Abstract
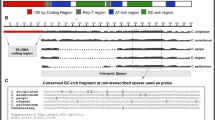
Approximately seven hundred 45S rRNA genes (rDNA) in the Arabidopsis thaliana genome are organised in two 4 Mbp-long arrays of tandem repeats arranged in head-to-tail fashion separated by an intergenic spacer (IGS). These arrays make up 5 % of the A. thaliana genome. IGS are rapidly evolving sequences and frequent rearrangements inside the rDNA loci have generated considerable interspecific and even intra-individual variability which allows to distinguish among otherwise highly conserved rRNA genes. The IGS has not been comprehensively described despite its potential importance in regulation of rDNA transcription and replication. Here we describe the detailed sequence variation in the complete IGS of A. thaliana WT plants and provide the reference/consensus IGS sequence, as well as genomic DNA analysis. We further investigate mutants dysfunctional in chromatin assembly factor-1 (CAF-1) (fas1 and fas2 mutants), which are known to have a reduced number of rDNA copies, and plant lines with restored CAF-1 function (segregated from a fas1xfas2 genetic background) showing major rDNA rearrangements. The systematic rDNA loss in CAF-1 mutants leads to the decreased variability of the IGS and to the occurrence of distinct IGS variants. We present for the first time a comprehensive and representative set of complete IGS sequences, obtained by conventional cloning and by Pacific Biosciences sequencing. Our data expands the knowledge of the A. thaliana IGS sequence arrangement and variability, which has not been available in full and in detail until now. This is also the first study combining IGS sequencing data with RFLP analysis of genomic DNA.





Similar content being viewed by others
References
Abou-Ellail M, Cooke R, Saez-Vasquez J (2011) Variations on a team major and minor variants of Arabidopsis thaliana rDNA genes. Nucleus 2:294–299. doi:10.4161/nucl.2.4.16561
Cavallero S, De Liberato C, Friedrich KG, Di Cave D, Masella V, D’Amelio S, Berrilli F (2015) Genetic heterogeneity and phylogeny of Trichuris spp. from captive non-human primates based on ribosomal DNA sequence data. Infect Genet Evol 34:450–456. doi:10.1016/j.meegid.2015.06.009
Chandrasekhara C, Mohannath G, Blevins T, Pontvianne F, Pikaard CS (2016) Chromosome-specific NOR inactivation explains selective rRNA gene silencing and dosage control in Arabidopsis Gene Dev 30:177–190. doi:10.1101/gad.273755.115
Copenhaver GP, Pikaard CS (1996a) RFLP and physical mapping with an rDNA-specific endonuclease reveals that nucleolus organizer regions of Arabidopsis thaliana adjoin the telomeres on chromosomes 2 and 4. Plant J 9:259–272. doi:10.1046/j.1365-313X.1996.09020259.x
Copenhaver GP, Pikaard CS (1996b) Two-dimensional RFLP analyses reveal megabase-sized clusters of rRNA gene variants in Arabidopsis thaliana, suggesting local spreading of variants as the mode for gene homogenization during concerted evolution. Plant J 9:273–282. doi:10.1046/j.1365-313X.1996.09020273.x
Copenhaver GP, Doelling JH, Gens S, Pikaard CS (1995) Use of RFLPs larger than 100 kbp to map the position and internal organization of the nucleolus organizer region on chromosome 2 in Arabidopsis thaliana. Plant J 7:273–286
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA minipreparation: Version II. Plant Mol Biol Report 1:19–21
Durut N et al (2014) A duplicated NUCLEOLIN gene with antagonistic activity is required for chromatin organization of silent 45S rDNA in Arabidopsis. Plant Cell 26:1330–1344. doi:10.1105/tpc.114.123893
Dvorackova M, Fojtova M, Fajkus J (2015) Chromatin dynamics of plant telomeres and ribosomal genes. Plant J 83:18–37. doi:10.1111/tpj.12822
Earley K et al (2006) Erasure of histone acetylation by Arabidopsis HDA6 mediates large-scale gene silencing in nucleolar dominance. Genes Dev 20:1283–1293. doi:10.1101/gad.1417706
Earley KW et al (2010) Mechanisms of HDA6-mediated rRNA gene silencing: suppression of intergenic Pol II transcription and differential effects on maintenance versus siRNA-directed cytosine methylation. Genes Dev 24:1119–1132. doi:10.1101/gad.1914110
Eickbush TH, Eickbush DG (2007) Finely orchestrated movements: evolution of the ribosomal RNA genes. Genetics 175:477–485. doi:10.1534/genetics.107.071399
Elliott TA, Stage DE, Crease TJ, Eickbush TH (2013) In and out of the rRNA genes: characterization of Pokey elements in the sequenced Daphnia genome. Mobile DNA. doi:10.1186/1759-8753-4-20
Endo M et al (2006) Analysis of Arabidopsis CAF-1 mutants showing enhanced homologous recombination. Plant Cell Physiol 47:S60–S60
Exner V, Taranto P, Schonrock N, Gruissem W, Hennig L (2006) Chromatin assembly factor CAF-1 is required for cellular differentiation during plant development. Development 133:4163–4172 doi:10.1242/dev.02599
Fajkus J, Reich J (1991) Evaluation of restriction endonuclease cleavage of plant nuclear-DNA using contaminating chloroplast DNA. Folia Biol Prague 37:224–226
Fojtova M, Fulneckova J, Fajkus J, Kovarik A (2002) Recovery of tobacco cells from cadmium stress is accompanied by DNA repair and increased telomerase activity. J Exp Bot 53:2151–2158
Garcia S, Kovarik A (2013) Dancing together and separate again: gymnosperms exhibit frequent changes of fundamental 5S and 35S rRNA gene (rDNA) organisation. Heredity 111:23–33. doi:10.1038/hdy.2013.11
Garcia S, Panero JL, Siroky J, Kovarik A (2010) Repeated reunions and splits feature the highly dynamic evolution of 5S and 35S ribosomal RNA genes (rDNA) in the Asteraceae family. BMC Plant Biol 10:176. doi:10.1186/1471-2229-10-176
Geiser C, Mandakova T, Arrigo N, Lysak MA, Parisod C (2016) Repeated whole-genome duplication, karyotype reshuffling, and biased retention of stress-responding genes in buckler mustard. Plant Cell 28:17–27. doi:10.1105/tpc.15.00791
Gruendler P, Unfried I, Pointner R, Schweizer D (1989) Nucleotide-Sequence of the 25S–18S ribosomal gene spacer from Arabidopsis thaliana. Nucleic Acids Res 17:6395–6396. doi:10.1093/nar/17.15.6395
Gruendler P, Unfried I, Pascher K, Schweizer D (1991) rDNA intergenic region from Arabidopsis thaliana. Structural analysis, intraspecific variation and functional implications. J Mol Biol 221:1209–1222
Grummt I, Pikaard CS (2003) Epigenetic silencing of RNA polymerase I transcription. Nat Rev Mol Cell Bio 4:641–649. doi:10.1038/nrm1171
Grummt I, Kuhn A, Bartsch I, Rosenbauer H (1986) A transcription terminator located upstream of the mouse rDNA initiation site affects rRNA synthesis. Cell 47:901–911
Han EH, Cho K, Goo Y, Kim M, Shin YW, Kim YH, Lee SW (2016) Development of molecular markers, based on chloroplast and ribosomal DNA regions, to discriminate three popular medicinal plant species, Cynanchum wilfordii, Cynanchum auriculatum, and Polygonum multiflorum. Mol Biol Reports. doi:10.1007/s11033-016-3959-1
Kaya H, Shibahara KI, Taoka KI, Iwabuchi M, Stillman B, Araki T (2001) FASCIATA genes for chromatin assembly factor-1 in arabidopsis maintain the cellular organization of apical meristems. Cell 104:131–142
Kelly LJ et al (2015) Analysis of the giant genomes of Fritillaria (Liliaceae) indicates that a lack of DNA removal characterizes extreme expansions in genome size. New Phytol 208:596–607. doi:10.1111/nph.13471
Kirik A, Pecinka A, Wendeler E, Reiss B (2006) The chromatin assembly factor subunit FASCIATA1 is involved in homologous recombination in plants. Plant Cell 18:2431–2442. doi:10.1105/tpc.106.045088
Kobayashi T, Horiuchi T, Tongaonkar P, Vu L, Nomura M (2004) SIR2 regulates recombination between different rDNA repeats, but not recombination within individual rRNA genes in yeast. Cell 117:441–453
Konstantinova P, Yli-Mattila T (2004) IGS-RFLP analysis and development of molecular markers for identification of Fusarium poae, Fusarium langsethiae, Fusarium sporotrichioides and Fusarium kyushuense. Int J Food Microbiol 95:321–331. doi:10.1016/j.ijfoodmicro.2003.12.010
Lauro FM, Chastain RA, Blankenship LE, Yayanos AA, Bartlett DH (2007) The unique 16S rRNA genes of piezophiles reflect both phylogeny and adaptation. Appl Environ Microbiol 73:838–845. doi:10.1128/AEM.01726-06
Leyser HMO, Furner IJ (1992) Characterization of 3 shoot apical meristem mutants of Arabidopsis thaliana. Development 116:397–403
Lin RQ, Shu L, Zhao GH, Cheng T, Zou SS, Zhang Y, Weng YB (2014) Characterization of the intergenic spacer rDNAs of two pig nodule worms, Oesophagostomum dentatum and O. quadrispinulatum. Sci World J 2014:147963. doi:10.1155/2014/147963
Long Q et al (2013) Massive genomic variation and strong selection in Arabidopsis thaliana lines from Sweden. Nat Genet 45:884–890. doi:10.1038/ng.2678
Mandakova T, Lysak MA (2008) Chromosomal phylogeny and karyotype evolution in x = 7 crucifer species (Brassicaceae). Plant Cell 20:2559–2570. doi:10.1105/tpc.108.062166
Marcilla A et al (2001) The ITS-2 of the nuclear rDNA as a molecular marker for populations, species, and phylogenetic relationships in Triatominae (Hemiptera: Reduviidae), vectors of Chagas disease. Mol Phylogenet Evol 18:136–142 doi:10.1006/mpev.2000.0864
Mozgova I, Mokros P, Fajkus J (2010) Dysfunction of chromatin assembly factor 1 induces shortening of telomeres and loss of 45S rDNA in Arabidopsis thaliana. Plant Cell 22:2768–2780. doi:10.1105/tpc.110.076182
Muchova V, Amiard S, Mozgova I, Dvorackova M, Gallego ME, White C, Fajkus J (2015) Homology-dependent repair is involved in 45S rDNA loss in plant CAF-1 mutants. Plant J 81:198–209. doi:10.1111/tpj.12718
Pavlištová V, Dvořáčková M, Jež M, Mozgová I, Mokroš P, Fajkus J (2016) Phenotypic reversion in mutants of by reintroduction of genes: variable recovery of telomeres with major spatial rearrangements and transcriptional reprogramming of 45S rDNA genes. Plant J. doi:10.1111/tpj.13257
Pikaard CS et al (1990) Enhancers for RNA polymerase I in mouse ribosomal DNA. Mol Cell Biol 10:4816–4825
Pontes O et al (2003) Natural variation in nucleolar dominance reveals the relationship between nucleolus organizer chromatin topology and rRNA gene transcription in Arabidopsis. P Natl Acad Sci USA 100:11418–11423. doi:10.1073/pnas.1932522100
Pontvianne F et al (2010) Nucleolin is required for DNA methylation state and the expression of rRNA gene variants in Arabidopsis thaliana. PLoS Genet 6:e1001225. doi:10.1371/journal.pgen.1001225
Pontvianne F et al (2013) Subnuclear partitioning of rRNA genes between the nucleolus and nucleoplasm reflects alternative epiallelic states. Gene Dev 27:1545–1550. doi:10.1101/gad.221648.113
Preuss S, Pikaard CS (2007) rRNA gene silencing and nucleolar dominance: insights into a chromosome-scale epigenetic on/off switch. Bba Gene Struct Expr 1769:383–392 doi:10.1016/j.bbaexp.2007.02.005
Pruitt RE, Meyerowitz EM (1986) Characterization of the genome of Arabidopsis thaliana. J Mol Biol 187:169–183. doi:10.1016/0022-2836(86)90226-3
Ramirez-Parra E, Gutierrez C (2007) E2F regulates FASCIATA1, a chromatin assembly gene whose loss switches on the endocycle and activates gene expression by changing the epigenetic status. Plant Physiol 144:105–120. doi:10.1104/pp.106.094979
Reinholz E (1966) Radiation induced mutants showing changed inflorescence characteristics Arabid Inf Serv 3:19–20
Schonrock N, Exner V, Probst A, Gruissem W, Hennig L (2006) Functional genomic analysis of CAF-1 mutants in Arabidopsis thaliana. J Biol Chem 281:9560–9568. doi:10.1074/jbc.M513426200
Schubert I, Wobus U (1985) In situ hybridisation confirms jumping nucleolus organizing regions in Allium. Chromosoma 92:143–148. doi:10.1007/BF00328466
Smith S, Stillman B (1989) Purification and characterization of CAF-I, a human cell factor required for chromatin assembly during DNA replication in vitro. Cell 58:15–25
Symonova R et al (2013) Genome differentiation in a species pair of coregonine fishes: an extremely rapid speciation driven by stress-activated retrotransposons mediating extensive ribosomal DNA multiplications. BMC Evol Biol 13:42. doi:10.1186/1471-2148-13-42
Takeda S et al (2004) BRU1, a novel link between responses to DNA damage and epigenetic gene silencing in Arabidopsis. Gene Dev 18:782–793. doi:10.1101/gad.295404
Weider LJ, Elser JJ, Crease TJ, Mateos M, Cotner JB, Markow TA (2005) The functional significance of ribosomal (r)DNA variation: Impacts on the evolutionary ecology of organisms. Annu Rev Ecol Evol S 36:219–242. doi:10.1146/annurev.ecolsys.36.102003.152620
Acknowledgments
We thank Veronika Pavlištová for providing us with the revertant A. thaliana lines. Access to computing and storage facilities owned by parties and projects contributing to the National Grid Infrastructure MetaCentrum, provided under the programme “Projects of Large Research, Development, and Innovations Infrastructures” (CESNET LM2015042) is greatly appreciated. This work was supported by GACR 13-11563P, 16-04166Y and by the Ministry of Education, Youth and Sports of the Czech Republic-projects KONTAKT II no. LH15189 and CEITEC 2020 (LQ1601). Work in the laboratory of C.G. is supported by Grants BFU2012-34821 and BFU2015-68396-R.
Author contributions
J.F. and M.D. designed the study, K.H., M.D., I.M. and L.V. performed experiments, all authors performed analysis and interpretation of results, K.H., M.D. and J.F. wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Havlová, K., Dvořáčková, M., Peiro, R. et al. Variation of 45S rDNA intergenic spacers in Arabidopsis thaliana . Plant Mol Biol 92, 457–471 (2016). https://doi.org/10.1007/s11103-016-0524-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-016-0524-1