Abstract
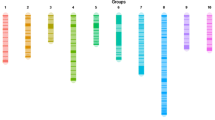
Auricularia auricula-judae is a traditional edible fungus that is cultivated widely in China. In this study, a genetic linkage map for A. auricula-judae was constructed using a mapping population consisting of 138 monokaryons derived from a hybrid strain (A119-5). The monokaryotic parent strains A14-5 and A18-119 were derived from two cultivated varieties, A14 (Qihei No. 1) and A18 (Qihei No. 2), respectively. In total, 130 simple sequence repeat markers were mapped. These markers were developed using the whole genome sequence of A. auricula-judae and amplified in A14-5, A18- 119, and the mapping population. The map consisted of 11 linkage groups (LGs) spanning 854 cM, with an average interval length of 6.57 cM. A testcross population was derived from crossing between the monokaryon A184-57 (from the wild strain A184 as a tester strain) and the mapping population. Important agronomic trait-related QTLs, including mycelium growth rate on potato dextrose agar for the mapping population, mycelium growth rate on potato dextrose agar and sawdust for the testcross population, growth period (days from inoculation to fruiting body harvesting), and yield for the testcross population, were identified using the composite interval mapping method. Six mycelium growth raterelated QTLs were identified on LG1 and LG4, two growth period-related QTLs were identified on LG2, and three yieldrelated QTLs were identified on LG2 and LG6. The results showed no linkage relationship between mycelium growth rate and growth period. The present study provides a foundation for locating genes for important agronomic characteristics in A. auricula-judae in the future.
Similar content being viewed by others
References
Au, C.H., Man, K.C., Man, C.W., Chu, A.K.K., Law, P.T.W., and Kwan, H.S. 2013. Rapid genotyping by low-coverage resequencing to construct genetic linkage maps of fungi: a case study in Lentinula edodes. BMC Res. Notes 6, 307.
Chen, Y., Yao, F.J., Zhang, Y.M., Chen, L., Cong, S., and Fang, M. 2014a. A new Auricularia auricula cultivar‘Jihei 3’. Acta Hortic. Sinica 41, 1751–1752.
Chen, Y., Yao, F.J., Zhang, Y.M., and Fang, M. 2014b. Numerical classification of cultivated germplasm of Auricularia auricula-judae. Mycosystema 33, 984–996.
Du, P., Cui, B.K., Zhang, C.F., and Dai, Y.C. 2013. Genetic diversity of wild Auricularia auricula-judae revealed by ISSR analysis. Biochem. Syst. Ecol. 48, 199–205.
Dutech, C., Enjalbert, J., Fournier, E., Delmotte, F., Barrès, B., Carlier, J., Tharreau, D., Giraud, T. 2007. Challenges of microsatellite isolation in fungi. Fungal Genet. Biol. 44, 933–949.
Fan, X.Z., Zhou, Y., Xiao, Y., and Bian, Y.B. 2013. Cloning and characterization of two allelic glyceraldehyde-3-phosphate dehydrogenase genes in Auricularia auricula-judae. World J. Microbiol. Biotechnol. 30, 181–189.
Fan, X.Z., Zhou, Y., Xiao, Y., Xu, Z., and Bian, Y.B. 2014. Cloning, expression and phylogenetic analysis of a divergent laccase multigene family in Auricularia auricula-judae. Microbiol. Res. 169, 453–462.
Fang, M., Yao, F.J., Wang, X., Chen, Y., Zhang, Y.M., and Ren, Y.Y. 2013. A new Auricularia auricula cultivar‘Jihei 2’. Acta Hortic. Sinica 40, 1215–1216.
Floudas, D., Binder, M., Riley, R., Barry, K., Blanchette, R.A., Henrissat, B., Martínez, A.T., Otillar, R., Spatafora, J.W., Yadav., J.S., et al. 2012. The paleozoic origin of enzymatic lignin decomposition reconstructed from 31 fungal genomes. Science 336, 1715–1719.
Foulongne-Oriol, M. 2012. Genetic linkage mapping in fungi: current state, applications, and future trends. Appl. Microbiol. Biotechnol. 95, 891–904.
Foulongne-Oriol, M., Rocha de Brito, M., Cabannes, D., Clément, A., Spataro, C., Moinard, M., Souza Dias, E., Callac, P., and Savoie, J. 2016. The genetic linkage map of the medicinal mushroom Agaricus subrufescens reveals highly conserved macrosynteny with the congeneric species Agaricus bisporus. 3G-Genes Genom. Genet. 11, 1217–1226.
Foulongne-Oriol, M., Spataro, C., Cathalot, V., Monllor, S., and Savoie, J.M. 2010. An expanded genetic linkage map of an intervarietal Agaricus bisporus var. bisporus × A. bisporus var. burnettii hybrid based on AFLP, SSR and CAPS markers sheds light on the recombination behaviour of the species. Fungal Genet. Biol. 47, 226–236.
Gong, W.B., Liu, W., Lu, Y.Y., Bian, Y.B., Yan, Z., Kwan, H.S., Cheung, M.K., and Xiao, Y. 2014a. Constructing a new integrated genetic linkage map and mapping quantitative trait loci for vegetative mycelium growth rate in Lentinula edodes. Fungal Biol. 118, 295–308.
Gong, W.B., Xu, R., Xiao, Y., Zhou, Y., and Bian, Y.B. 2014b. Phenotypic evaluation and analysis of important agronomic traits in the hybrid and natural populations of Lentinula edode. Sci. Hortic. 179, 271–276.
Im, C.H., Park, Y.H., Hammel, K.E., Park, B., Kwon, S.W., Ryu, H., and Ryu, J.S. 2016. Construction of a genetic linkage map and analysis of quantitative trait loci associated with the agronomically important traits of Pleurotus eryngii. Fungal Genet. Biol. 92, 50–64.
Joshi, C.P. and Chiang, V.L. 1998. Conserved sequence motifs in plant S-adenosyl-L-methionine-dependent methyltransferases. Plant Mol. Biol. 37, 663–674.
Kerrigan, R.W., Royer, J.C., Baller, L.M., Kohli, Y., Horgen, P.A., and Anderson, J.B. 1993. Meiotic behavior and linkage relationships in the secondarily homothallic fungus Agaricus bisporus. Genet. 133, 225–236.
Khaskheli, S.G., Zheng, W., Sheikh, S.A., Khaskheli, A.A., Liu, Y., Soomro, A.H., Feng, X., Sauer, M.B., Wang, Y.F., and Huang, W. 2015. Characterization of Auricularia auricula polysaccharides and its antioxidant properties in fresh and pickled product. Int. J. Biol. Macromol. 81, 387–395.
Labbé, J. and Tacon, F.L. 2008. A genetic linkage map for the ectomycorrhizal fungus Laccaria bicolor and its alignment to the whole-genome sequence assemblies. New Phytol. 180, 316–328.
Larraya, L.M., Alfonso, M., Pisabarro, A.G., and Ramírez, L. 2003. Mapping of genomic regions (quantitative trait loci) controlling production and quality in industrial cultures of the edible basidiomycete Pleurotus ostreatus. Appl. Environ. Microbiol. 69, 3617–3625.
Larraya, L.M., Idareta, E., Arana, D., Ritter, E., Pisabarro, A.G., and Ramírez, L. 2002. Quantitative trait loci controlling vegetative growth rate in the edible basidiomycete Pleurotus ostreatus. Appl. Environ. Microbiol. 68, 1109–1114.
Larraya, L.M., Pérez, G., Ritter, E., Pisabarro, A.G., and Ramírez, L. 2000. Genetic linkage map of the edible basidiomycete Pleurotus ostreatus. Appl. Environ. Microbiol. 66, 5290–5300.
Li, Y. 2001. Wood ear of China, pp. 3–10. Changchun Press, Changchun, Jilin, China.
Li, L., Li, J., Zou, L., Bai, S.Y., Niu, L.M., and Ma, Y.K. 2007. RAPD analysis of genetic diversity of nine strains of Auricularia auricula cultivated in Heilongjiang Province. J. Forest. Res. 18, 136–138.
Li, H.M., Shang, X.D., Tan, Q., Pan, Y.J., and Chen, M.J. 2009. Genetic differentiation of self-crossing S1 generations of Pleurotus eryngii. Mycosystema 28, 541–547.
Liu, G.J. 2011. Master’s thesis. Jilin Agricultural University, Changchun, Jilin, China.
Lu, Y.P., Lian, L.D., Guo, L.X., Xie, B., Wang, W., Chen, B.Z., Peer, A.F., Li, S.J., Wu, T.J., and Xie, B.G. 2015. The accordant trend of both parameters (rgs expression and cAMP content) follows the pattern of development of fruiting body in Volvariella volvacea. Currt. Microbiol. 71, 1–6.
Miyazaki, K., Huang, F., Zhang, B., Shiraishi, S., Sakai, M., Shimaya, C., and Kazuo, S. 2008. Genetic map of a basidiomycete fungus, Lentinula edodes (shiitake mushroom), constructed by tetrad analysis. Breed. Sci. 58, 23–30.
Murray, M.G. and Thompson, W.F. 1980. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 8, 4321–4326.
Okuda, Y., Ueda, J., Obatake, Y., Murakami, S., Fukumasa, Y., and Matsumoto, T. 2012. Construction of a genetic linkage map based on amplified fragment length polymorphism markers and development of sequence-tagged site markers for marker-assisted selection of the sporeless trait in the oyster mushroom (Pleurotus eryngii). Appl. Environ. Microbiol. 78, 1496–1504.
Reza, A., Choi, M.J., Damte, D., Jo, W.S., Lee, S.J., Lee, J.S., and Park, S.C. 2011. Comparative antitumor activity of different solvent fractions from an Auricularia auricula-judae ethanol extract in P388D1 and sarcoma 180 cells. Toxicol. Res. 27, 77–83.
Rozen, S. and Skaletsky, H. 2000. Primer 3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 132, 365–386.
Salmones, D., Gaitán-Hernández, R., Pérez, R., and Guzmán, G. 1997. Studies on genus Pleurotus. VIII. Interaction between mycelial growth and yield. Rev. Iberoam. Micol. 14, 173–176.
Sivolapova, A.B., Shnyreva, A.V., Sonnenberg, A., and Baars, I. 2012. DNA marking of some quantitative trait loci in the cultivated edible mushroom Pleurotus ostreatus (Fr.) Kumm. Russ. J. Genet. 48, 383–389.
Sun, S.J., Zhang, X.J., Chen, W.X., Zhang, L.Y., and Zhu, H. 2016. Production of natural edible melanin by Auricularia auricula and its physicochemical properties. Food Chem. 196, 486–492.
Terashima, K., Matsumoto, T., Hayashi, E., and Fukumasa-Nakai, Y. 2002. A genetic linkage map of Lentinula edodes (shiitake) based on AFLP markers. Mycol. Res. 106, 911–917.
Van Ooijen, J.W. 2006. JoinMap 4.0: Software for the calculation of genetic Linkage maps in experimental populations. Kyazma BV, Wageningen, Netherlands.
Wang, X.E. 2013. Ph. D. thesis. Jilin Agricultural University, Changchun, Jilin, China.
Wang, S., Basten, C., and Zeng, Z.B. 2012. Windows QTL cartographer 2.5. Raleigh, NC: Department of Statistics, North Carolina State University. USA.
Wang, X.E., Zhang, Y.M., Chen, Y., Fang, M., Wang, W., and Yao, F.J. 2013. A new Auricularia auricula cultivar‘Jihei 1’. Acta Hortic. Sinica 40, 601–602.
Wieland, T. and Chen, C.K. 1999. Regulators of G-protein signaling: a novel protein family involved in timely deactivation and desensitization of signalling via heterotrimeric G proteins. Naunyn-Schmiedeberg’s Arch. Pharmacol. 360, 14–26.
Wu, F., Yuan, Y., Malysheva, V.F., Du, P., and Dai, Y.C. 2014. Species clarification of the most important and cultivated Auricularia mushroom “Heimuer”: evidence from morphological and molecular data. Phytotaxa 186, 241–253.
Xiang, X.J., Li, C., Li, L., Bian, Y.B., Kwan, H.S., Nong, W.Y., Cheung, M.K., and Xiao, Y. 2016. Genetic diversity and population structure of Chinese Lentinula edodes revealed by InDel and SSR markers. Mycol. Prog. 15, 37.
Yao, F.J. and Bian, Y.B. 2011. Graphic illustration on key cultivation techniques of Auricularia auricula-judae, pp. 1–78. China Agriculture Press, Beijing, China.
Yao, F.J., Zhang, Y.M., and Li, Y. 2005. Analysis of the gene linkage of Pleurotus citrinopileatus by use of auxotrophic mutation. Mycosystema 24, 36–41.
Yao, F.J., Zhang, Y.M., Lu, L.X., and Fang, M. 2015. Research progress on genetics and breeding of Auricularia auricula-judae. J. Fungal Res. 13, 125–128.
Zeng, Z.B. 1994. Precision mapping of quantitative trait loci. Genetics 136, 1457–1468.
Zhang, P. 2011. Master’s thesis. Jilin Agricultural University, Changchun, Jilin, China.
Zhang, R.Y., Hu, D.D., Gu, J.G., Hu, Q.X., Zuo, X.M., and Wang, H.X. 2012. Development of SSR markers for typing cultivars in the mushroom Auricularia auricula-judae. Mycol. Prog. 11, 587–592.
Zhang, R.Y., Hu, D.D., Zuo, X.M., and Hu, Q.X. 2010. Strategies to develop SSR markers in edible mushrooms: a review. J. Fungal Res. 8, 239–244.
Zhang, Y., Yao, A.Y., and Song, K.Y. 2016. Torrefaction of cultivation residue of Auricularia auricula-judae to obtain biochar with enhanced fuel properties. Bioresour. Technol. 206, 211–216.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplemental material for this article may be found at http://www.springerlink.com/content/120956.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Lu, LX., Yao, FJ., Wang, P. et al. Construction of a genetic linkage map and QTL mapping of agronomic traits in Auricularia auricula-judae . J Microbiol. 55, 792–799 (2017). https://doi.org/10.1007/s12275-017-7241-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-017-7241-6