Abstract
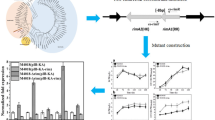
The two-component system “AfsQ1/Q2” plays a crucial role to activate the production of antibiotics ACT, RED, and CDA through directly binding the promoters of pathway-specific activator genes actII-ORF4, redZ, and cdaR respectively when grown under glutamate-supplemented minimal medium in Streptomyces coelicolor. In this report, we demonstrated that the RspA1/A2 (a homologous protein of two-component system AfsQ1/Q2) plays a regulatory role in salinomycin biosynthesis in Streptomyces albus. Gene deletion and complementation experiments showed that the RspA1/A2 promoted salinomycin production but inhibited cell growth when cultured in YMG medium supplemented with 3% soybean oil. More importantly, RspA1/A2 strengthens salinomycin biosynthesis by directly affecting the transcription of the pathway-specific activator gene slnR. Meanwhile, RspA1/A2 plays a negative role in the regulation of nitrogen assimilation and urea decarboxylation by interacting with the promoters of genes gdhA, glnA, amtB, and SLNWT_1828/1829. Gene sigW is located downstream of rspA1/A2 and encodes an extracytoplasmic function sigma factor. Moreover, it negatively regulates the salinomycin biosynthesis and promotes cell growth, which antagonizes the function of RspA1/A2. In short, these useful findings are proved helpful to enrich the understanding of the regulatory pathways of antibiotic biosynthesis by an ECF σ factor-TCS signal transduction system in Streptomyces.






Similar content being viewed by others
References
Worthen, D. B. (2008). Streptomyces in Nature and Medicine: The Antibiotic Makers. Journal of the History of Medicine and Allied Sciences, 63(272), 273–274.
Gumila, C., Ancelin, M. L., Delort, A. M., Jeminet, G., & Vial, H. J. (1997). Characterization of the potent in vitro and in vivo antimalarial activities of ionophore compounds. Antimicrobial Agents and Chemotherapy, 41(3), 523–529.
Jianqiang, H., Jing, S., Virginie, M., BjRn, S., David, W., Bibb, M. J., Nitsara, K., Chih-Jian, L., Kao, C. M., & Buttner, M. J. (2010). Cross-regulation among disparate antibiotic biosynthetic pathways of Streptomyces coelicolor. Molecular Microbiology, 58, 1276–1287.
Zhu, Z., Li, H., Yu, P., Guo, Y., Luo, S., Chen, Z., Mao, X., Guan, W., & Li, Y. (2017). SlnR is a positive pathway-specific regulator for salinomycin biosynthesis in Streptomyces albus. Applied Microbiology and Biotechnology.
Chang, H. M., Chen, M. Y., Shieh, Y. T., Bibb, M. J., & Chen, C. W. (1996). The CutR/S signal transduction system of Streptomyces lividans represses the biosynthesis of the polyketide antibiotic actinorhodin. Molecular Microbiology, 21, 1075–1085.
Mascher, T., Helmann, J. D., & Unden, G. (2006). Stimulus perception in bacterial signal-transducing histidine kinases. Microbiology and Molecular Biology Reviews, 70(4), 910–938.
Krell, T., Lacal, J., Busch, A., Silvajiménez, H., Guazzaroni, M. E., & Ramos, J. L. (2010). Bacterial sensor kinases: Diversity in the recognition of environmental signals. Annual Review of Microbiology, 64(1), 539–559.
Bourret, R. B., & Silversmith, R. E. (2010). Two-component signal transduction. Annual Review of Biochemistry, 13, 113–115.
Alberto, S. L., Antonio, R. G., Alexander Kristian, A., & Martín, J. F. (2008). Target genes and structure of the direct repeats in the DNA-binding sequences of the response regulator PhoP in Streptomyces coelicolor. Nucleic Acids Research, 36, 1358–1368.
Hutchings, M., Hong, H., & Buttner, M. (2010). The vancomycin resistance VanR/S two-component signal transduction system of Streptomyces coelicolor. Molecular Microbiology, 59, 923–935.
Zaburannyi, N., Rabyk, M., Ostash, B., Fedorenko, V., & Luzhetskyy, A. (2014). Insights into naturally minimised Streptomyces albus J1074 genome. BMC Genomics, 15, 97. https://doi.org/10.1186/1471-2164-15-97.
Ishizuka, H., Horinouchi, S., Kieser, H. M., Hopwood, D. A. and Beppu, T.(1992). A putative two-component regulatory system involved in secondary metabolism in Streptomyces spp. Journal of Bacteriology, 174, 7585-7594, 23.
Shu, D., Chen, L., Wang, W., Yu, Z., Ren, C., Zhang, W., & Yang, S. (2009). afsQ1-Q2-sigQ is a pleiotropic but conditionally required signal transduction system for both secondary metabolism and morphological development in Streptomyces coelicolor. Applied Microbiology and Biotechnology, 81(6), 1149–1160.
Rui, W., Yvonne, M., Jin, W., Weiwen, Z., Guoping, Z., Wolfgang, W., Yinhua, L., & Weihong, J. (2013). Identification of two-component system AfsQ1/Q2 regulon and its cross-regulation with GlnR in Streptomyces coelicolor. Molecular Microbiology, 87, 30–48.
Chen, S., Zheng, G., Zhu, H., He, H., Chen, L., Zhang, W., Jiang, W., & Lu, Y. (2016). Roles of two-component system AfsQ1/Q2 in regulating biosynthesis of the yellow-pigmented coelimycin P2 in Streptomyces coelicolor. FEMS Microbiology Letters, 363, fnw160.
Shuai, L., Di, S., Jianya, Z., Zhi, C., Ying, W., & Jilun, L. (2014). An extracytoplasmic function sigma factor, σ(25), differentially regulates avermectin and oligomycin biosynthesis in Streptomyces avermitilis. Applied Microbiology and Biotechnology, 98, 7097–7112.
Zhang, K., Mohsin, A., Dai, Y., Chen, Z., Zhuang, Y., Chu, J. and Guo, M. (2019).Combinatorial effect of ARTP mutagenesis and ribosome engineering on an industrial strain of Streptomyces albus S12 for enhanced biosynthesis of salinomycin. Frontiers in Bioengieering and Biotechnology, pp. 212.
Bierman, M., Logan, R., O’Brien, K., Seno, E. T., Rao, R. N., & Schoner, B. E. (1992). Plasmid cloning vectors for the conjugal transfer of DNA from Escherichia coli to Streptomyces spp. Gene, 116(1), 43–49.
Livak, K. J., & Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods, 25(4), 402–408.
You, D., Wang, M. M., & Ye, B. C. (2017). Acetyl-CoA synthetases of Saccharopolyspora erythraea are regulated by the nitrogen response regulator GlnR at both transcriptional and post-translational levels. Molecular Microbiology, 103(5), 845–859.
Lu, C., Zhang, X., Jiang, M., & Bai, L. (2016). Enhanced salinomycin production by adjusting the supply of polyketide extender units in Streptomyces albus. Metabolic Engineering, 35, 129–137.
Zhang, X., Lu, C., & Bai, L. (2017). Mechanism of salinomycin overproduction in Streptomyces albus as revealed by comparative functional genomics. Applied Microbiology and Biotechnology, 101(11), 4635–4644.
Mobley, H. L., Island, M. D., & Hausinger, R. P. (1995). Molecular biology of microbial ureases. Microbiology and Molecular Biology Reviews, 59, 451–480.
Mobley, H. L., & Hausinger, R. P. (1989). Microbial ureases: significance, regulation, and molecular characterization. Microbiological Reviews, 53(1), 85–108.
Zhao, J., Zhu, L., Fan, C., Wu, Y., & Xiang, S. (2017). Structure and function of urea amidolyase. Bioscience Reports, 38, BSR20171617.
Rodríguez, H., Rico, S., Díaz, M., & Santamaría, R. I. (2013). Two-component systems in Streptomyces: Key regulators of antibiotic complex pathways. Microbial Cell Factories, 12(1), 127.
Apel, A. K., Alberto, S. L., Antonio, R. G., & Martín, J. F. (2007). Phosphate control of phoA, phoC and phoD gene expression in Streptomyces coelicolor reveals significant differences in binding of PhoP to their promoter regions. Microbiology, 153(10), 3527–3537.
Zhenyu, Y., Hong, Z., Fujun, D., Weiwen, Z., Zhongjun, Q., Sheng, Y., Huarong, T., Yinhua, L., & Weihong, J. (2012). Differential regulation of antibiotic biosynthesis by DraR-K, a novel two-component system in Streptomyces coelicolor. Molecular Microbiology, 85, 535–556.
Hee-Jeon, H., Paget, M. S. B., & Buttner, M. J. (2008). A signal transduction system in Streptomyces coelicolor that activates the expression of a putative cell wall glycan operon in response to vancomycin and other cell wall-specific antibiotics. Molecular Microbiology, 69, 1199–1211.
Acknowledgments
We thank Zhejiang Biok Biology Co., Ltd. for providing strains and other experimental help.
Funding
This research was supported by the National Natural Science Foundation of China (81373286) and Fundamental Research Funds for the China Central Universities (No.22221818014 and No.22221817014) and 111 Project (B18022) to Meijin Guo.
Author information
Authors and Affiliations
Contributions
All authors saw and approved the manuscript. All authors contributed significantly to the work. Meijin Guo and Kuipu Zhang conceived the project. Meijin Guo, Kuipu Zhang designed experiments and analyzed results. Kuipu Zhang wrote the manuscript with the help of Ali Mohsin, Muhammad Fahad Ali, Meijin Guo, Yingping Zhuang, and Ju Chu. Kuipu Zhang performed experiments, supported by Yichen Dai, Zhongbing Chen.
Corresponding author
Ethics declarations
Conflict of Interest
Author Zhongbing Chen was employed by the company Zhejiang Biok Biology Co., Ltd. All other authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The presenting author of this manuscript in ACB2019 is Kuipu Zhang.
Electronic Supplementary Materials
ESM 1
(DOCX 40 kb)
Rights and permissions
About this article
Cite this article
Zhang, K., Mohsin, A., Dai, Y. et al. Role of a Two-Component Signal Transduction System RspA1/A2 in Regulating the Biosynthesis of Salinomycin in Streptomyces albus. Appl Biochem Biotechnol 193, 1296–1310 (2021). https://doi.org/10.1007/s12010-020-03357-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-020-03357-z