Abstract
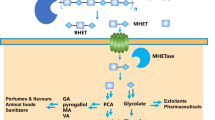
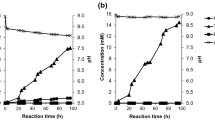
Poly(ethylene terephthalate) (PET) is a petroleum-based plastic that is massively produced and used worldwide. A promising PET recycling process to circumvent petroleum feedstock consumption and help to reduce environmental pollution is microbial or enzymatic biodegradation of post-consumer (PC) PET packages to its monomers—terephthalic acid (TPA) and ethylene glycol (EG)—or to key intermediates in PET synthesis—such as mono- and bis-(2-hydroxyethyl) terephthalate (MHET and BHET). Two species of filamentous fungi previously characterized as lipase producers (Penicillium restrictum and P. simplicissimum) were evaluated in submerged fermentation for induction of lipase production by two inducers (BHET and amorphous PET), and for biodegradation of two substrates (BHET and PC-PET). BHET induced lipase production in P. simplicissimum, achieving a peak of 606.4 U/L at 49 h (12.38 U/L.h), representing an almost twofold increase in comparison to the highest peak in the control (without inducers). Microbial biodegradation by P. simplicissimum after 28 days led to a 3.09% mass loss on PC-PET fragments. In contrast, enzymatic PC-PET depolymerization by cell-free filtrates from a P. simplicissimum culture resulted in low concentrations of BHET, MHET and TPA (up to 9.51 µmol/L), suggesting that there are mechanisms at the organism level that enhance biodegradation. Enzymatic BHET hydrolysis revealed that P. simplicissimum extracellular enzymes catalyze the release of MHET as the predominant product. Our results show that P. simplicissimum is a promising biodegrader of PC-PET that can be further explored for monomer recovery in the context of feedstock recycling processes.






Similar content being viewed by others
Accession numbers
Penicillium simplicissimum 28f2 (acession number 071113-05 at the International Depository Authority of Canada—IDAC).
Abbreviations
- BHET:
-
Bis-(2-hydroxyethyl) terephthalate
- EG:
-
Ethylene glycol
- MHET:
-
Mono-(2-hydroxyethyl) terephthalate
- PC-PET:
-
Post-consumer PET
- PET:
-
Poly(ethylene terephthalate)
- SSF:
-
Solid-state fermentation
- TPA:
-
Terephthalic acid
References
Bai H, Wang H, Sun J, Irfan M, Han M, Huang Y, Han X, Yang Q (2013) Production, purification and characterization of novel beta glucosidase from newly isolated Penicillium simplicissimum H-11 in submerged fermentation. EXCLI J 12:528–540
Barth M, Oeser T, Wei R, Then J, Schmidt J, Zimmermann W (2015) Effect of hydrolysis products on the enzymatic degradation of polyethylene terephthalate nanoparticles by a polyester hydrolase from Thermobifida fusca. Biochem Eng J 93:222–228. https://doi.org/10.1016/j.bej.2014.10.012
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Carniel A, Valoni E, Junior JN, Gomes AC, Castro AM (2017) Lipase from Candida antarctica (CALB) and cutinase from Humicola insolens act synergistically for PET hydrolysis to terephthalic acid. Process Biochem 59:84–90. https://doi.org/10.1016/j.procbio.2016.07.023
Carniel A, Waldow VA, Castro AM (2021) A comprehensive and critical review on key elements to implement enzymatic PET depolymerization for recycling purposes. Biotech Adv 52:107811. https://doi.org/10.1016/j.biotechadv.2021.107811
Castro AM, Carniel A, Junior JN, Gomes AC, Valoni E (2017) Screening of commercial enzymes for poly(ethylene terephthalate) (PET) hydrolysis and synergy studies on different substrate sources. J Ind Microbiol Biotechnol 44:835–844. https://doi.org/10.1007/s10295-017-1942-z
Copley SD (2017) Shining a light on enzyme promiscuity. Curr Opin Struc Biol 47:167–175. https://doi.org/10.1016/j.sbi.2017.11.001
Costa AM, Oliveira Lopes VR, Vidal L, Nicaud JM, Castro AM, Coelho MAZ (2020) Poly(ethylene terephthalate) (PET) degradation by Yarrowia lipolytica: investigations on cell growth, enzyme production and monomers consumption. Process Biochem 95:81–90. https://doi.org/10.1016/j.procbio.2020.04.001
Crippa M, Morico B (2019) PET depolymerization: a novel process for plastic waste chemical recycling. Stud Surf Sci Catal 179:215–229. https://doi.org/10.1016/B978-0-444-64337-7.00012-4
Eberl A, Heumann S, Brückner T, Araujo R, Cavaco-Paulo A, Kaufmann F, Kroutil W, Guebitz GM (2009) Enzymatic surface hydrolysis of poly(ethylene terephthalate) and bis(benzoyloxyethyl) terephthalate by lipase and cutinase in the presence of surface active molecules. J Biotechnol 143(3):207–212. https://doi.org/10.1016/j.jbiotec.2009.07.008
Espino-Rammer L, Ribitsch D, Przylucka A, Marold A, Greimel KJ, Acero EH, Guebitz GM, Kubicek CP, Druzhinina IS (2013) Two novel class II hydrophobins from Trichoderma spp. stimulate enzymatic hydrolysis of poly(ethylene terephthalate) when expressed as fusion proteins. Appl Environ Microbiol 79(14):4230–4238. https://doi.org/10.1128/AEM.01132-13
Freire DM, Gomes PM, Bon EPS, Sant’Anna GL Jr (1997a) Lipase production by a new promising strain of Penicillium restrictum. Rev Microbiol 28(Suppl. 1):6–12
Freire DM, Teles EMF, BonSant’Anna EPSGL Jr (1997b) Lipase production by Penicillium restrictum in a bench-scale fermenter: effect of carbon and niltrogen nutrition, agitation, and aeration. Appl Biochem Biotechnol 63:09–421. https://doi.org/10.1007/BF02920442
Freire DMG (1996) Seleção de microorganismos lipolíticos e estudo da produção de lipase por Penicillium restrictum. Dissertation, Universidade Federal do Rio de Janeiro
Gan Z, Zhang H (2019) PMBD: a comprehensive plastics microbial biodegradation database. Database 2019:1–11. https://doi.org/10.1093/database/baz119
Gautam G, Mishra V, Verma P, Pandey AK, Negi S (2014) A cost effective strategy for production of bio-surfactant from locally isolated Penicillium chrysogenum SNP5 and its applications. J Bioproces Biotech 4:6. https://doi.org/10.4172/2155-9821.1000177
Geyer R, Jambeck JR, Law KL (2017) Production, use, and fate of all plastics ever made. Sci Adv 3:e1700782. https://doi.org/10.1126/sciadv.1700782
Godoy MG, Gutarra MLE, Castro AM, Machado OLT, Freire DMG (2011) Adding value to a toxic residue from the biodiesel industry: production of two distinct pool of lipases from Penicillium simplicissimum in castor bean waste. J Ind Microbiol Biotechnol 38(8):945–953. https://doi.org/10.1007/s10295-010-0865-8
Guebitz GM, Cavaco-Paulo A (2007) Enzymes go big: surface hydrolysis and functionalisation of synthetic polymers. Trends Biotechnol 26(1):32–38. https://doi.org/10.1016/j.tibtech.2007.10.003
Gutarra MLE, Cavalcanti EDC, Castilho LR, Freire DMG, Sant’Anna GL Jr (2005) Lipase production by solid-state fermentation: cultivation conditions and operation of tray and packed-bed bioreactors. Appl Biochem Biotechnol 121(1–3):105–116. https://doi.org/10.1385/ABAB:121:1-3:0105
Gutarra MLE, Godoy MG, Castilho LR, Freire DMG (2007) Inoculum strategies for Penicillium simplicissimum lipase production by solid-state fermentation using a residue from the babassu oil industry. J Chem Technol Biotechnol 82(3):313–318. https://doi.org/10.1002/jctb.1674
Gutarra MLE, Godoy MG, Silva JN, Guedes IA, Lins U, Castilho LR, Freire DMG (2009a) Lipase production and Penicillium simplicissimum morphology in solid-state and submerged fermentations. Biotechnol J 4:1450–1459. https://doi.org/10.1002/biot.200800298
Gutarra MLE, Godoy MG, Maugeri F, Rodrigues MI, Freire DMG, Castilho LR (2009b) Production of an acidic and thermostable lipase of the mesophilic fungus Penicillium simplicissimum by solid-state fermentation. Bioresour Technol 100(21):5249–5254. https://doi.org/10.1016/j.biortech.2008.08.050
Hall CAS (2017) Will EROI be the primary determinant of our economic future? The view of the natural scientist versus the economist. Joule 1(4):635–638. https://doi.org/10.1016/j.joule.2017.09.010
Hantani Y, Imamura H, Yamamoto SA, Yamagami Y, Kato M, Kawai F, Oda M (2018) Functional characterizations of polyethylene terephthalate-degrading cutinase-like enzyme Cut190 mutants using bis(2-hydroxyethyl) terephthalate as the model substrate. AIMS Biophys 5(4):290–302. https://doi.org/10.3934/biophy.2018.4.290
Hasan F, Shah AA, Hameed A (2009) Methods for detection and characterization of lipases: a comprehensive review. Biotech Adv 27:782–798. https://doi.org/10.1016/j.biotechadv.2009.06.001
Hooker J, Hinks D, Montero G, Icherenska M (2003) Enzyme-catalyzed hydrolysis of poly(ethylene terephthalate) cyclic trimer. J Appl Polym Sci 89:2545–2552. https://doi.org/10.1002/app.11963
Hyde KD et al (2019) The amazing potential of fungi: 50 ways we can exploit fungi industrially. Fungal Divers 97:1–136. https://doi.org/10.1007/s13225-019-00430-9
Ion S, Voicea S, Sora C, Gheorghita G, Tudorache M, Parvulescu VI (2020) Sequential biocatalytic decomposition of BHET as valuable intermediator of PET recycling strategy. Catal Today 366:177–184. https://doi.org/10.1016/j.cattod.2020.08.008
Kawai F, Kawabata T, Oda M (2019) Current knowledge on enzymatic PET degradation and its possible application to waste stream management and other fields. Appl Microbiol Biotechnol 103:4253–4268. https://doi.org/10.1007/s00253-019-09717-y
Kawai F, Kawabata T, Oda M (2020) Current state and perspectives related to the PET hydrolases available for biorecycling. ACS Sustain Chem Eng 8(24):8894–8908. https://doi.org/10.1021/acssuschemeng.0c01638
Kim DY, Rhee YH (2003) Biodegradation of microbial and synthetic polyesters by fungi. Appl Microbiol Biotechnol 61:300–308. https://doi.org/10.1007/s00253-002-1205-3
Koshti R, Mehta L, Samarth N (2018) Biological recycling of polyethylene terephthalate: a mini-review. J Polym Environ 26:3520–3529. https://doi.org/10.1007/s10924-018-1214-7
Kumar AG, Anjana K, Hinduja M, Sujitha K, Dharani G (2020) Review on plastic wastes in marine environment—biodegradation and biotechnological solutions. Mar Pollut Bull 150:110733. https://doi.org/10.1016/j.marpolbul.2019.110733
Lepoittevin B, Roger P (2011) Poly(ethylene terephthalate). In: Thomas S, Visakh PM (eds) Handbook of engineering and speciality thermoplastics: polyethers and polyesters, vol 3. Scrivener Publishing, Beverly, pp 97–126. https://doi.org/10.1002/9781118104729.ch4
Liebminger S, Eberl A, Sousa F, Heumann S, Fischer-Colbrie G, Cavaco-Paulo A, Guebitz GM (2007) Hydrolysis of PET and 3PET with a new polyesterase from Penicillium citrinum. Biocatal Biotransform 25(2–4):171–177. https://doi.org/10.1080/10242420701379734
Malafatti-Picca L, Chaves MRB, Castro AM, Valoni E, Oliveira VM, Marsaioli AJ, Angelis DF, Attili-Angelis D (2019) Hydrocarbon-associated substrates reveal promising fungi for poly(ethylene terephthalate) (PET) depolymerization. Braz J Microbiol 50:633–648. https://doi.org/10.1007/s42770-019-00093-3
Markit IHS (2018) Chemical economics handbook—PET polymer. https://ihsmarkit.com/products/pet-polymer-chemical-economics-handbook.html. Accessed 10 Sept 2020
Nimchua T, Punnapayak H, Zimmermann W (2007) Comparison of the hydrolysis of polyethylene terephthalate fibers by a hydrolase from Fusarium oxysporum LHC I and Fusarium solani f. sp. pisi. Biotechnol J 2:361–364. https://doi.org/10.1002/biot.200600095
Nimchua T, Eveleigh DE, Sangwatanaroj U, Punnapayak H (2008) Screening of tropical fungi producing polyethylene terephthalate-hydrolyzing enzyme for fabric modification. J Ind Microbiol Biotechnol 35:843–850. https://doi.org/10.1007/s10295-008-0356-3
Nowak B, Pajak J, Labuzek S, Rymarz G, Talik E (2011) Biodegradation of poly(ethylene terephthalate) modified with polyester “Bionolle®” by Penicillium funiculosum. Polimery 56(1):35–44
Qiu L, Yin X, Liu T, Zhang H, Chen G, Wu S (2020) Biodegradation of bis(2‐hydroxyethyl) terephthalate by a newly isolated Enterobacter sp. HY1 and characterization of its esterase properties. J Basic Microbiol 60:699–711. https://doi.org/10.1002/jobm.202000053
Raheem AB, Noor ZZ, Hassan A, Hamid MKA, Samsudin SA, Sabeen AH (2019) Current developments in chemical recycling of post-consumer polyethylene terephthalate wastes for new materials production: a review. J Clean Prod 225:1052–1064. https://doi.org/10.1016/j.jclepro.2019.04.019
Ronkvist AM, Xie W, Lu W, Gross RA (2009) Cutinase-catalyzed hydrolysis of poly(ethylene terephthalate). Macromolecules 42:5128–5138. https://doi.org/10.1021/ma9005318
Sales JCS, Castro AM, Ribeiro BD, Coelho MAZ (2020) Supplementation of watermelon peels as an enhancer of lipase and esterase production by Yarrowia lipolytica in solid-state fermentation and their potential use as biocatalysts in poly(ethylene terephthalate) (PET) depolymerization reactions. Biocatal Biotransform 38:457–468. https://doi.org/10.1080/10242422.2020.1782387
Sales JCS, Castro AM, Ribeiro BD, Coelho MAZ (2021) Improved production of biocatalysts by Yarrowia lipolytica using natural sources of the biopolyesters cutin and suberin, and their application in hydrolysis of poly (ethylene terephthalate) (PET). Bioprocess Biosyst Eng. https://doi.org/10.1007/s00449-021-02603-w
Salvador M, Abdulmutalib U, Gonzalez J, Kim J, Smith AA, Faulon JP, Wei R, Zimmermann W, Jimenez JI (2019) Microbial genes for a circular and sustainable bio-PET economy. Genes 10:373. https://doi.org/10.3390/genes10050373
Sánchez C (2020) Fungal potential for the degradation of petroleum-based polymers: an overview of macro- and microplastics biodegradation. Biotechnol Adv 40:107501. https://doi.org/10.1016/j.biotechadv.2019.107501
Sharon C, Sharon M (2012) Studies on biodegradation of polyethylene terephthalate: a synthetic polymer. J Microbiol Biotech Res 2(2):248–257
Silva CM, Carneiro F, O’Neill A, Fonseca LP, Cabral JSM, Guebitz G, Cavaco-Paulo A (2005) Cutinase—a new tool for biomodification of synthetic fibres. J Polym Sci, Part A: Polym Chem 43:2448–2450. https://doi.org/10.1002/pola.20684
Sinha V, Patel MR, Patel JV (2010) PET waste management by chemical recycling: a review. J Polym Environ 18:8–25. https://doi.org/10.1007/s10924-008-0106-7
Syedd-León R, Sandoval-Barrantes M, Trimiño-Vásquez H, Villegas-Peñaranda LR, Rodríguez-Rodríguez G (2020) Revisiting the fundamentals of p-nitrophenol analysis for its application in the quantification of lipases activity. A graphical update. Uniciencia 34(2):31–43. https://doi.org/10.15359/ru.34-2.2
Taniguchi I, Yoshida S, Hiraga K, Miyamoto K, Kimura Y, Oda K (2019) Biodegradation of PET: current status and application aspects. ACS Catal 9(5):4089–4105. https://doi.org/10.1021/acscatal.8b05171
Tournier V, Topham CM, Gilles A, David B, Folgoas C, Moya-Leclair E, Kamionka E, Desrousseaux ML, Texier H, Gavalda S, Cot M, Guémard E, Dalibey M, Nomme J, Cioci G, Barbe S, Chateau M, André I, Duquesne S, Marty A (2020) An engineered PET depolymerase to break down and recycle plastic bottles. Nature 580(7802):216–219. https://doi.org/10.1038/s41586-020-2149-4
Vertommen MAME, Nierstrasz VA, Veer MVD, Warmoeskerken MMCG (2005) Enzymatic surface modification of poly(ethylene terephthalate). J Biotechnol 120:376–386. https://doi.org/10.1016/j.jbiotec.2005.06.015
Wang X, Lu D, Jönsson LJ, Hong F (2008) Preparation of a PET-hydrolyzing lipase from Aspergillus oryzae by the addition of bis(2-hydroxyethyl) terephthalate to the culture medium and enyzmatic modification of PET fabrics. Eng Life Sci 8(3):268–276. https://doi.org/10.1002/elsc.200700058
Wei R, Zimmermann W (2017a) Biocatalysis as a green route for recycling the recalcitrant plastic polyethylene terephthalate. Microb Biotechnol 10(6):1302–1307. https://doi.org/10.1111/1751-7915.12714
Wei R, Zimmermann W (2017b) Microbial enzymes for the recycling of recalcitrant petroleum-based plastics: how far are we? Microb Biotechnol 10(6):1308–1322. https://doi.org/10.1111/1751-7915.12710
Wierckx N, Narancic T, Eberlein C, Wei R, Drzyzga O, Magnin A, Ballerstedt H, Kenny ST, Pollet E, Avérous L, O’Connor KE, Zimmermann W, Heipieper HJ, Prieto A, Jiménez J, Blank LM (2018) Plastic biodegradation: callenges and opportunities. In: Steffan R (ed) Consequences of microbial interactions with hydrocarbons, oils, and lipids: biodegradation and bioremediation, handbook of hydrocarbon and lipid microbiology. Springer International Publishing AG, Basel, pp 1–29. https://doi.org/10.1007/978-3-319-44535-9_23-1
Yamada-Onodera K, Mukomoto H, Katsuyaya Y, Saiganji A, Tani Y (2001) Degradation of polyethylene by a fungus, Penicillium simplicissimum YK. Polym Degrad Stab 72:323–327. https://doi.org/10.1016/S0141-3910(01)00027-1
Yoshida S, Hiraga K, Takehana T, Taniguchi I, Yamaji H, Maeda Y, Toyohara K, Miyamoto K, Kimura Y, Oda K (2016) A bacterium that degrades and assimilates poly(ethylene terephthalate). Science 351(6278):1196–1199. https://doi.org/10.1126/science.aad6359
Zhang J, Wang X, Gong J, Gu Z (2004) A study on the biodegradability of polyethylene terephthalate fiber and diethylene glycol terephthalate. J Appl Polym Sci 93(3):1089–1096. https://doi.org/10.1002/app.20556
Zheng J, Suh S (2019) Strategies to reduce the global carbon footprint of plastics. Nat Clim Chang 9:374–378. https://doi.org/10.1038/s41558-019-0459-z
Zimmermann W (2020) Biocatalytic recycling of polyethylene terephthalate plastic. Phil Trans R Soc A 378:20190273. https://doi.org/10.1098/rsta.2019.0273
Zimmermann W, Billig S (2011) Enzymes for the biofunctionalization of poly(ethylene terephthalate). Adv Biochem Eng/biotechnol 125:97–120. https://doi.org/10.1007/10_2010_87
Acknowledgements
This study was funded by Petrobras (Petróleo Brasileiro S.A.).
Author information
Authors and Affiliations
Contributions
DNS and DAT planned and executed the laboratory work; VAW wrote the original draft; DMGF reviewed the original draft; AMC planned and supervised the laboratory work and reviewed the original draft.
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Rights and permissions
About this article
Cite this article
Moyses, D.N., Teixeira, D.A., Waldow, V.A. et al. Fungal and enzymatic bio-depolymerization of waste post-consumer poly(ethylene terephthalate) (PET) bottles using Penicillium species. 3 Biotech 11, 435 (2021). https://doi.org/10.1007/s13205-021-02988-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-021-02988-1