Abstract
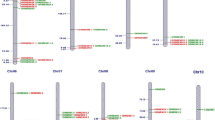
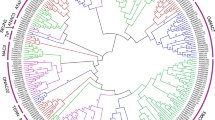
The plant-specific NAC transcription factor (TFs) plays crucial role in plant growth as well as in stress resistance. In the present study, 87 Zea mays NAC TFs were obtained from the transcriptome analysis using drought-resistant maize inbred line Y882 as experimental material under PEG stress and rewatering treatment. Comprehensive analyses were conducted including genes structure, chromosomal localization, phylogenetic tree and motif prediction, cis-elements and expression patterns. The results showed that the 87 ZmNAC genes distributed on 10 chromosomes and were categorized into 15 groups based on their conserved gene structure and motifs. Phylogenetic tree analysis was also constructed referencing to the counterparts of Arabidopsis and rice, and the stress-related cis-elements in the promoter region were also analyzed. 87 ZmNAC genes exhibited different expression levels at 3 treatment points, indicating different response to drought stress. This genome-wide analysis of 87 ZmNAC genes will provide basis for further gene function detection.









Similar content being viewed by others
References
Borrill P, Harrington SA, Uauy C (2017) Genome-wide sequence and expression analysis of the NAC transcription factor family in polyploid wheat. G3 Genes Genomes Genet 7:3019–3029. https://doi.org/10.1534/g3.117.043679
Bruce WB, Edmeades GO, Barker TC (2002) Molecular and physiological approaches to maize improvement for drought tolerance. J Exp Bot 53:13–25. https://doi.org/10.1093/jexbot/53.366.13
Bu LD, Zhang RH, Han MM, Xue JQ, Chang Y (2009) The physiological mechanism of compensation effect in maize leaf by rewatering after draught stress. Acta Agric Boreal Occident Sin 18:88–92. https://doi.org/10.3969/j.issn.1004-1389.2009.02.020
Capella M, Re DA, Arce AL, Chan RL (2014) Plant homeodomain-leucine zipper I transcription factors exhibit different functional AHA motifs that selectively interact with TBP or/and TFIIB. Plant Cell Rep 33:955–967. https://doi.org/10.1007/s00299-014-1576-9
Dung Tien L, Nishiyama R, Watanabe Y, Mochida K, Yamaguchi-Shinozaki K, Shinozaki K, Lam-Son Phan T (2011) Genome-wide survey and expression analysis of the plant-specific NAC transcription factor family in soybean during development and dehydration stress. DNA Res 18:263–276. https://doi.org/10.1093/dnares/dsr015
Fang Y, Liao K, Du H, Xu Y, Song H, Li X, Xiong L (2015) A stress-responsive NAC transcription factor SNAC3 confers heat and drought tolerance through modulation of reactive oxygen species in rice. J Exp Bot 66:6803–6817. https://doi.org/10.1093/jxb/erv386
Ge SS, Tang GY, Bi YP, Liu ZJ (2015) Genome-wide identification and analysis of NAC gene family in maize. Shandong Agric Sci 47:1–6. https://doi.org/10.14083/j.issn.1001-4942.2015.02.001
Gong X, Zhao L, Song X, Lin Z, Gu B, Yan J, Zhang S, Tao S, Huang X (2019) Genome-wide analyses and expression patterns under abiotic stress of NAC transcription factors in white pear (Pyrus bretschneideri). BMC Plant Biol. https://doi.org/10.1186/s12870-019-1760-8
Kadier Y, Zu Y-y, Dai Q-m, Song G, Lin S-w, Sun Q-p, Pan J-b, Lu M (2017) Genome-wide identification, classification and expression analysis of NAC family of genes in sorghum Sorghum bicolor (L.) Moench. Plant Growth Regul 83:301–312. https://doi.org/10.1007/s10725-017-0295-y
Kamoshita A, Rodriguez R, Yamauchi A, Wade LJ (2004) Genotypic variation in response of rainfed lowland rice to prolonged drought and rewatering. Plant Prod Sci 7:406–420. https://doi.org/10.1626/pps.7.406
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL (2013) TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. https://doi.org/10.1186/gb-2013-14-4-r36
Kou X, Wang S, Wu M, Guo R, Xue Z, Meng N, Tao X, Chen M, Zhang Y (2014) Molecular characterization and expression analysis of NAC family transcription factors in tomato. Plant Mol Biol Rep 32:501–516. https://doi.org/10.1007/s11105-013-0655-3
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359. https://doi.org/10.1038/nmeth.1923
Liu Y, Sun J, Wu Y (2016) Arabidopsis ATAF1 enhances the tolerance to salt stress and ABA in transgenic rice. J Plant Res 129:955–962. https://doi.org/10.1007/s10265-016-0833-0
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta C(T)) method. Methods (San Diego, Calif) 25:402–408. https://doi.org/10.1006/meth.2001.1262
Lu M, Ying S, Zhang D-F, Shi Y-S, Song Y-C, Wang T-Y, Li Y (2012) A maize stress-responsive NAC transcription factor, ZmSNAC1, confers enhanced tolerance to dehydration in transgenic Arabidopsis. Plant Cell Rep 31:1701–1711. https://doi.org/10.1007/s00299-012-1284-2
Mao H, Yu L, Han R, Li Z, Liu H (2016) ZmNAC55, a maize stress-responsive NAC transcription factor, confers drought resistance in transgenic Arabidopsis. Plant Physiol Biochem 105:55–66. https://doi.org/10.1016/j.plaphy.2016.04.018
Nakashima K, Takasaki H, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) NAC transcription factors in plant abiotic stress responses. Biochim Biophys Acta Gene Regul Mech 1819:97–103. https://doi.org/10.1016/j.bbagrm.2011.10.005
Nuruzzaman M, Manimekalai R, Sharoni AM, Satoh K, Kondoh H, Ooka H, Kikuchi S (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465:30–44. https://doi.org/10.1016/j.gene.2010.06.008
Peng X, Zhao Y, Li X, Wu M, Chai W, Sheng L, Wang Y, Dong Q, Jiang H, Cheng B (2015) Genomewide identification, classification and analysis of NAC type gene family in maize. J Genet 94:377–390. https://doi.org/10.1007/s12041-015-0526-9
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17:369–381. https://doi.org/10.1016/j.tplants.2012.02.004
Saidi MN, Mergby D, Brini F (2017) Identification and expression analysis of the NAC transcription factor family in durum wheat (Triticum turgidum L. ssp. durum). Plant Physiol Biochem 112:117–128. https://doi.org/10.1016/j.plaphy.2016.12.028
Schwartz S, Meshorer E, Ast G (2009) Chromatin organization marks exon-intron structure. Nat Struct Mol Biol 16:990–995. https://doi.org/10.1038/nsmb.1659
Shan L (2003) Issues of science and technology on water saving agricultural development in China. Agric Res Arid Areas. https://doi.org/10.3321/j.issn:1000-7601.2003.01.001
Shang H, Wang Z, Zou C, Zhang Z, Li W, Li J, Shi Y, Gong W, Chen T, Liu A, Gong J, Ge Q, Yuan Y (2016) Comprehensive analysis of NAC transcription factors in diploid Gossypium: sequence conservation and expression analysis uncover their roles during fiber development. Sci China Life Sci 59:142–153. https://doi.org/10.1007/s11427-016-5001-1
Shiriga K, Sharma R, Kumar K, Yadav SK, Hossain F, Thirunavukkarasu N (2014) Genome-wide identification and expression pattern of drought-responsive members of the NAC family in maize. Meta Gene 2:407–417. https://doi.org/10.1016/j.mgene.2014.05.001
So H-A, Lee J-H (2019) NAC transcription factors from soybean (Glycine max L.) differentially regulated by abiotic stress. J Plant Biol 62:147–160. https://doi.org/10.1007/s12374-018-0285-2
Sun H, Hu M, Li J, Chen L, Li M, Zhang S, Zhang X, Yang X (2018) Comprehensive analysis of NAC transcription factors uncovers their roles during fiber development and stress response in cotton. BMC Plant Biol. https://doi.org/10.1186/s12870-018-1367-5
Tran LSP, Nakashima K, Sakuma Y, Simpson SD, Fujita Y, Maruyama K, Fujita M, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2004) Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought-responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell 16:2481–2498. https://doi.org/10.1105/tpc.104.022699
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7:562–578. https://doi.org/10.1038/nprot.2012.016
Wang Z, Dane F (2013) NAC (NAM/ATAF/CUC) transcription factors in different stresses and their signaling pathway. Acta Physiol Plant 35:1397–1408. https://doi.org/10.1007/s11738-012-1195-4
Wu Y, Deng Z, Lai J, Zhang Y, Yang C, Yin B, Zhao Q, Zhang L, Li Y, Yang C, Xie Q (2009) Dual function of Arabidopsis ATAF1 in abiotic and biotic stress responses. Cell Res 19:1279–1290. https://doi.org/10.1038/cr.2009.108
Xu Z, Gongbuzhaxi Wang C, Xue F, Zhang H, Ji W (2015) Wheat NAC transcription factor TaNAC29 is involved in response to salt stress. Plant Physiol Biochem 96:356–363. https://doi.org/10.1016/j.plaphy.2015.08.013
Xue G-P, Way HM, Richardson T, Drenth J, Joyce PA, McIntyre CL (2011) Overexpression of TaNAC69 leads to enhanced transcript levels of stress up-regulated genes and dehydration tolerance in bread wheat. Mol Plant 4:697–712. https://doi.org/10.1093/mp/ssr013
Yang X, Wang X, Ji L, Yi Z, Fu C, Ran J, Hu R, Zhou G (2015) Overexpression of a Miscanthus lutarioriparius NAC gene MlNAC5 confers enhanced drought and cold tolerance in Arabidopsis. Plant Cell Rep 34:943–958. https://doi.org/10.1007/s00299-015-1756-2
Acknowledgements
The manuscript “Genome-Wide Analysis of NAC Transcription Factor Family in Maize under Drought Stress and Rewatering” was supported by the National Natural Science Foundation of China (No. 31471452), and the National Key Research and Development Program of China (No. 2017YFD0301106).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, G., Yuan, Z., Zhang, P. et al. Genome-wide analysis of NAC transcription factor family in maize under drought stress and rewatering. Physiol Mol Biol Plants 26, 705–717 (2020). https://doi.org/10.1007/s12298-020-00770-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-020-00770-w