liu X, Quan Z, Cheng S, Xu X, Pan S, Xie M, Zeng P, Yue Z, Wang W, Tao Y, Bian C, Han C, Xia QP, X, Cao R, Yang X, Zhan D, Hu, J Zhang Y, Li H, Li H, Li N, Wang J, Wang, C, Wang R, Guo T, Cai Y, Liu C, Xiang, H, Shi Q, Huang P, Chen Q, Li Y, Wang, J, Zhao, Z, Wang J, (2012) Genome sequence of foxtail millet (Setaria italica) provides insights into grass evolution and biofuel potential. Nat Biotechnol 30:549–554
Acknowledgements
We are grateful to the NARO Genebank, Japan, the USDA genebank, and Iwate Agricultural Research Center for providing plant materials. This work was supported by JSPS KAKENHI Grant Number 20K06098.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
All the authors inform that the manuscript have not been submitted to other journals and not have been published elsewhere in any form. If there is suspicion of misbehavior or alleged fraud, the Journal and/or Publisher can carry out an investigation following COPE guidelines.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10722_2021_1165_MOESM1_ESM.docx
Supplemental Figure 1. Sequence of SvLes1 gene of A10.1 and primers used for this study. Primer directions are indicated by arrows. (DOCX 41 kb)
10722_2021_1165_MOESM2_ESM.docx
Supplemental Figure 2. Alignment of the SiLes-TE allele among five accessions, Yugu1, JP71640, JP3913, Otsuchi10, and Nisatai-zairai. Sequences of other four landraces were retrieved from the whole genome sequences determined in this study. Exons were highlighted in light blue. Red arrows and large arrows indicate starts and ends of TE sequences and LTRs, respectively. Red arrows indicate primers used for genotyping and their directions. *Nisatai-zairai (DOCX 80 kb)
Rights and permissions
About this article
Cite this article
Fukunaga, K., Matsuyama, S., Abe, A. et al. Insertion of a transposable element in Less Shattering1 (SvLes1) gene is not always involved in foxtail millet (Setaria italica) domestication. Genet Resour Crop Evol 68, 2923–2930 (2021). https://doi.org/10.1007/s10722-021-01165-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-021-01165-w