Abstract
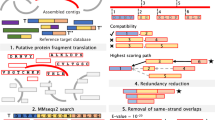
Currently, there is a need of non-computationally-intensive bioinformatics tools to cope with the increase of large datasets produced by Next Generation Sequencing technologies. We present a simple and robust bioinformatics pipeline to search for novel enzymes in metagenomic sequences. The strategy is based on pattern searching using as reference conserved motifs coded as regular expressions. As a case study, we applied this scheme to search for novel proteases S8A in a publicly available metagenome. Briefly, (1) the metagenome was assembled and translated into amino acids; (2) patterns were matched using regular expressions; (3) retrieved sequences were annotated; and (4) diversity analyses were conducted. Following this pipeline, we were able to identify nine sequences containing an S8 catalytic triad, starting from a metagenome containing 9,921,136 Illumina reads. Identity of these nine sequences was confirmed by BLASTp against databases at NCBI and MEROPS. Identities ranged from 62 to 89% to their respective nearest ortholog, which belonged to phyla Proteobacteria, Actinobacteria, Planctomycetes, Bacterioidetes, and Cyanobacteria, consistent with the most abundant phyla reported for this metagenome. All these results support the idea that they all are novel S8 sequences and strongly suggest that our methodology is robust and suitable to detect novel enzymes.

Similar content being viewed by others
References
Afgan E, Baker D, Batut B et al (2018) The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res 46:W537–W544. https://doi.org/10.1093/nar/gky379
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Amann R, Ludwig W, Schleifer K (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59:143–169
Andrews S (2010) FastQC. In: Qual. Control Tool High Throughput Seq. Data. www.bioinformatics.babraham.ac.uk/projects/fastqc. Accessed 4 Oct 2018
Bailey TL, Johnson J, Grant CE, Noble WS (2015) The MEME Suite. Nucleic Acids Res 43:W39–W49. https://doi.org/10.1093/nar/gkv416
Chen I-MA, Chu K, Palaniappan K et al (2019) IMG/M vol 5.0: an integrated data management and comparative analysis system for microbial genomes and microbiomes. Nucleic Acids Res 47:D666–D677. https://doi.org/10.1093/nar/gky901
DeLange R, Smith E (1967) Subtilisin Carlsberg. Amino acid composition; isolation and composition of peptides from the tryptic hydrolysate. J Biol Chem 243:2134–2142
Guindon S, Dufayard J-F, Lefort V et al (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. https://doi.org/10.1093/sysbio/syq010
Jisha VN, Smitha RB, Pradeep S et al (2013) Versatility of microbial proteases. Adv Enzyme Res 01:39–51. https://doi.org/10.4236/aer.2013.13005
Katoh K, Standley DM (2013) MAFFT multiple alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Keegan KP, Glass EM, Meyer F (2016) MG-RAST, a metagenomics service for analysis of microbial community structure and function. In: Martin F, Uroz S (eds) Microbial environmental genomics (MEG). Springer, New York, pp 207–233
Laskar M, James RE, Chatterjee A et al (2011) Modeling and structural analysis of evolutionarily diverse S8 family serine proteases. Bioinformation 7(5):239–245
Margulies M, Egholm M, Altman WE et al (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature. https://doi.org/10.1038/nature03959
Mitchell AL, Scheremetjew M, Denise H et al (2018) EBI Metagenomics in 2017: enriching the analysis of microbial communities, from sequence reads to assemblies. Nucleic Acids Res 46:D726–D735. https://doi.org/10.1093/nar/gkx967
NCBI Resource Coordinators (2012) Database resources of the national center for biotechnology information. Nucleic Acids Res 41:D8–D20. https://doi.org/10.1093/nar/gks1189
Nielsen HB, Almeida M, Juncker AS et al (2014) Identification and assembly of genomes and genetic elements in complex metagenomic samples without using reference genomes. Nat Biotechnol 32:822
Rawlings ND, Barrett AJ, Thomas PD et al (2018) The MEROPS database of proteolytic enzymes, their substrates and inhibitors in 2017 and a comparison with peptidases in the PANTHER database. Nucleic Acids Res 46:D624–D632. https://doi.org/10.1093/nar/gkx1134
Rice P, Longden I, Bleasby A (2000) EMBOSS: the European molecular biology open software suite. Trends Genet 16:276–277. https://doi.org/10.1016/S0168-9525(00)02024-2
Shendure J, Balasubramanian S, Church GM et al (2017) DNA sequencing at 40: past, present and future. Nature 550:345–353. https://doi.org/10.1038/nature24286
Thézé J, Li T, du Plessis L et al (2018) Genomic epidemiology reconstructs the introduction and spread of Zika virus in Central America and Mexico. Cell Host Microbe 23:855-864.e7. https://doi.org/10.1016/j.chom.2018.04.017
Acknowledgements
The authors wish to express their gratitude to National Science and Technology Council, Mexico for providing the financial support for this research (Project No. INFR-2016-01-269833). The authors thank César de los Santos-Briones and Mildred R. Carrillo-Pech for their technical assistance.
Author information
Authors and Affiliations
Contributions
All the authors contributed to this work. Góngora-Castillo and Ramirez-Prado designed and performed the experiments and analyzed the data; Caamal-Pech, Contreras-De la Rosa and Apolinar-Hernández participated in performing the experiments and the data analysis. López-Ochoa and Quiroz-Moreno participated in drafting the paper and discussing results. O’Connor-Sanchez, Ramirez-Prado and Góngora-Castillo conceived and designed the research and wrote the paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
Each of the authors confirms that this manuscript is original, has not been previously published and is not currently under consideration by any other journal. Additionally, all of the authors have approved the contents of this paper and have agreed to the 3 Biotech’s submission policies. The manuscript has two corresponding authors, who are Dr. Jorge H Ramírez-Prado and Dr. Aileen O’Connor-Sánchez.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Góngora-Castillo, E., López-Ochoa, L.A., Apolinar-Hernández, M.M. et al. Data mining of metagenomes to find novel enzymes: a non-computationally intensive method. 3 Biotech 10, 78 (2020). https://doi.org/10.1007/s13205-019-2044-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-019-2044-6