Abstract
The freshwater pearl mussel Margaritifera margaritifera is one of the most threatened freshwater bivalves worldwide. In this study, we aimed (i) to study the processes by which water quality might affect freshwater mussels in situ and (ii) to provide insights into the ecotoxicological significance of water pollution to natural populations in order to provide necessary information to enhance conservation strategies. M. margaritifera specimens were sampled in two close sites located upstream or downstream from an illegal dumping site. The renal transcriptome of these animals was assembled and gene transcription determined by RNA-seq. Correlations between transcription levels of each single transcript and the bioaccumulation of nine trace metals, age (estimated by sclerochronology), and condition index were determined in order to identify genes likely to respond to a specific factor. Amongst the studied metals, Cr, Zn, Cd, and Ni were the main factors correlated with transcription levels, with effects on translation, apoptosis, immune response, response to stimulus, and transport pathways. However, the main factor explaining changes in gene transcription appeared to be the age of individuals with a negative correlation with the transcription of retrotransposon-related genes. To investigate this effect further, mussels were classified into three age classes. In young, middle-aged and old animals, transcription levels were mainly explained by Cu, Zn and age, respectively. This suggests differences in the molecular responses of this species to metals during its lifetime that must be better assessed in future ecotoxicology studies.


Similar content being viewed by others
References
Alexa A, Rahnenfuhrer J (2016) topGO: enrichment analysis for gene ontology. R Package Version 2280
Auffret M, Oubella R (1994) Cytometric parameters of bivalve molluscs: effect of environmental factors. Modulators of fish immune responses. In: Models Environ. Toxicol. Biomark. Immunostimul. Stolen, J.S., Fletcher, T.C., pp 22–32
Baillon L, Pierron F, Coudret R et al (2015) Transcriptome profile analysis reveals specific signatures of pollutants in Atlantic eels. Ecotoxicol Lond Engl 24:71–84. https://doi.org/10.1007/s10646-014-1356-x
Bellas J, Albentosa M, Vidal-Liñán L et al (2014) Combined use of chemical, biochemical and physiological variables in mussels for the assessment of marine pollution along the N-NW Spanish coast. Mar Environ Res 96:105–117. https://doi.org/10.1016/j.marenvres.2013.09.015
Bouchut A, Roger E, Coustau C et al (2006) Compatibility in the Biomphalaria glabrata/Echinostoma caproni model: potential involvement of adhesion genes. Int J Parasitol 36:175–184. https://doi.org/10.1016/j.ijpara.2005.09.009
Cabau C, Escudié F, Djari A, et al (2016) Compacting and correcting Trinity and Oases RNA-Seq de novo assemblies. PeerJ Preprints
Camacho C, Coulouris G, Avagyan V et al (2009) BLAST+: architecture and applications. BMC Bioinformatics 10:421. https://doi.org/10.1186/1471-2105-10-421
Canesi L, Viarengo A (1997) Age-related differences in glutathione metabolism in mussel tissues (Mytilus edulis L.) Comp Biochem Physiol - B Biochem Mol Biol 116:217–221. https://doi.org/10.1016/S0305-0491(96)00223-4
Carella F, Villari G, Maio N, De Vico G (2016) Disease and disorders of freshwater unionid mussels: a brief overview of recent studies. Front Physiol. https://doi.org/10.3389/fphys.2016.00489
Casacuberta E, González J (2013) The impact of transposable elements in environmental adaptation. Mol Ecol 22:1503–1517. https://doi.org/10.1111/mec.12170
Causeur D, Friguet C, Houee-Bigot M, Kloareg M (2011) Factor analysis for multiple testing (FAMT): an R package for large-scale significance testing under dependence. J Stat Softw. 10.18637/jss.v040.i14
Chambers JE, Dalton LE, Clarke HJ et al (2015) Actin dynamics tune the integrated stress response by regulating eukaryotic initiation factor 2α dephosphorylation. elife. https://doi.org/10.7554/eLife.04872
Coles JA, Farley SR, Pipe R (1995) Alteration of the immune response of the common marine mussel Mytilus edulis resulting from exposure to cadmium. Dis Aquat Org 22:59–65. https://doi.org/10.3354/dao022059
Connon RE, Geist J, Werner I (2012) Effect-based tools for monitoring and predicting the ecotoxicological effects of chemicals in the aquatic environment. Sensors 12:12741–12771. https://doi.org/10.3390/s120912741
Conrado AB, D’Angelantonio M, Torreggiani A et al (2014) Reactivity of hypotaurine and cysteine sulfinic acid toward carbonate radical anion and nitrogen dioxide as explored by the peroxidase activity of Cu,Zn superoxide dismutase and by pulse radiolysis. Free Radic Res 48:1300–1310. https://doi.org/10.3109/10715762.2014.951839
Cope WG, Bringolf RB, Buchwalter DB et al (2008) Differential exposure, duration, and sensitivity of unionoidean bivalve life stages to environmental contaminants. J North Am Benthol Soc 27:451–462. https://doi.org/10.1899/07-094.1
Cousins RJ (1998) A role of zinc in the regulation of gene expression. Proc Nutr Soc 57:307–311. https://doi.org/10.1079/PNS19980045
Cuttelod A, Seddon M, Neubert E (2011) European Red List of Non-marine Molluscs. Publications Office of the European Union, Luxembourg
de Almagro MC, Vucic D (2012) The inhibitor of apoptosis (IAP) proteins are critical regulators of signaling pathways and targets for anti-cancer therapy. Exp Oncol 34:200–211
De Cecco M, Criscione SW, Peterson AL et al (2013) Transposable elements become active and mobile in the genomes of aging mammalian somatic tissues. Aging 5:867–883
de Nadal E, Ammerer G, Posas F (2011) Controlling gene expression in response to stress. Nat Rev Genet 12:833–845. https://doi.org/10.1038/nrg3055
De Schutter K, Van Damme EJM (2015) Protein-carbohydrate interactions as part of plant defense and animal immunity. Molecules 20:9029–9053. https://doi.org/10.3390/molecules20059029
Defo MA, Pierron F, Spear PA et al (2012) Evidence for metabolic imbalance of vitamin A2 in wild fish chronically exposed to metals. Ecotoxicol Environ Saf 85:88–95. https://doi.org/10.1016/j.ecoenv.2012.08.017
Dunca E, Soderberg H, Norrgrann O (2011) Shell growth and age determination in the freshwater pearl mussel Margaritifera margaritifera in Sweden: natural versus limed streams. In: Rearing of unionoid mussels (with special emphasis on the Freshwater Pearl Mussel Margaritifera margaritifera). Thielen Frank (ed), pp 48–58
Edgar R, Domrachev M, Lash AE (2002) Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30:207–210. https://doi.org/10.1093/nar/30.1.207
Ferreira VP, Pangburn MK, Cortés C (2010) Complement control protein factor H: the good, the bad, and the inadequate. Mol Immunol 47:2187–2197. https://doi.org/10.1016/j.molimm.2010.05.007
Flores-Nunes F, Gomes T, Company R et al (2015) Changes in protein expression of pacific oyster Crassostrea gigas exposed in situ to urban sewage. Environ Sci Pollut Res Int 22:17267–17279. https://doi.org/10.1007/s11356-014-3821-8
Friguet C, Kloareg M, Causeur D (2009) A factor model approach to multiple testing under dependence. J Am Stat Assoc 104:1406–1415. https://doi.org/10.1198/jasa.2009.tm08332
Geist J (2010) Strategies for the conservation of endangered freshwater pearl mussels (Margaritifera margaritifera L.): a synthesis of conservation genetics and ecology. Hydrobiologia 644:69–88. https://doi.org/10.1007/s10750-010-0190-2
Gilgès D, Vinit M-A, Callebaut I et al (2000) Polydom: a secreted protein with pentraxin, complement control protein, epidermal growth factor and von Willebrand factor A domains. Biochem J 352:49–59. https://doi.org/10.1042/bj3520049
Gillis PL, McInnis R, Salerno J et al (2017) Freshwater mussels in an urban watershed: impacts of anthropogenic inputs and habitat alterations on populations. Sci Total Environ 574:671–679. https://doi.org/10.1016/j.scitotenv.2016.09.110
Gonzalez P, Pierron F (2015) Omics in aquatic ecotoxicology: the ultimate response to biological questions? In: Amiard J-C, Mouneyrac C (eds) Amiard-Triquet C. Academic Press, Aquatic Ecotoxicology, pp 183–203
González VL, Andrade SCS, Bieler R et al (2015) A phylogenetic backbone for Bivalvia: an RNA-seq approach. Proc Biol Sci 282:20142332. https://doi.org/10.1098/rspb.2014.2332
Grabherr MG, Haas BJ, Yassour M et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29:644–652. https://doi.org/10.1038/nbt.1883
Gruber H, Wessels W, Boynton P, et al (2015) Age-related cellular changes in the long-lived bivalve A. islandica. Age. doi: https://doi.org/10.1007/s11357-015-9831-8
Gupta A, Lutsenko S (2009) Human copper transporters: mechanism, role in human diseases and therapeutic potential. Future Med Chem 1:1125–1142. https://doi.org/10.4155/fmc.09.84
Harding HP, Zhang Y, Scheuner D et al (2009) Ppp1r15 gene knockout reveals an essential role for translation initiation factor 2 alpha (eIF2α) dephosphorylation in mammalian development. Proc Natl Acad Sci U S A 106:1832–1837. https://doi.org/10.1073/pnas.0809632106
Husmann G, Abele D, Rosenstiel P et al (2014) Age-dependent expression of stress and antimicrobial genes in the hemocytes and siphon tissue of the Antarctic bivalve, Laternula elliptica, exposed to injury and starvation. Cell Stress Chaperones 19:15–32. https://doi.org/10.1007/s12192-013-0431-1
Ivanov VA, Melnikov AA, Siunov AV et al (1991) Authentic reverse transcriptase is coded by jockey, a mobile Drosophila element related to mammalian LINEs. EMBO J 10:2489–2495
Kerambrun E, Rioult D, Delahaut L et al (2016) Variations in gene expression levels in four European zebra mussel, Dreissena polymorpha, populations in relation to metal bioaccumulation: a field study. Ecotoxicol Environ Saf 134(Part 1):53–63. https://doi.org/10.1016/j.ecoenv.2016.08.018
Kültz D (2005) Molecular and evolutionary basis of the cellular stress response. Annu Rev Physiol 67:225–257. https://doi.org/10.1146/annurev.physiol.67.040403.103635
Lagesen K, Hallin P, Rødland EA et al (2007) RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 35:3100–3108. https://doi.org/10.1093/nar/gkm160
Lauer MM, de Oliveira CB, Yano NLI, Bianchini A (2012) Copper effects on key metabolic enzymes and mitochondrial membrane potential in gills of the estuarine crab Neohelice granulata at different salinities. Comp Biochem Physiol Part C Toxicol Pharmacol 156:140–147. https://doi.org/10.1016/j.cbpc.2012.08.001
Leulier F, Lhocine N, Lemaitre B, Meier P (2006) The Drosophila inhibitor of apoptosis protein DIAP2 functions in innate immunity and is essential to resist gram-negative bacterial infection. Mol Cell Biol 26:7821–7831. https://doi.org/10.1128/MCB.00548-06
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li H, Handsaker B, Wysoker A et al (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25:2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Liu H, Chen X, Kang IJ et al (2016) The valve movement response of three freshwater mussels Corbicula fluminea Müller 1774, Hyriopsis cumingii Lea 1852, and Anodonta woodiana Lea 1834 exposed to copper. Hydrobiologia 770:1–13. https://doi.org/10.1007/s10750-015-2560-2
Lopes-Lima M, Teixeira A, Froufe E et al (2014) Biology and conservation of freshwater bivalves: past, present and future perspectives. Hydrobiologia 735:1–13. https://doi.org/10.1007/s10750-014-1902-9
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Lummer E-M, Auerswald K, Geist J (2016) Fine sediment as environmental stressor affecting freshwater mussel behavior and ecosystem services. Sci Total Environ 571:1340–1348. https://doi.org/10.1016/j.scitotenv.2016.07.027
Lydeard C, Cowie RH, Ponder WF et al (2004) The global decline of nonmarine mollusks. Bioscience 54:321–330. https://doi.org/10.1641/0006-3568(2004)054[0321:TGDONM]2.0.CO;2
Machordom A, Araujo R, Erpenbeck D, Ramos M-Á (2003) Phylogeography and conservation genetics of endangered European Margaritiferidae (Bivalvia: Unionoidea). Biol J Linn Soc 78:235–252. https://doi.org/10.1046/j.1095-8312.2003.00158.x
Mariette J, Noirot C, Nabihoudine I et al (2014) RNAbrowse: RNA-Seq de novo assembly results browser. PLoS One 9:e96821. https://doi.org/10.1371/journal.pone.0096821
McKenna A, Hanna M, Banks E et al (2010) The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20:1297–1303. https://doi.org/10.1101/gr.107524.110
Meng J, Zhang L, Li L et al (2015) Transcription factor CgMTF-1 regulates CgZnT1 and CgMT expression in Pacific oyster (Crassostrea gigas) under zinc stress. Aquat Toxicol 165:179–188. https://doi.org/10.1016/j.aquatox.2015.05.023
Miousse IR, Chalbot M-CG, Lumen A et al (2015) Transposable elements in response to environmental stressors. Mutat Res Rev Mutat Res 765:19–39. https://doi.org/10.1016/j.mrrev.2015.05.003
Moise AR, Kuksa V, Imanishi Y, Palczewski K (2004) Identification of all-trans-retinol:all-trans-13,14-dihydroretinol saturase*. J Biol Chem 279:50230–50242. https://doi.org/10.1074/jbc.M409130200
Moise AR, Lobo GP, Erokwu B et al (2010) Increased adiposity in the retinol saturase-knockout mouse. FASEB J 24:1261–1270. https://doi.org/10.1096/fj.09-147207
Nishimura T, Duereh M, Sugita Y et al (2015) Protective effect of hypotaurine against oxidative stress-induced cytotoxicity in rat placental trophoblasts. Placenta 36:693–698. https://doi.org/10.1016/j.placenta.2015.02.014
Ozato K, Shin D-M, Chang T-H, Morse HC (2008) TRIM family proteins and their emerging roles in innate immunity. Nat Rev Immunol 8:849–860. https://doi.org/10.1038/nri2413
Paul-Pont I, Gonzalez P, Baudrimont M et al (2010) Interactive effects of metal contamination and pathogenic organisms on the marine bivalve Cerastoderma edule. Mar Pollut Bull 60:515–525. https://doi.org/10.1016/j.marpolbul.2009.11.013
Petris MJ, Mercer JF, Culvenor JG et al (1996) Ligand-regulated transport of the Menkes copper P-type ATPase efflux pump from the Golgi apparatus to the plasma membrane: a novel mechanism of regulated trafficking. EMBO J 15:6084–6095
Philipp EER, Abele D (2010) Masters of longevity: lessons from long-lived bivalves—a mini-review. Gerontology 56:55–65. https://doi.org/10.1159/000221004
Pierron F, Normandeau E, Defo MA et al (2011) Effects of chronic metal exposure on wild fish populations revealed by high-throughput cDNA sequencing. Ecotoxicology 20:1388–1399. https://doi.org/10.1007/s10646-011-0696-z
Polishchuk R, Lutsenko S (2013) Golgi in copper homeostasis: a view from the membrane trafficking field. Histochem Cell Biol 140:285–295. https://doi.org/10.1007/s00418-013-1123-8
Quayle DB, Newkirk GF (1989) Farming bivalve molluscs: methods for study and development. World Aquaculture Society in association with the International Development Research Centre. Baton Rouge, LA
Quevillon E, Silventoinen V, Pillai S et al (2005) InterProScan: protein domains identifier. Nucleic Acids Res 33:W116–W120. https://doi.org/10.1093/nar/gki442
Regier N, Baerlocher L, Münsterkötter M et al (2013) Analysis of the Elodea nuttallii transcriptome in response to mercury and cadmium pollution: development of sensitive tools for rapid ecotoxicological testing. Environ Sci Technol 47:8825–8834. https://doi.org/10.1021/es401082h
Regoli F, Giuliani ME (2014) Oxidative pathways of chemical toxicity and oxidative stress biomarkers in marine organisms. Mar Environ Res 93:106–117. https://doi.org/10.1016/j.marenvres.2013.07.006
Regoli F, Winston GW, Gorbi S et al (2003) Integrating enzymatic responses to organic chemical exposure with total oxyradical absorbing capacity and DNA damage in the European eel Anguilla anguilla. Environ Toxicol Chem 22:2120–2129. https://doi.org/10.1897/02-378
Reynders H, van der Ven K, Moens LN et al (2006) Patterns of gene expression in carp liver after exposure to a mixture of waterborne and dietary cadmium using a custom-made microarray. Aquat Toxicol 80:180–193. https://doi.org/10.1016/j.aquatox.2006.08.009
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. Methods Mol Biol 132:365–386
Sahin E, DePinho RA (2010) Linking functional decline of telomeres, mitochondria and stem cells during ageing. Nature 464:520–528. https://doi.org/10.1038/nature08982
Saleem M, Qadir MI, Perveen N et al (2013) Inhibitors of apoptotic proteins: new targets for anticancer therapy. Chem Biol Drug Des 82:243–251. https://doi.org/10.1111/cbdd.12176
Sato-Nishiuchi R, Nakano I, Ozawa A et al (2012) Polydom/SVEP1 is a ligand for integrin α9β1. J Biol Chem 287:25615–25630. https://doi.org/10.1074/jbc.M112.355016
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675. https://doi.org/10.1038/nmeth.2089
Schöne BR, Dunca E, Fiebig J, Pfeiffer M (2005) Mutvei’s solution: an ideal agent for resolving microgrowth structures of biogenic carbonates. Palaeogeogr Palaeoclimatol Palaeoecol 228:149–166. https://doi.org/10.1016/j.palaeo.2005.03.054
Sciamanna I, De Luca C, Spadafora C (2016) The reverse transcriptase encoded by LINE-1 retrotransposons in the genesis, progression, and therapy of cancer. Front Chem. https://doi.org/10.3389/fchem.2016.00006
Seiler GR, Morse MP (1988) Kidney and hemocytes of Mya arenaria (Bivalvia): normal and pollution-related ultrastructural morphologies. J Invertebr Pathol 52:201–214
Sheir SK, Handy RD, Henry TB (2013) Effect of pollution history on immunological responses and organ histology in the marine mussel Mytilus edulis exposed to cadmium. Arch Environ Contam Toxicol 64:701–716. https://doi.org/10.1007/s00244-012-9868-y
Smith AFA, Hubley R, Green P RepeatMasker Open-4.0
Strayer DL, Downing JA, Haag WR et al (2004) Changing perspectives on pearly mussels, North America’s most imperiled animals. Bioscience 54:429–439. https://doi.org/10.1641/0006-3568(2004)054[0429:CPOPMN]2.0.CO;2
Stubblefield WA, Steadman BL, La Point TW, Bergman HL (1999) Acclimation-induced changes in the toxicity of zinc and cadmium to rainbow trout. Environ Toxicol Chem 18:2875–2881. https://doi.org/10.1002/etc.5620181231
Supek F, Bošnjak M, Škunca N, Šmuc T (2011) REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS One 6:e21800. https://doi.org/10.1371/journal.pone.0021800
Taylor HH, Anstiss JM (1999) Copper and haemocyanin dynamics in aquatic invertebrates. Mar Freshw Res 50:907–931
Thomas GR, Taylor J, Garcia de Leaniz C (2010) Captive breeding of the endangered freshwater pearl mussel Margaritifera margaritifera. Endanger Species Res 12:1
Tomanek L (2012) Environmental proteomics of the mussel Mytilus: implications for tolerance to stress and change in limits of biogeographic ranges in response to climate change. Integr Comp Biol 52:648–664. https://doi.org/10.1093/icb/ics114
Uren Webster TM, Bury N, van Aerle R, Santos EM (2013) Global transcriptome profiling reveals molecular mechanisms of metal tolerance in a chronically exposed wild population of brown trout. Environ Sci Technol 47:8869–8877. https://doi.org/10.1021/es401380p
Vaughn CC, Hakenkamp CC (2001) The functional role of burrowing bivalves in freshwater ecosystems. Freshw Biol 46:1431–1446. https://doi.org/10.1046/j.1365-2427.2001.00771.x
Williams TD, Diab AM, George SG et al (2006) Development of the GENIPOL European flounder (Platichthys flesus) microarray and determination of temporal transcriptional responses to cadmium at low dose. Environ Sci Technol 40:6479–6488. https://doi.org/10.1021/es061142h
Yang J, Huang T, Petralia F et al (2015) Synchronized age-related gene expression changes across multiple tissues in human and the link to complex diseases. Sci Rep 5:15145. https://doi.org/10.1038/srep15145
Young M (2005) A literature review of the water quality requirements of the freshwater pearl mussel (Margaritifera margaritifera) and related freshwater bivalves. Scott. Nat. Herit. Comm. Rep. No 084 ROAME No F01AC609d
Zhang G, Fang X, Guo X et al (2012) The oyster genome reveals stress adaptation and complexity of shell formation. Nature 490:49–54. https://doi.org/10.1038/nature11413
Acknowledgements
This work was supported by the European Union program LIFE+ Nature (project LIFE13 NAT/FR/000506). We are grateful to all the staff members of the Perigord-Limousin national park for their contribution to this project. We thank the four anonymous reviewers for their careful reading and insightful comments on our work.
Author information
Authors and Affiliations
Contributions
AB, FP, JT, CK, PG, and MB wrote the manuscript; MB designed the experiment; AB, FP, and CK analyzed the results. JT and JB performed sclerochronological analysis. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
The collection of M. margaritifera for this study was allowed by the prefectoral decree 60/2008 delivered by the Préfecture de la Dordogne (DIREN Aquitaine) on the 31st of October 2008. In France, no ethical permits are required to carry out research on bivalves. Therefore, all procedures were conducted according to the ethical guidelines of France to ensure ethical appropriateness.
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Responsible editor: Philippe Garrigues
Electronic supplementary material
Online Resource 1
Complete annotation results, including GO terms, of the transcriptome sequenced from M. margaritifera kidney (XLSX 2808 kb)
Online Resource 2
List of genes tested by real-time quantitative PCR. (E: primers efficiency, CV: coefficient of variation for control genes) (XLSX 14 kb)
Online Resource 3
Comparison of expression profiles obtained from RNA-seq and RT-qPCR (PDF 167 kb)
Online Resource 4
List of contigs that varied significantly (FDR ≤ 0.05) between upstream and downstream sites (XLSX 48 kb)
Online Resource 5
Heatmap of sample-to-sample distances computed with the DESeq2 package. Site and year are indicated at the bottom of the figure (Down: downstream, Up: upstream, 09: 2009, 10:2010). The age class is indicated on the right (PDF 122 kb)
Online Resource 6
List of the 11,264 contigs with expression level that correlates with one specific factor based on the FAMT analysis (FDR ≤ 0.01) (XLSX 232 kb)
Online Resource 7
List of contigs belonging to Gene Ontology terms that are enriched in each FAMT categories (r: Pearson correlation coefficient) (XLSX 22 kb)
Online Resource 8
List of contigs belonging to the most represented GO Biological Processes related to Ni, Cd and Pb (r: Pearson correlation coefficient) (XLSX 18 kb)
Online Resource 9
Details of GO enrichment test (FDR ≤ 0.05) for contigs related to Cu in young mussels and list of contigs involved in transport, catalytic activity and fatty acid metabolism (r: Pearson correlation coefficient) (XLSX 52 kb)
Online Resource 10
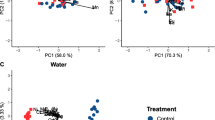
Principal components analysis (PCA) plot of M. margaritifera digestive glands contamination profile. Variables are represented with black arrows (As and Ni overlap). The two first principal components are plotted with the proportion of variance explained by each component shown on the axis. Symbols correspond either to (A) sampling group or (B) age class (PDF 92 kb)
Online Resource 11
Details of GO enrichment test (FDR ≤ 0.05) for contigs related to Zn in middle-aged mussels and list of contigs involved in gene expression and response to chemical metabolism (r: Pearson correlation coefficient) (XLSX 20 kb)
Online Resource 12
Details of GO enrichment test (FDR ≤ 0.05) for contigs related to Age in old mussels and list of contigs from the GO category “mitochondrion” (r: Pearson correlation coefficient) (XLSX 20 kb)
Online Resource 13
Correlations between the original variables and the Principal Components for the PCA analysis of metal concentrations (XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Bertucci, A., Pierron, F., Thébault, J. et al. Transcriptomic responses of the endangered freshwater mussel Margaritifera margaritifera to trace metal contamination in the Dronne River, France. Environ Sci Pollut Res 24, 27145–27159 (2017). https://doi.org/10.1007/s11356-017-0294-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-017-0294-6