Abstract
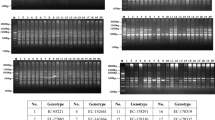
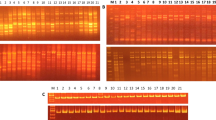
A total of 24 pigeonpea (Cajanus cajan L. Millspaugh) cultivars representing different maturity groups were evaluated for genetic diversity analysis using 10 pigeonpea specific and 66 cross-genera microsatellite markers. Of the cross-genera microsatellite markers, only 12 showed amplification. A total of 45 alleles were amplified by the 22 markers. Nine markers showed 100 % polymorphism. Markers Lc 14, BMd 48 and CCB 9 amplified maximum number (5) of alleles each. One genotype specific unique band in Pusa 9 was generated by markers CCB 8. Maximum genetic diversity (74 %) was observed between cultivars MA 3 and CO 6, while the minimum diversity (12 %) was observed between NDA 1 and DA 11. The average diversity among the cultivars was estimated to be 45.6 %. SSR primers from pigeonpea were found to be more polymorphic (37 %) as compared to common bean and lentil markers. The arithmetic mean heterozygosity (Hav) and marker index (MI) were found to be 0.014 and 0.03, respectively, indicating the potential of common bean and lentil microsatellite markers for genetic mapping, diversity analysis and genotyping in Cajanus.
Similar content being viewed by others
References
Abdelnoor RV, Barros EG and Moreira MA (1995). Determination of genetic diversity within Brazilian soybean germplasm using random amplified DNA amplification techniques and comparative analysis with pedigree data. Braz. J. Genet. 18: 265–273
Anderson JA, Churchill GA, Autrique JE, Sollers ME and Tanksley SD (1993). Optimizing parental selection for genetic linkage maps. Genome 36: 181–186
Barrett BA, Kidwell KK and Fox PN (1998). Comparison of AFLP and pedigree based genetic diversity assessment methods using wheat cultivars from the Pacific Northwest. Crop Sci. 38: 1271–1278
Blair MW, Pedraza F, Buendia HF, Gaitan-Solis E, Beebe SE, Gepts P and Tohme J (2003). Development of a genome-wide anchored microsatellite map for common bean (Phaseolus vulgaris). Theor. Appl. Genet. 107: 1362–1374
Burns MJ, Edwards KJ, Newburg HJ, Ford Lloyd BV and Baggott CD (2001). Development of simple sequence repeat (SSR) markers for the assessment of gene flow and genetic diversity in pigeonpea (Cajanus cajan). Mol. Ecol. Notes 4: 283–285
Choi HK, Mun JH, Kin DJ, Zhu H, Baek JM, Mudjge J, Roe B, Ellis N, Doyle J, Kiss GB, Young ND and Cook DR (2004). Estimating genome conservation between crop and model legume species. Proc. Natl. Acad. Sci. 101: 15289–15294
Choudhary S, Sethy NK, Shokeen B and Bhatia S (2009). Development of chickpea EST-SSR markers and analysis of allelic variation across related species. Theor. Appl. Genet. 118: 591–608
Choumane W, Winter P, Weigand F and Kahl G (2004). Conservation of microsatellite flanking sequences in different taxa of leguminosae. Euphytica 138: 239–245
Datta S, Kaashyap M and Kumar S (2009). Amplification of chickpea-specific SSR primers in Cajanus species and their validity in diversity analysis. Plant Breed. doi 10.1111/j.1439-0523.2009.01678.x
DeWoody JA, Honeycutt RL and Skow LC (1995). Microsatellite markers in white-tailed deer. J. Hered. 86: 317–319
Doldi ML, Vollmann J and Lelley T (1997). Genetic diversity in soybean as determined by RAPD and microsatellite analysis. Plant Breed. 116: 331–335
Ford R, Le Roux K, Itman C, Brouwer JB and Taylor PWJ (2002). Diversity analysis and genotyping in Pisum with sequence tagged microsatellite site (STMS) primers. Euphytica 124: 397–405
Gupta SC, Reddy LJ, Sharma D, Green JM, Murthi AN and Saxena KB (1980). Maintenance of pigeonpea cultivars. In: Proceedings of International Workshop on Pigeonpea, (Eds. Nene Y.L.), Vol 2, ICRISAT, Patancheru, India, pp 295–301.
Gutierrez MV, Vaz Patto MC, Huguet T, Cubero JI, Moreno MT and Torres AM (2005). Cross-species amplification of Medicago truncatula microsatellites across three major pulse crops. Theor. Appl. Genet. 110: 1210–1217
Hamwieh A, Udupa SM, Choumane W, Sarker A, Dreyer F, Jung C and Baum M (2005). A genetic linkage map of Lens sp. based on microsatellite and AFLP markers and the localization of fusarium vascular wilt resistance. Theor. Appl. Genet. 110: 669–677
Hamwieh A, Udupa SM, Sarker A, Jung C and Baum M (2009). Development of new microsatellite markers and their application in the analysis of genetic diversity in lentils. Breed. Sci. 59: 77–86
Jaccard P (1908), Nouvelle recherches sur La distribution florale. Bulletin Bull. Soc. Vaud. Sci. Nat. 44: 223–270
Kumar S and Ali M (2006). GE interaction and its breeding implications in pulses. The Botanica 56: 31–36
L’taief B, Horres R, Jungmann R, Molina C, Sifi B, Lachaal M, Winter P and Kahl G (2008). Locus-specific microsatellite markers in common bean (Phaseolus vulgaris L.) isolation and characterization. Euphytica 162: 301–310
Nadimpalli RG, Jarret RL, Phatak SC and Kochert G (1992). Phylogenetic relationships of the pigeonpea (Cajanus cajan) based on nuclear restriction fragment length polymorphisms. Genome 36: 216–223
Nei M and Li WH (1979). Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. 79: 5269–5273
Odeny DA, Jayashree B, Ferguson M, Hoisngton D, Crouch J and Gebhardt C (2007). Development, characterization and utilization of microsatellite markers in pigeonpea. Plant Breed. 126:130–136
Odeny DA, Jayashree B, Gebhardt C and Crouch J (2009). New microsatellite markers for pigeonpea (Cajanus cajan (L) millsp.) BMC Res. Notes 2: 35. doi:10.1186/1756-0500-2-35
Pandian A, Ford R and Paul WJ (2000). Transferability of sequence tagged microsatellite site (STMS) primers across four major pulses. Plant Mol. Biol. Rep. 18:395a–395h
Panguluri SK, Janaiah K, Govil JN, Kumar PA and Sharma PC (2006). AFLP fingerprinting in pigeonpea (Cajanus cajan (L.) Millsp.) and its wild relatives. Genet. Res. Crop. Evol. 53: 523–531
Peakall R, Gilmore S, Keys W, Morgante M and Rafalski A (1998). Cross-species amplification of soybean (Glycine max) simple sequence repeats (SSRs) within the genus and other legume genera; Implications for the transferability of SSRs in plants. Mol. Biol. Evol. 15: 1275–1287
Powell W, Morgante M, Andre C, Hanafey V, Vogel T, Tingey S and Rafalski JA (1996). The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol. Breed. 3: 225–238
Roa A, P Chavarriaga-Agguirre, Duque MC, Maya MM, Bonier-bale MW, Iglesias C and Tohme J (2000). Crosss-pecies amplification of cassava (Manihot esculenta) (Euphorbiaceae) microsatellite: allelic polymorphism and degree of relationship. Am. J. of Bot. 87: 1647–1655
Ratnaparkhe MB, Gupta VS, Ven Meurthy MR and Ranjekar PK (1995). Genetic fingerprinting of Pigeonpea (Cajanus cajan L. Millsp) and wild relatives using RAPD markers. Theor. Appl. Genet. 91: 893–898
Rohlf FJ (1998). NTSYS-PC Numerical taxonomy and Multivariate analysis system Version 2.1, Exeter Software, Applied Biostatistics, New York, USA
Sethy NK, Shokeen B, Edwards KJ and Bhatia S (2006). Development of microsatellite markers and analysis of intraspecific genetic variability in chickpea (Cicer arietinum L.). Theor. Appl. Genet. 112: 1416–1428
Singh F, Kumar S and Majumder ND (2006). Genetic base of pigeonpea varieties released in India. Indian J. Pulses Res. 19: 179–183
Souframanien J, Manjaya JG, Krishna TG and Pawar ME (2003). Random amplified polymorphic DNA analyses of cytoplasmic male sterile and male fertile pigeon pea (Cajanus cajan (L.) Millsp). Euphytica 129: 293–299
Stepien L, Mohler V, Bocianowski and Koczyk G (2004). Assessing genetic diversity of Polish wheat (Triticum aestivum) varieties using microsatellite markers. Genet. Res. Crop. Evol. 54: 1499–1506
Tautz D (1989). Hyper variability of simple sequences as a general source of polymorphism DNA markers. Nucl. Ac. Res. 17: 6463–6471
Tautz D and Renz M (1984). Simple sequences are ubiquitous repetitive components of eukaryotic genomes. Nucl. Ac. Res. 12: 4127–4137
Upadhyaya HD, Reddy KN, Gowda CLL and Singh S (2007). Phenotypic diversity in the pigeonpea core collection. Genet. Resour. Crop. Evol. 54: 1167–1184
Varshney RK, Garner A and Sorell ME (2005). Genic microsatellite markers in plants: Features and application. Trends Biotech. 23: 48–55
Wilde J, Waugh JR and Powell W (1992). Genetic fingerprinting of Theobroma clones using RAPD markers. Theor. Appl. Genet. 83:871–877
Zeng L, Kwon TR, Liu X, Wilson C, Grieve CM and Gregorio GB (2006) Genetic diversity analyzed by microsatellite markers among rice genotypes with different adaptation to saline soil. Plant Sci. 166: 1275–1285
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Datta, S., Mahfooz, S., Singh, P. et al. Cross-genera amplification of informative microsatellite markers from common bean and lentil for the assessment of genetic diversity in pigeonpea. Physiol Mol Biol Plants 16, 123–134 (2010). https://doi.org/10.1007/s12298-010-0014-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-010-0014-x