Abstract
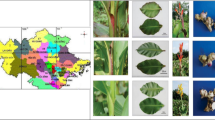
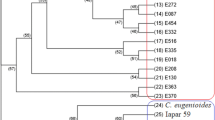
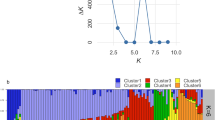
Curcuma alismatifolia is a showy ornamental of the genus Curcuma. Thirty-two C. alismatifolia accessions with high morphologic diversity were collected in this study. A total of 139 alleles were detected by 27 SSR markers. The average values of Shannon's Information index (I) and polymorphism information content were 1.267 and 0.667, respectively. Population structure about the C. alismatifolia accessions was first reported, and the 32 C. alismatifolia accessions were divided into two groups. The genetic differentiation (Fst) between these two groups was 0.127. Phylogenetic analysis showed that two clusters of the whole accessions in the study were related to pedicel length, inflorescence length and yield. The results suggested a high level of genetic diversity among these accessions. Taken together, this study provides a reference for C. alismatifolia industry and a new insight into the genetic diversity and population structure, as well as promotes the breeding program of C. alismatifolia.





Similar content being viewed by others
References
Boonsrangsom T (2020) Genetic diversity of ‘Wan Chak Motluk’ (Curcuma comosa Roxb.) in Thailand using morphological characteristics and random amplification of polymorphic DNA (RAPD) markers. S Afr J Bot 130:224–230. https://doi.org/10.1016/j.sajb.2020.01.005
Cheng Y, Guo W, Yi H, Pang X, Deng X (2003) An efficient protocol for genomic DNA extraction from Citrus species. Plant Mol Biol Report 21:177–178. https://doi.org/10.1007/BF02774246
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resourc 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Favero BT, Dole J, Lima G (2017) Curcuma alismatifolia vase life. Ornam Hortic 23:101–106. https://doi.org/10.14295/oh.v23i1.989
Govindaraj M, Vetriventhan M, Srinivasan M (2015) Importance of genetic diversity assessment in crop plants and its recent advances: an overview of its analytical perspectives. Genet Res Int. https://doi.org/10.1155/2015/431487
Gupta AK, Mishra R, Lal RK (2015) Genetic resources, diversity, characterization and utilization of agronomical traits in turmeric (Curcuma longa L.). Ind Crops Prod 77:708–712. https://doi.org/10.1016/j.indcrop.2015.09.030
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129. https://doi.org/10.1093/bioinformatics/bti282
Mao L, Liu J, Ding H, Xu X, Tian D (2018) Microsatellite characterization analysis and primers design of the whole transcriptome of Curcuma alismatifolia. Mol Plant Breed 16:7407–7414
Mao L, Zou Q, Liu J, Ding H, Tian D (2020a) Genetic diversity and genetic relationships among Curcuma accessions based on SRAP and ISSR analysis. Ecol Genet Genomics 15:100053. https://doi.org/10.1016/j.egg.2020.100053
Mao L, Liu J, Ding H, Zou Q, Tian D (2020b) Development of SSR markers and their application to genetic diversity analysis of Curcuma alismatifolia varieties. Bot Sci 99:124–131. https://doi.org/10.17129/botsci.2619
Paisooksantivatana Y, Kako S, Seko H (2001a) Genetic diversity of Curcuma alismatifolia Gagnep. (Zingiberaceae) in Thailand as revealed by allozyme polymorphism. Genet Resour Crop Evol 48:459–465. https://doi.org/10.1023/A:1012078728003
Paisooksantivatana Y, Kako S, Seko H (2001b) Isozyme polymorphism in Curcuma alismatifolia Gagnep. (Zingiberaceae) populations from Thailand. Sci Hortic 88:299–307. https://doi.org/10.1016/S0304-4238(00)00218-1
Paw M, Munda S, Borah A, Pandey SK, Lal M (2020) Estimation of variability, genetic divergence, correlation studies of Curcuma caesia Roxb. J Appl Res Med Aromatic Plants 17:100251. https://doi.org/10.1016/j.jarmap.2020.100251
Peakall ROD, Smouse PE (2006) Genalex 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Sasikumar B (2005) Genetic resources of Curcuma: diversity, characterization and utilization. Plant Genet Resour 3:230–251. https://doi.org/10.1079/PGR200574
Sigrist MS, Pinheiro JB, Azevedo-Filho JA, Colombo CA, Bajay MM, Lima PF, Camilo FR, Sandhu S, Souza AP, Zucchi MI (2010) Development and characterization of microsatellite markers for turmeric (Curcuma longa). Plant Breed 129:570–573. https://doi.org/10.1111/j.1439-0523.2009.01720.x
Sihanat A, Theanphong O, Rungsihirunrat K (2020) Assessment of phylogenetic relationship among twenty Curcuma species in Thailand using amplified fragment length polymorphism marker. J Adv Pharm Technol Res 11:134–141. https://doi.org/10.4103/japtr.JAPTR_24_20
Singh S, Panda MK, Nayak S (2012) Evaluation of genetic diversity in turmeric (Curcuma longa L.) using RAPD and ISSR markers. Ind Crops Prod 37:284–291. https://doi.org/10.1016/j.indcrop.2011.12.022
Syamkumar S, Sasikumar B (2007) Molecular marker based genetic diversity analysis of Curcuma species from India. Sci Hortic 112:235–241. https://doi.org/10.1016/j.scienta.2006.12.021
Taheri S, Abdullah T, Abdullah N, Ahmad Z (2012) Genetic relationships among five varieties of Curcuma alismatifolia (Zingiberaceae) based on ISSR markers. Genet Mol Res 11:3069–3076. https://doi.org/10.4238/2012.August.31.4
Taheri S, Abdullah T, Abdullah N, Ahmad Z, Karimi E, Shabanimofrad M (2014) Assessing the genetic relationships of Curcuma alismatifolia varieties using simple sequence repeat markers. Genet Mol Res 13:7339–7346. https://doi.org/10.4238/2014.September.5.12
Taheri S, Abdullah TL, Rafii MY, Harikrishna JA, Werbrouck SPO, Teo CH, Sahebi M, Azizi P (2019) De novo assembly of transcriptomes, mining, and development of novel EST-SSR markers in Curcuma alismatifolia (Zingiberaceae family) through Illumina sequencing. Sci Rep 9:3047. https://doi.org/10.1038/s41598-019-39944-2
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Thohirah LA, Flora C, Kamalakshi N (2010) Breaking bud dormancy and different shade levels for production of pot and cut Cucurma alismatifolia. Am J Agric Biol Sci 5:385–383. https://doi.org/10.3844/ajabssp.2010.385.388
Ullah H, Rabbani MA, Shinwari ZK (2011) Assessment of genetic diversity of indigenous turmeric (Curcuma longa L.) germplasm from Pakistan using RAPD markers. J Med Plant Res 5:823–830. https://doi.org/10.5897/JMPR.9000277
Wright JS (1978) Evolution and the genetics of natural populations, Volume 4. Variability within and among natural populations. University of Chicago Press, Chicago
Yeh FC, Boyle TJB (1997) Population genetic analysis of codominant and dominant markers and quantitative traits. Belgian J Bot 129:157–163
Yu H, Cheng Y, Xu Q, Deng X (2018) SSR polymorphism analysis in citrus. Bio-Protocol. https://doi.org/10.21769/BioProtoc.1010214
Acknowledgements
The authors appreciate the assistance of Zhangzhou Jinluan Horticulture Co. LTD, Zhangzhou city, Fujian province, China.
Funding
This project was supported by Natural Science Foundation of Fujian Province (2020J05168), Educational Research Project for Young and Middle-aged Teachers in Fujian Province (JAT190389), Regional Development Project of Science and Technology Department of Fujian Province (2020N3010) and the Principal Foundation of Minnan Normal University (4206/L21816, 4206/L21715).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Plant materials were collected by JS Lin and YY Li. Experimentation, and data collection were performed by YL Xiao, SJ Chen and YH Li. The analysis of the data was implemented by LJ Ke and QL Tian. The manuscript was written by HW Yu and LM Lu.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yu, HW., Ke, LJ., Xiao, YL. et al. Genetic diversity and population structure of Curcuma alismatifolia Gagnep. accessions revealed by SSR markers. Genet Resour Crop Evol 69, 1661–1669 (2022). https://doi.org/10.1007/s10722-021-01329-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-021-01329-8