Abstract
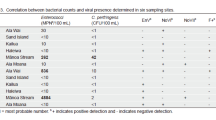
Enteric viruses in the aquatic environment are a concern due to the potential for waterborne disease transmission to humans. In Nepal, the Bagmati River serves as a source of drinking and irrigation water; therefore, the detection of waterborne enteric pathogens is integral to maintaining human health. The objective of this study was to quantify the crAssphage marker in surface water samples from the Bagmati River between November 2015 and September 2016. Concentrations of crAssphage were then compared to those of other enteric viruses and indicator organisms found in the samples in order to examine the potential of crAssphage as a marker for fecal contamination. CrAssphage was detected in 17% (1/6) of samples from Sundarijal, 100% (6/6) of samples from Thapathali, and 100% (6/6) samples from Chovar, with the highest average concentrations recorded in May 2016 and the lowest average concentrations recorded in September 2016. Overall, crAssphage was present in 72% (13/18) of samples and was strongly correlated with the presence of fecal indicator bacteria Escherichia coli (r = 0.89) and Enterococcus (r = 0.92) and several enteric viruses. The strongest viral correlations were to salivirus (r = 0.84), pepper mild mottle virus (r = 0.77), Aichi virus 1 (r = 0.75), enteroviruses (r = 0.76), and tobacco mosaic virus (r = 0.71). These results provide evidence for the potential use of crAssphage as a marker for human fecal contamination in river water.


Similar content being viewed by others
References
Ahmed, W., Hughes, B., & Harwood, V. J. (2016). Current status of marker genes of bacteroides and related taxa for identifying sewage pollution in environmental waters. Water, 8(231). https://doi.org/10.3390/w8060231.
Ahmed, W., Wan, C., Goonetilleke, A., & Gardner, T. (2010). Evaluating sewage-associated JCV and BKV polyomaviruses for sourcing human fecal pollution in a coastal river in Southeast Queensland, Australia. Journal of Environmental Quality, 39(5), 1743–1750, 334. https://doi.org/10.2134/jeq2010.0062.
Ahmed, W., Lobos, A., Senkbeil, J., Peraud, J., Gallard, J., & Harwood, V. J. (2018a). Evaluation of the novel crAssphage marker for sewage pollution tracking in storm drain outfalls in Tampa, 297 Florida. Water Research, 131, 142–150. https://doi.org/10.1016/j.watres.2017.12.011.
Ahmed, W., Payyappat, S., Cassidy, M., Besley, C., & Power, K. (2018b). Novel crAssphage marker genes ascertain sewage pollution in a recreational lake receiving urban stormwater runoff. Water Research, 145, 769–778. https://doi.org/10.1016/j.watres.2018.08.049.
Ballesté, E., Pascual-Benito, M., Martín-Díaz, J., Blanch, A. R., Lucena, F., Muniesa, M., Jofre, J., & García-Aljaro, C. (2019). Dynamics of crAssphage as a human source tracking marker in potentially faecally polluted environments. Water Research, 233–244. https://doi.org/10.1016/j.watres.2019.02.042.
Bibby, K., Crank, K., Greaves, J., Li, X., Wu, Z., Hamza, I., & Stachler, E. (2019). Metagenomics and the development of viral water quality tools. npj Clean Water, 2(1), 1–13. https://doi.org/10.1038/s41545-019-0032-3.
Chen, H., Bai, X., Li, Y., Jing, L., Chen, R., & Teng, Y. (2019a). Characterization and source- tracking of antibiotic resistomes in the sediments of a peri-urban river. Science of the Total Environment, 679, 88–96. https://doi.org/10.1016/j.scitotenv.2019.05.063.
Chen, H., Bai, X., Li, Y., Jing, L., Chen, R., & Teng, Y. (2019b). Source identification of antibiotic resistance genes in a peri-urban river using novel crAssphage marker genes and metagenomic 359 signatures. Water Research, 167, 115098. https://doi.org/10.1016/j.watres.2019.115098.
Farkas, K., Adriaenssens, E. M., Walker, D. I., McDonald, J. E., Malham, S. K., & Jones, D. L. (2019). Critical evaluation of crAssphage as a molecular marker for human-derived wastewater contamination in the aquatic environment. Food and Environmental Virology, 11(2) 252, 113–119. https://doi.org/10.1007/s12560-019-09369-1.
Feachem, R. G., Bradley, D. J., Garelick, H., & Mara, D. D. (1983). Sanitation and disease: health aspects of excreta and wastewater management. New York: Wiley.
García-Aljaro, C., Ballesté, E., Muniesa, M., & Jofre, J. (2017a). Determination of crAssphage in water samples and applicability for tracking human faecal pollution. Microbial Biotechnology, 10(6), 1775–1780. https://doi.org/10.1111/1751-7915.12841.
García-Aljaro, C., Martín-Díaz, J., Viñas-Balada, E., Calero-Cáceres, W., Lucenca, F., & Blanch, A. R. (2017b). Mobilisation of microbial indicators, microbial source tracking markers and pathogens after rainfall events. Water Research, 112, 248–253. https://doi.org/10.1016/j.watres.2017.02.003.
Ghaju Shrestha, R., Tandukar, S., Bhandari, D., Sherchan, S. P., Tanaka, Y., Sherchand, J. B., & Haramoto, E. (2019). Prevalence of Arcobacter and other pathogenic bacteria in river water 294 in Nepal. Water, 11, 1416. https://doi.org/10.3390/w11071416.
Guerin, E., Shkoporov, A., Stockdale, S. R., Clooney, A. G., Ryan, F. J., Sutton, T. D. S., Draper, L. A., Gonzalez-Tortuero, E., Ross, R. P., & Hill, C. (2018). Biology and taxonomy of crAss- like bacteriophages, the most abundant virus in the human gut. Cell Host & Microbe, 24(5), 653–664. https://doi.org/10.1016/j.chom.2018.10.002.
Haramoto, E., Yamada, K., & Nishida, K. (2011). Prevalence of protozoa, viruses, coliphages and indicator bacteria in groundwater and river water in the Kathmandu Valley, Nepal. Transactions of the Royal Society of Tropical Medicine and Hygiene, 105, 711–716, 267. https://doi.org/10.1016/j.trstmh.2011.08.004.
Haramoto, E., Kitajima, M., Hata, A., Torrey, J. R., Masago, Y., Sano, D., & Katayama, H. (2018). A review on recent progress in the detection methods and prevalence of human enteric viruses 316 in water. Water Research, 135, 168–186. https://doi.org/10.1016/j.wateres.2018.02.004.
Karkman, A., Pärnänen, J., & Larsson, D. G. (2019). Fecal pollution can explain antibiotic resistance gene abundances in anthropogenically impacted environments. Nature Communications, 10. https://doi.org/10.1038/s41467-018-07992-3.
Kongprajug, A., Mongkolsuk, S., & Sirikanchana, K. (2019). CrAssphage as a potential human sewage marker for microbial source tracking in Southeast Asia. Environmental Science and Technology Letters, 6(3), 159–164. https://doi.org/10.1021/acs.estlett.9b00041.
Malla, B., Ghaju Shrestha, R., Tadukar, S., Sherchand, J. B., & Haramoto, E. (2019a). Performance evaluation of human-specific viral markers and application of pepper mild mottle virus and crAssphage to environmental water samples as fecal pollution markers in the Kathmand Valley. Nepal. Food and Environmental Virology, 11(3), 274–287. https://doi.org/10.1007/s12560-019-09389-x.
Malla, B., Makise, K., Nakaya, K., Mochizuki, T., Yamada, T., & Haramoto, E. (2019b). Evaluation of human- and animal-specific viral markers and application of CrAssphage, pepper mild mottle virus, and tobacco mosaic virus as potential fecal pollution markers to river water in Japan. Food and Environmental Virology. https://doi.org/10.1007/s12560-019-09398-w.
Mishra, B. K., Regmi, R. K., Masago, Y., Fukushi, K., Kumar, P., & Saraswat, C. (2017). Assessment of Bagmati river pollution in Kathmandu Valley: scenario-based modeling and analysis for sustainable urban development. Sustainability of Water Quality and Ecology, 9(10), 67–77. https://doi.org/10.1016/j.swaqe.2017.06.001.
Nielsen, A. C. Y., Gyhrs, M. L., Nielsen, L. P., Pedersen, C., & Bӧttiger, B. (2013). Gastroenteritis and the novel picornaviruses Aichi virus, cosavirus, saffold virus, and salivirus in young children. Journal of Clinical Virology, 57(3), 239–242. https://doi.org/10.1016/j.jcv.2013.03.015.
Panthi, J., Li, F., Wang, H., Aryal, S., Dahal, P., Ghimire, S., & Kabenge, M. (2017). Evaluating climactic and non-climactic stresses for declining surface water quality in Bagmati River of Nepal. Environmental Monitoring and Assessment, 189(6), 1–15. https://doi.org/10.1007/s10661-017-6000-9.
Shrestha, S., Nakamura, T., Magome, J., Aihara, Y., Kondo, N., Haramoto, E., Malla, B., Shindo, J., & Nishida, K. (2018a). Groundwater use and diarrhoea in urban Nepal: Novel application of a geostatistical interpolation technique linking environmental and epidemiologic survey data. International Health, 10(5), 324–332. https://doi.org/10.1093/inthealth/ihy037.
Shrestha, S., Shrestha, S., Shindo, J., Sherchand, J. B., & Haramoto, E. (2018b). Virological quality of irrigation water sources and pepper mild mottle virus and tobacco mosaic virus as index of pathogenic virus contamination level. Food and Environmental Virology, 10(1), 107–120. https://doi.org/10.1007/s12560-017-9324-2.
Stachler, E., & Bibby, K. (2014). Metagenomic evaluation of the highly abundant human gut bacteriophage crAssphage for source tracking of human fecal pollution. Environmental Science and Technology Letters, 1(10), 405–409. https://doi.org/10.1021/ez500266s.
Stachler, E., Kelty, C., Sivaganesan, M., Li, X., Bibby, K., & Shanks, O. C. (2017). Quantitative crAssphage PCR assays for human fecal pollution measurement. Environmental Science and Technology, 51(16), 9146–9154. https://doi.org/10.1021/acs.est.7b02703.
Stachler, E., Akyon, B., Aquino de Carvalho, N., Ference, C., & Bibby, K. (2018). Correlation of crAssphage qPCR Markers with culturable and molecular indicators of human fecal pollution in an impacted urban watershed. Environmental Science and Technology, 52(13), 7505–7512, 256. https://doi.org/10.1021/acs.est.8b00638.
Stachler, E., Crank, K., & Bibby, K. (2019). Co-occurrence of crAssphage with antibiotic resistance genes in an impacted urban watershed. Environmental Science and Technology Letters, 6(4), 216–221. https://doi.org/10.1021/acs.estlett.9b00130.
Symonds, E. M., Rosario, K., & Breitbart, M. (2019). Pepper mild mottle virus: agricultural menace turned effective tool for microbial water quality monitoring and assessing (waste)water treatment technologies. PLoS Pathogens, 15(4). https://doi.org/10.1371/journal.ppat.1007639.
Tandukar, S., Sherchand, J., Bhandari, D., Sherchan, S., Malla, B., Ghaju Shrestha, R., & Haramoto, E. (2018). Presence of human enteric viruses, protozoa, and indicators of pathogens in the Bagmati River, Nepal. Pathogens, 7(2), 38. https://doi.org/10.3390/pathogens7020038.
Tornevi, A., Bergstedt, O., & Forsberg, B. (2014). Precipitation effects on microbial pollution in a river: Lag structures and seasonal effect modification. PLoS One, 9(5). https://doi.org/10.1371/journal.pone.0098546.
Funding
This study was supported by startup funds to Dr. Samendra Sherchan and Kurita Water and Environment Foundation, the Science and Technology Research Partnership for Sustainable Development (SATREPS) project entitled “Hydro-microbiological approach for the water security in Kathmandu Valley, Nepal,” and the Japan Society for the Promotion of Science (JSPS) through a Grant-in-Aid for Scientific Research (B) (grant number JP17H03332) and the Fund for the Promotion of Joint International Research (Fostering Joint International Research (B)) (grant number JP18KK0297) to Dr. Eiji Haramoto.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ward, L.M., Ghaju Shrestha, R., Tandukar, S. et al. Evaluation of CrAssphage Marker for Tracking Fecal Contamination in River Water in Nepal. Water Air Soil Pollut 231, 282 (2020). https://doi.org/10.1007/s11270-020-04648-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-020-04648-1