Abstract
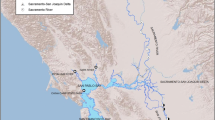
Burbot (Lota lota) occur in the Wind River Basin in central Wyoming, USA, at the southwestern extreme of the species’ native range in North America. The most stable and successful of these populations occur in six glacially carved mountain lakes on three different tributary streams and one large main stem impoundment (Boysen Reservoir) downstream from the tributary populations. Burbot are rarely found in connecting streams and rivers, which are relatively small and high gradient, with a variety of potential barriers to upstream movement of fish. We used high-throughput genomic sequence data for 11,197 SNPs to characterize the genetic diversity, population structure, and connectivity among burbot populations on the Wind River system. Fish from Boysen Reservoir and lower basin tributary populations were genetically differentiated from those in the upper basin tributary populations. In addition, fish within the same tributary streams fell within the same genetic clusters, suggesting there is movement of fish between lakes on the same tributaries but that populations within each tributary system are isolated and genetically distinct from other populations. Observed genetic differentiation corresponded to natural and anthropogenic barriers, highlighting the importance of barriers to fish population connectivity and gene flow in human-altered linked lake-stream habitats.





Similar content being viewed by others
References
Bergstedt, L. & E. P. Bergersen, 1992. Impacts of Water Management on the Fishery Resources of the Wind River on the Wind River Indian Reservation. Annual Report by CSU Cooperative Fish & Wildlife Research Unit, Fort Collins, CO.
Burger, C. V., W. J. Spearman & M. A. Cronin, 1997. Genetic differentiation of sockeye salmon subpopulations from a geologically young Alaskan lake system. Transactions of the American Fisheries Society 126: 926–938.
Bjorn, E. E., 1940. Preliminary observations and experimental study of the ling, Lota maculosa (LeSueur), in Wyoming. Transactions of the American Fisheries Society 69: 192–196.
Carlsson, J., K. H. Olsen, J. Nilsson, O. Overli & O. B. Stabell, 1999. Microsatellites reveal fine-scale genetic structure in stream-living brown trout. Journal of Fish Biology 55: 1290–1303.
Dillen, A., J. Coeck, & D. Monnier, 2008. Habitat Use and Seasonal Migrations of Burbot in Lowland Rivers in North France. In Paragamian, V. L. & D. H. Bennett (eds), Burbot: Biology, Management, and Culture. American Fisheries Society, Symposium 59, Bethesda, MD: 29–42.
Dunham, J. B. & B. E. Rieman, 1999. Metapopulation structure of bull trout: influences of physical, biotic, and geometrical landscape characteristics. Ecological Applications 92: 642–655.
Dunnigan, J. L., & L. S. Cameron, 2008. Home Range and Movement Patterns of Burbot in Koocanusa Reservoir, Montana. In Paragamian, V. L. & D. H. Bennett (eds), Burbot: Biology, Management, and Culture. American Fisheries Society, Symposium 59, Bethesda, MD: 43–54.
Elshire, R. J., J. C. Glaubitz, Q. Sun, J. A. Poland, K. Kawamoto, E. S. Buckler & S. E. Mitchell, 2011. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS One 6: e19379.
Evenson, M. J., 1993. Seasonal movements of radio-implanted burbot in the Tanana River drainage. Alaska Department of Fish and Game, Division of Sport Fish 93–97: 1–25.
Falush, D., M. Stephens & J. K. Pritchard, 2003. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164: 1567–1587.
Fitzpatrick, S. W., J. C. Gerberich, J. A. Kronenberger, L. M. Angeloni & W. C. Funk, 2015. Locally adapted traits maintained in the face of high gene flow. Ecology Letters 18: 37–47.
Fontaine, P. M., J. J. Dodson, L. Bernatchez & A. Slettan, 1997. A genetic test of metapopulation structure in Atlantic salmon (Salmo salar) using microsatellites. Canadian Journal of Fisheries and Aquatic Sciences 54: 2434–2442.
Gardunio, E. I., 2014. Jumping and Swimming Performance of Burbot and White Suckers: Implications for Barrier Design. Master’s thesis, Colorado State University, Fort Collins, CO.
Gompert, Z., L. K. Lucas, C. A. Buerkle, M. L. Forister, J. A. Fordyce & C. C. Nice, 2014. Admixture and the organization of genetic diversity in a butterfly species complex revealed through common and rare genetic variants. Molecular Ecology 23: 455–4573.
Hagen, G. O., 1952. Ling Hatching Experiment, Cokeville, Wyoming. Cheyenne, WY: Report to the Wyoming Game and Fish Commission
Hawkins, D., B. Adams, S. Roth & D. Skates, 2011. Genetic characterization of Burbot (Lota lota maculosa) from three lakes in the Wind River watershed, WY. Project Report, U.S. Fish and Wildlife Service, Lander, WY.
Hohenlohe, P. A., M. D. Day, S. J. Amish, M. R. Miller, N. Kamps-Hughes, M. C. Boyer, C. C. Muhlfeld, F. W. Allendorf, E. A. Johnson & G. Luikart, 2013. Genomic patterns of introgression in rainbow and westslope cutthroat trout illuminated by overlapping paired-end rad sequencing. Molecular Ecology 22: 3002–3013.
Hubert, W., D. Dufek, J. Deromedi, K. Johnson, S. Roth, & D. Skates, 2008. Burbot in the Wind River Drainage of Wyoming: Knowledge of Stocks and Management Issues. In Paragamian V. L. & D. H. Bennett (eds), Burbot: Biology, Management, and Culture. American Fisheries Society, Symposium 59, Bethesda, MD: 187–200.
Hudson, R. R., M. Slatkin & W. P. Maddison, 1992. Estimation of levels of gene flow from DNA sequence data. Genetics 132: 583–589.
Krueger, K. L., 1996. Assessment of Lentic Sport Fisheries for Burbot and Sauger in the Bighorn/Wind River Drainage, Wyoming. Master’s thesis, Universtiy of Wyoming, Laramie, WY.
Li, H., 2011. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27: 2987–2993.
Li, H. & R. Durbin, 2009. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25: 1754–1760.
Li, H., B. Handsaker, A. Wysoker, T. Fennell, J. Ruan, N. Homer, G. Marth, G. Abecasis & R. Durbin, 2009. The sequence alignment/map format and SAMtools. Bioinformatics 25: 2078–2079.
Mandeville, E. G., T. L. Parchman, D. B. McDonald & C. A. Buerkle, 2015. Highly variable reproductive isolation among pairs of Catostomus species. Molecular Ecology 24: 1856–1872.
McDermid, J. L., S. Nienhuis, M. Al-Shamlih, T. J. Haxton & C. C. Wilson, 2014. Evaluating the genetic consequences of river fragmentation in lake sturgeon (Acipenser fulvescens Rafinesque, 1817) populations. Applied Icthyology 30: 1514–1523.
McPhail, J. D., & V. L. Paragamian, 2000. Burbot Biology and Life History. In Paragamian, V. L. & D. W. Willis (eds), Burbot: Biology, Ecology, and Management American Fisheries Society, Fisheries Management Section, Publication Number 1, Bethesda, MD: 11–23
Paragamian, V. L., 2000. The Effects of Variable Flows on Burbot Spawning Migrations in the Kootenai River, Idaho, USA, and British Columbia, Canada. In Paragamian, V. L. & D. W. Willis (eds), Burbot: Biology, Ecology, and Management American Fisheries Society, Fisheries Management Section, Publication Number 1, Bethesda, MD: 111–123
Paragamian V. L. & D. H. Bennett (eds), 2008. Burbot: Biology, Management, and Culture. American Fisheries Society, Symposium 59, Bethesda, MD.
Paragamian, V. L. & M. A. Stapanian, 2011. Preface. Journal of Applied Ichthyology 27: 3.
Paragamian, V. L., & V. D. Wakkinen, 2008. Seasonal Movement of Burbot in Relation to Temperature and Discharge in the Kootenai River, Idaho, USA and British Columbia, Canada. In Paragamian V. L. & D. H. Bennett (eds), Burbot: Biology, Management, and Culture. American Fisheries Society, Symposium 59, Bethesda, MD: 55–77.
Paragamian, V. L., M. Powell & J. Foler, 1999. Mitochondrial DNA analysis of burbot in the Kootenai River basin of British Columbia, Montana, and Idaho. Transactions of the American Fisheries Society 128: 854–886.
Paragamian, V. L., R. Hardy & B. Gunderman, 2005. Effects of regulated discharge on burbot migration. Journal of Fish Biology 66: 1199–1213.
Parchman, T. L., Z. Gompert, J. Mudge, F. Schilkey, C. W. Benkman & C. A. Buerkle, 2012. Genome-wide association genetics of an adaptive trait in lodgepole pine. Molecular ecology 21: 2991–3005.
Pickrell, J. K. & J. K. Pritchard, 2012. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS genetics 8: e1002967.
Pritchard, J. K., M. Stephens & P. Donnelly, 2000. Inference of population structure using multilocus genotype data. Genetics 155: 945–959.
Roach, S. M., & M. J. Evenson, 1993. A Geometric Approach to Estimating and Predicting the Fecundity of Tanana River Burbot. Alaska Department of Fish and Game. Juneau, AK. Fisheries Data Series 93–98.
Rogers, K. B., 2007. A Suggested Protocol for Collecting Cutthroat Trout Tissues for Subsequent Genetic Analysis. Colorado Divison of Wildlife, Fort Collins, CO.
Spence, C., & M. Neufeld, 2002. Radio Telemetry Studies of Duncan Reservoir Burbot. Report prepared by the Ministry of Water, Land and Air Protection, for the BC Habitat Conservation Trust Fund and the Bonneville Power Administration, Nelson BC.
Stapanian, M. A., V. L. Paragamian, C. P. Madenjian, J. R. Jackson, J. Lappalainen, M. J. Evenson & M. D. Neufeld, 2010. Worldwide status of burbot and conservation measures. Fish and Fisheries 11: 34–56.
Thorrold, S. R., C. Latkoczy, P. K. Swart & C. M. Jones, 2001. Natal homing in a marine fish metapopulation. Science 291: 297–299.
Wagner, C. E., I. Keller, S. Wittwer, O. M. Selz, S. Mwaiko, L. Greuter, A. Sivasundar & O. Seehausen, 2013. Genome-wide rad sequence data provide unprecedented resolution of species boundaries and relationships in the Lake Victoria cichlid adaptive radiation. Molecular Ecology 22: 787–798.
Williams, F., 1959. Progress Report on Life History Investigations of the Burbot. Wyoming Game and Fish Commission, Daniel.
Woodford, D. J. & A. R. McIntosh, 2010. Evidence of source-sink metapopulations in a vulnerable native galaxid fish driven by introduced trout. Ecological Applications 20: 967–977.
Acknowledgements
We could not have accomplished this research without extensive help in the field and lab. Joe Deromedi, Paul Gerrity, and Kevin Johnson with the Wyoming Game and Fish Department, and Mike Mazur with US Fish and Wildlife collected the majority of our fish for genetic sampling, and were invaluable resources during the planning stages of this project. Sean Lewandoski also collected a number of burbot for us. Alex Buerkle provided direction and advice relating to the genetics sample collection and analyses, and provided extensive computing and lab resources. David Underwood provided figure design support. Carlin Girard, Eric Gardunio, and Mary Kathryn Hooley offered intellectual contributions to data interpretation and discussed initial drafts of this manuscript. Dave McDonald and two anonymous reviewers provided feedback that further improved the manuscript. All fish were treated humanely and anesthetized before all surgical procedures in accordance with University of Wyoming Animal Care and Use Committee protocol #A-3216-01. Funding was provided by the Wyoming Game and Fish Department and the U.S. Geological Survey. Use of trade, product, or firm names does not imply endorsement by the U.S. Government.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: M. Power
This article was originally intended to appear in ‘Ecology, Culture, and Management of Burbot’ guest edited by Martin A. Stapanian & Christopher A. Myrick (published in volume 757 of Hydrobiologia).
Data accessibility Raw sequence data (.fastq files) are available at the National Center for Biotechnology Information (NCBI) SRA: SRX1078958. SNP files and necessary scripts are available at http://datadryad.org; doi:10.5061/dryad.7842r.
Rights and permissions
About this article
Cite this article
Underwood, Z.E., Mandeville, E.G. & Walters, A.W. Population connectivity and genetic structure of burbot (Lota lota) populations in the Wind River Basin, Wyoming. Hydrobiologia 765, 329–342 (2016). https://doi.org/10.1007/s10750-015-2422-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10750-015-2422-y