Abstract
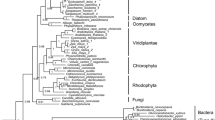
The coding product of alginate-c5-mannuronan-epimerase gene (algG gene) can catalyze the conversion of mannuronate to guluronate and determine the M/G ratio of alginate. Most of the current knowledge about genes involved in the alginate biosynthesis comes from bacterial systems. In this article, based on some algal and bacterial algG genes registered on GenBank and EMBL databases, we predicted 94 algG genes open reading frame (ORF) sequences of brown algae from the 1 000 Plant Transcriptome Sequencing Project (OneKP). By method of transcriptomic sequence analysis, gene structure and gene localization analysis, multiple sequence alignment and phylogenetic tree construction, we studied the algal algG gene family characteristics, the structure modeling and conserved motifs of AlgG protein, the origin of alginate biosynthesis and the variation incidents that might have happened during evolution in algae. Although there are different members in the algal algG gene family, almost all of them harbor the conserved epimerase region. Based on the phylogenetic analysis of algG genes, we proposed that brown algae acquired the alginate biosynthesis pathway from an ancient bacterium by horizontal gene transfer (HGT). Afterwards, followed by duplications, chromosome disorder, mutation or recombination during evolution, brown algal algG genes were divided into different types.
Similar content being viewed by others
References
Arnold K L, Bordoli J, Kopp, et al. 2006. The SWISS-MODEL Workspace: a web-base environment for protein structure homology modeling. Bioinformatics, 22: 195–201
Chitnis C E, Ohman D E. 1990. Cloning of Pseudomonas aeruginosa algG, which controls alginate structure. Bacteriol, 172(6): 2894–2900
Chitnis C E, Ohman D E. 1993. Genetic analysis of the alginate biosynthetic gene cluster of Pseudomonas aeruginosa shows evidence of an operonic structure. Mol Microbiol, 8: 583–590
Cook J M, Sterck L, Rouzé P, et al. 2010. The Ectocarpus genome and the independent evolution of multicellularity in brown algae. Nature, 465: 617–621
Douglas S E. 1998. Plastid evolution: origins, diversity, trends. Curr Opin Genet Dev, 8: 655–661
Ertesvåg H, Doseth B, Larsen B, et al. 1994. Cloning and expression of an Azotobacter vinelandii mannuronan C-5-epimerase gene. J Bacteriol, 176: 2846–2853
Ertesvåg H, Høidal H K, Hals I K, et al. 1995. A family of modular type mannuronan C-5-epimerase genes controls alginate structure in Azotobacter vinelandii. Mol Microbiol, 9: 719–731
Ertesvåg H, Høidal H K, Skjåk-Bræk G, et al. 1998. The Azotobacter vinelandii mannuronan C-5-epimerase AlgE1 consists of two separate catalytic domains. J Biol Chem, 273: 30927–30932
Ertesvåg H, Valla S. 1999. The A modules of the Azotobacter vinelandii mannuronan-C-5-epimerase AlgE1 are sufficient for both epimerization and binding of Ca2+. J Bacteriol, 181: 3033–3038
Flagel L, Wendel J. 2009. Gene duplication and evolutionary novelty in plants. New Phytologist, 183: 557–564
Franklin M J, Chitnis C E, Gacesa P, et al. 1994. Pseudomonas aeruginosa AlgG is a polymer level alginate C5-mannuronan epimerase. J Bacteriol, 176: 1821–1830
Gimmestad M, Sletta H, Ertesvag H, et al. 2003. The Pseudomonas fluorescens AlgG protein, but not itsmannuronan C-5-epimerase activity, is needed for alginate polymer formation. J Bacteriol, 185: 3515–3523
Gorin P A, Spencer J F. 1966. Exocellular alginic acid from Azotobacter vinelandii. Can J Chem, 44: 993–998
Govan J R W, Fyfe J A M, Jarman T R. 1981. Isolation of alginate producing mutants of Pseudomonas fluorescens, Pseudomonas putida and Pseudomonas mendocina. J Gen Microbiol, 125: 217–220
Guex N, Peitsch M C. 1997. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modelling. Electrophoresis, 18: 2714–2723
Gurvan M, Thierry T, Delphine S, et al. 2010. The cell wall polysaccharide metabolism of the brown alga Ectocarpus siliculosus. Insights into the evolution of extracellular matrix polysaccharides in Eukaryotes. New Phytologist, 188: 82–97
Rozeboom H J, Bjerkan T M, Kalk K H, et al. 2008. Structural and Mutational Characterization of the Catalytic A-module of the Mannuronan C-5-epimerase AlgE4 from Azotobacter vinelandii. J Biol Chem, 283: 23819–23828
Hughes T, Liberles D A. 2007. The pattern of evolution of smaller-scale gene duplicates in mammalian genomes is more consistent with neo-than subfunctionalisation. J Mol Evol, 65(5): 574–588
Jain S, Franklin M J, Ertesvag H, et al. 2003. The dual roles of AlgG in C-5-epimerization and secretion of alginate polymers in Pseudomonas aeruginosa. Mol Microbiol, 47: 1123–1133
Jain S, Ohman D E. 1998. Deletion of algK in mucoid Pseudomonas aeruginosa blocks alginate polymer formation and results in uronic acid secretion. J Bacteriol, 180: 634–641
Jenkins J, Shevchik V E, Hugouvieux-Cotte-Pattat N, et al. 2004. The crystal structure of pectate lyase Pel9A from Erwinia chrysanthemi. J Biol Chem, 279: 9139–9145
Kloareg B, Quatrano R S. 1988. Structure of the cell walls of marine algae and ecophysiological functions of the matrix polysaccharides. Oceanogr Mar Biol Annu Rev, 1988: 259–315
Lang B F, Seif E, Gray M W, et al. 1999. A comparative genomics approach to the evolution of eukaryotes and their mitochondria. J Eukaryot Microbiol, 46: 320–326
Linker A, Jones R S. 1966. A new polysaccharide resembling alginic acid isolated from pseudomonas. J Biol Chem, 241: 3845–385
Lynch M, Force A. 2000. The probability of duplicate gene preservation by subfunctionalization. Genetics, 154(1): 459–473
Morch Y A, Holtan S, Donati I, et al. 2008. Mechanical properties of C-5 epimerized alginates. Biomacromolecules, 9: 2360–2368
Moreira D, Le Guyader H, Philippe H. 2000. The origin of red algae and the evolution of chloroplasts. Nature, 405: 69–72
Nyvall P, Erwan C, Claire B, et al. 2003. Characterization of Mannuronan C-5-Epimerase Genes from the Brown Alga Laminaria digitata. Plant Physiology, 133(2): 726–735
Okasaki M, Furuya K, Tsukayama K, et al. 1982. Isolation and identification of alginic acid from a calcareous red alga Serraticardia maxima. Botanica Marina, 25: 123–131
Okasaki M, Shiroto C, Furuya K. 1984. Relationship between the location of polyuronides and calcification sites in the calcareous red algae Serraticardia maxima and Lithothamnion japonica (Rhodophyta, Corallinaceae). Jap J Phycol, 32: 364–372
Page R D. 1996. TREEVIEW: An application to display phylogenetic trees on personal computers. Computer Applications in the Biosciences, 12: 357–358
Panikkar R, Brasch D J. 1996. Composition and block structure of alginates from New Zealand brown seaweeds. Carbohydr Res, 293: 119–132
Pedersen S S, Hoiby N, Espersen F, et al. 1992. Role of alginate in infection with mucoid Pseudomonas aeruginosa in cystic fibrosis. Thorax, 47: 6–13
Penaloza-Vazquez A, Kidambi S P, Chakrabarty A M, et al. 1997. Characterization of the alginate biosynthetic gene cluster in Pseudomonas syringae pv. syringae. J Bacteriol, 179: 4464–4472
Posada D, Crandall K A. 1998. MODELTEST: testing the model of DNA substitution. Bioinformatics, 14: 817–818
Rehm B H A, Ertesvåg H, Valla S. 1996. A new Azotobacter vinelandii mannuronan C-5-epimerase gene (algG) is part of an alg gene cluster physically organized in a manner similar to that in Pseudomonas aeruginosa. Bacteriol, 178: 5884–5889
Rehm B H A, Valla S. 1997. Bacterial alginates: biosynthesis and applications. Appl Microbiol Biotechnol, 48: 281–288
Reyes-Prieto A, Weber A P, Bhattacharya D. 2007. The origin and establishment of the plastid in algae and plants. Annual Review of Genetics, 41: 147–168
Robles-Price A, Wong T Y, Sletta H, et al. 2004. AlgX is a periplasmic protein required for alginate biosynthesis in Pseudomonas aeruginosa. J Bacteriol, 186: 7369–7377
Ronquist F, Huelsenbeck J P. 2003. Mrbayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics, 19: 1572–1574
Sadoff H L. 1975. Encystment and germination of Azotobacter vinelandii. Bacteriol Rev, 39: 516–539
Schwede T, Kopp J, Guex N, et al. 2003. SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Research, 31: 3381–3385
Skjåk-Bræk G, Espevik T. 1996. Application of alginate gels in biotechnology and biomedicine. Carbohydr Eur, 14: 19–25
Smidsrød O, Draget K I. 1996. Chemistry and physical properties of alginates. Carbohydr Eur, 14: 6–13
Stephanie A, Douthit D, Mensur E O, et al. 2005. Epimerase Active Domain of Pseudomonas aeruginosa AlgG, a Protein That Contains a Right-Handed β-Helix. Journal of Bacteriology, 187(13): 4573–4583
Svanem B I, Skjåk-Bræk G, Ertesvåg H, et al. 1999. Cloning and expression of three new Azotobacter vinelandii genes closely related to a previously described gene family encoding mannuronan C-5-epimerases. Bacteriol, 181: 68–77
Thompson J D, Gibson T J, Plewniak F, et al. 1997. The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res, 25: 4876–4882
Usov A I, Bilan M I, Klochkova N G. 1995. Polysaccharides of algae. 48. Polysaccharide composition of several calcareous red algae: isolation of alginate from Corallina pilulifera P. et R. (Rhodophyta, Corallinaceae). Botanica Marina, 38: 43–52
Author information
Authors and Affiliations
Corresponding authors
Additional information
Foundation item: The National High Technology Research and Development Program of China under contract No. 2012AA10A406; the National Natural Science Foundation of China under contract Nos 41206116, 31140070 and 31271397; Technology Project of Ocean and Fisheries of Guangdong Province under contract No. A201201E03; the Fundamental Research Funds for the Central Universities under contract No. 201262003; China Postdoctoral Science Foundation under contract No. 2011M501167; the algal transcriptome sequencing was supported by 1KP Project (www.onekp.com).
Rights and permissions
About this article
Cite this article
Wang, R., Wang, X., Zhang, Y. et al. Origin and evolution of alginate-c5-mannuronan-epimerase gene based on transcriptomic analysis of brown algae. Acta Oceanol. Sin. 33, 73–85 (2014). https://doi.org/10.1007/s13131-014-0443-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13131-014-0443-4