Abstract
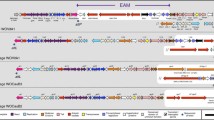
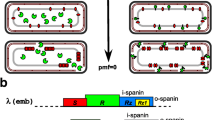
Genomes of the obligate intracellular alpha proteobacterium Wolbachia pipientis often encode prophage-like regions, and in a few cases, purified particles have been recovered. Because the structure of a conserved WO phage genome has been difficult to establish, we examined paired terminase and portal genes in Wolbachia phages and prophages, relative to those encoded by the gene transfer agent RcGTA from the free-living alpha proteobacterium Rhodobacter capsulatus. Terminase and portal proteins from Wolbachia have higher similarity to orthologs encoded by RcGTA than to orthologs encoded by bacteriophage lambda. In lambdoid phages, these proteins play key roles in assembly of mature phage particles, while in less well-studied gene transfer agents, terminase and portal proteins package random fragments of bacterial DNA, which could confound elucidation of WO phage genomes. In WO phages and prophages, terminase genes followed by a short gpW gene may be separated from the downstream portal gene by open-reading frames encoding a GH_25 hydrolase/muramidase, a PD-(D/E)XK nuclease, a hypothetical protein and/or a RelE/ParE toxin-antitoxin module. These aspects of gene organization, coupled with evidence for a low, non-inducible yield of WO phages, and the small size of WO phage particles described in the literature raise the possibility that Wolbachia prophage regions participate in processes that extend beyond conventional bacteriophage lysogeny and lytic replication. These intervening genes, and their possible relation to functions associated with GTAs, may contribute to variability among WO phage genomes recovered from physical particles and impact the ability of WO phages to act as transducing agents.














Similar content being viewed by others
Data availability
There are no large-scale data.
References
Landmann F (2019) The Wolbachia endosymbionts. Microbiol Spectr. https://doi.org/10.1128/microbiolspec.BAI-0018-2019
Klasson L, Westberg J, Sapountzis P, Näslund K, Lutnaes Y, Darby AC, Veneti Z, Chen L, Braig HR, Garrett R, Bourtzis K, Andersson SG (2009) The mosaic genome structure of the Wolbachia wRi strain infecting Drosophila simulans. Proc Natl Acad Sci USA 106(14):5725–5730. https://doi.org/10.1073/pnas.0810753106
Kent BN, Bordenstein SR (2010) Phage WO of Wolbachia: lambda of the endosymbiont world. Trends Microbiol 18(4):173–181. https://doi.org/10.1016/j.tim.2009.12.011
Kent BN, Funkhouser LJ, Setia S, Bordenstein SR (2011) Evolutionary genetics of a temperate bacteriophage in an obligate intracellular bacteria (Wolbachia). PLoS ONE 6(9):e24984
Bordenstein SR, Bordenstein SR (2016) Eukaryotic association module in phage WO genomes from Wolbachia. Nat Commun 7:13155. https://doi.org/10.1038/ncomms13155
Werren JH, Baldo L, Clark ME (2008) Wolbachia: master manipulators of invertebrate biology. Nat Rev Microbiol 6:741–751. https://doi.org/10.1038/nrmicro1969
Fraser JE, De Bruyne JT, Iturbe-Ormaetxe I, Stepnell J, Burns RL, Flores HA, O’Neill SL (2017) Novel Wolbachia-transinfected Aedes aegypti mosquitoes possess diverse fitness and vector competence phenotypes. PLoS Pathog 13(12):e1006751. https://doi.org/10.1371/journal.ppat.1006751
Sinkins S, Gould F (2006) Gene drive systems for insect disease vectors. Nat Rev Genet 7:427–435. https://doi.org/10.1038/nrg1870
Biliske JA, Batista PD, Grant CL, Harris HL (2011) The bacteriophage WORiC is the active phage element in wRi of Drosophila simulans and represents a conserved class of WO phages. BMC Microbiol 15(11):251. https://doi.org/10.1186/1471-2180-11-251
Christie GE, Dokland T (2012) Pirates of the Caudovirales. Virology 434(2):210–221. https://doi.org/10.1016/j.virol.2012.10.028
Lang AS, Zhaxybayeva O, Beatty JT (2012) Gene transfer agents: phage-like elements of genetic exchange. Nat Rev Microbiol 10:472–482. https://doi.org/10.1038/nrmicro2802
Westbye AB, Beatty JT, Lang AS (2017) Guaranteeing a captive audience: coordinated regulation of gene transfer agent (GTA) production and recipient capability by cellular regulators. Curr Opin Microbiol 38:122–129. https://doi.org/10.1016/j.mib.2017.05.003
Ding H, Moksa MM, Hirst M, Beatty JT (2014) Draft Genome Sequences of Six Rhodobacter capsulatus strains, YW1, YW2, B6, Y262, R121, and DE442. Genome Announc 2(1):e00050-14. https://doi.org/10.1128/genomeA.00050-14
Fallon AM, Baldridge GD, Higgins LA, Witthuhn BA (2013) Wolbachia from the planthopper Laodelphax striatellus establishes a robust, persistent, streptomycin-resistant infection in clonal mosquito cells. In Vitro Cell Dev Biol- Anim 49:66–73. https://doi.org/10.1007/s11626-012-9571-3
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. https://doi.org/10.1093/nar/25.17.3389
Altschul SF, Wootton JC, Gertz EM, Agarwala R, Morgulis A, Schaffer AA, Yu Y-K (2005) Protein database searches using compositionally adjusted substitution matrices. FEBS J 272:5101–5109. https://doi.org/10.1111/j.1742-4658.2005.04945.x
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30(14):3059–3066. https://doi.org/10.1093/nar/gkf436.PMID:12136088;PMCID:PMC135756
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30(4):772–780. https://doi.org/10.1093/molbev/mst010
Rao VB, Feiss M (2015) Mechanisms of DNA packaging by large double-stranded DNA viruses. Annu Rev Virol 2(1):351–378. https://doi.org/10.1146/annurev-virology-100114-055212
Wieczorek DJ, Didion L, Feiss M (2002) Alterations of the portal protein, gpB, of bacteriophage lambda suppress mutations in cosQ, the site required for termination of DNA packaging. Genetics 161(1):21–31. https://doi.org/10.1093/genetics/161.1.21
Isidro A, Henriques AO, Tavares P (2004) The portal protein plays essential roles at different steps of the SPP1 DNA packaging process. Virology 322(2):253–263
Ivanovska I, Wuite G, Jönsson B, Evilevitch E (2007) Internal DNA pressure modifies stability of WT phage. Proc Natl Acad Sci USA 104(23):9603–9608. https://doi.org/10.1073/pnas.0703166104
Sherlock D, Leong JX, Fogg PCM (2019) Identification of the first gene transfer agent (GTA) small terminase in Rhodobacter capsulatus and its role in GTA production and packaging of DNA. J Virol 93:e01328-e1419. https://doi.org/10.1128/JVI.01328-19
Maxwell KL, Davidson AR, Murialdo H, Gold M (2000) Thermodynamic and functional characterization of protein W from bacteriophage λ. J Biol Chem 275:18879–18886
Kupritz J, Martin J, Fischer K, Curtis KC, Fauver JR, Huang Y, Choi YJ, Beatty WL, Mitreva M, Fischer PU (2021) Isolation and characterization of a novel bacteriophage WO from Allonemobius socius crickets in Missouri. PLoS ONE 16(7):e0250051
Kosinski J, Feder M, Bujnicki JM (2005) The PD-(D/E)XK superfamily revisited: identification of new members among proteins involved in DNA metabolism and functional predictions for domains of (hitherto) unknown function. BMC Bioinform 6:172
Knizewski L, Kinch LN, Grishin NV, Rychlewski L, Ginalski K (2007) Realm of PD-(D/E)XK nuclease superfamily revisited: detection of novel families with modified transitive meta profile searches. BMC Struct Biol 7:40. https://doi.org/10.1186/1472-6807-7-40
Fallon AM (2020) Computational evidence for antitoxins associated with RelE/ParE, RatA, Fic, and AbiEii-family toxins in Wolbachia genomes. Mol Genet Genom 295:891–909. https://doi.org/10.1007/s00438-020-01662-0
Furukawa S, Tanaka K, Ikeda T, Fukatsu T, Sasaki T (2012) Quantitative analysis of the lytic cycle of WO phages infecting Wolbachia. Appl Entomol Zool 47:449–456. https://doi.org/10.1007/s13355-012-0142-6
Miao Y-h, Xiao J-h, Huang D-w (2020) Distribution and evolution of the bacteriophage WO and its antagonism with Wolbachia. Front Microbiol 11:595629. https://doi.org/10.3389/fmicb.2020.595629
Rapala J, Miller B, Garcia M, Dolan M, Bockman M, Hansson M et al (2021) Genomic diversity of bacteriophages infecting Rhodobacter capsulatus and their relatedness to its gene transfer agent RcGTA. PLoS ONE 16(11):e0255262
Fallon AM (2021) DNA recombination and repair in Wolbachia: RecA and related proteins. Mol Genet Genom 296:437–456
Masui S, Kamoda S, Sasaki T, Ishikawa H (2000) Distribution and evolution of bacteriophage WO in Wolbachia, the endosymbiont causing sexual alterations in arthropods. J Mol Evol 51:491–497. https://doi.org/10.1007/s002390010112
Pruss GJ, Calendar R (1978) Maturation of bacteriophage P2 DNA. Virology 86:454–467. https://doi.org/10.1016/0042-6822(78)90085-5
Six EW, Sunshine MG, Williams J, Haggard-Ljungquist E, Lindqvist BH (1991) Morphogenetic switch mutations of bacteriophage P2. Virology 182:34–46. https://doi.org/10.1016/0042-6822(91)90645-r
Chatterjee S, Rothenberg E (2012) Interaction of bacteriophage λ with its E. coli receptor. LamB Viruses 4:3162–3178. https://doi.org/10.3390/v4113162
Gavotte L, Vavre F, Henri H, Ravallec M, Stouthamer R, Boulétreau M (2004) Diversity, distribution and specificity of WO phage infection in Wolbachia of four insect species. Insect Mol Biol 13:147–153
Lang AS, Taylor TA, Beatty JT (2002) Evolutionary implications of phylogenetic analyses of the gene transfer agent (GTA) of Rhodobacter capsulatus. J Mol Evol 55:534–543. https://doi.org/10.1007/s00239-002-2348-7
Acknowledgements
This work was supported by the University of Minnesota Agricultural Experiment Station, St. Paul, MN.
Funding
This work was supported by the University of Minnesota Agricultural Experiment Station, St. Paul, MN, which provided salary to the author. There were no external grants.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares that she has no conflicts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by the author.
Additional information
Edited by A. Lorena Passarelli.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fallon, A.M. Muramidase, nuclease, or hypothetical protein genes intervene between paired genes encoding DNA packaging terminase and portal proteins in Wolbachia phages and prophages. Virus Genes 58, 327–349 (2022). https://doi.org/10.1007/s11262-022-01907-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-022-01907-7