Abstract
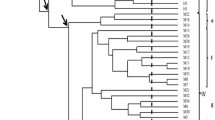
The genetic relationships in Staphylococcus aureus isolated from raw poultry obtained from various Brazilian broiler chicken processors were analyzed using repetitive extragenic palindromic-polymerase chain reaction (rep-PCR). The distribution of accessory gene regulator (agr) groups was determined, and the presence of biofilm-associated genes and phenotypic features, including biofilm formation and proteolytic, lipolytic, and β-hemolytic activities, was assessed. Isolates were grouped into three major clusters based on rep-PCR fingerprints typed with RW3A primer. The agr group I was the most common genotype identified (86.21 %), followed by groups II (10.34 %) and III (3.45 %). All strains were positive for the sasG gene; the next most frequent genes were icaA (93.1 %) and atlA (51.72 %). Twenty-six of the 29 isolates were biofilm producers. In this study, 96.55 %, 72.41 %, and 62.06 % of the isolates displayed lipolytic, β-hemolytic, and proteolytic activity, respectively. In conclusion, the rep-PCR results suggested a clonal relationship among the S. aureus isolated from raw poultry produced by different broiler chicken processors. Our results also showed that most isolates belonged to agr group I. The presence of biofilm-forming S. aureus strains in raw poultry, their ability to harbor biofilm-associated genes, and the spoilage features that they exhibit are indicative of their pathogenic potential, and may represent a serious problem in the food processing industry.


Similar content being viewed by others
References
Almeida LM, Zilta M, De Almeida PRB, Mendonça CL, Mamizuka LM (2013) Comparative analysis of agr groups and virulence genes among subclinical and clinical mastitis Staphylococcus aureus isolates from sheep flocks of the Northeast of Brazil. Braz J Microbiol 44:493–498
Bibalan MH, Shakeri F, Javid N, Ghaemi A, Ghaemi EA (2014) Accessory gene regulator types of Staphylococcus aureus isolated in Gorgan, North of Iran. J Clin Diagn Res 8:07–09
Del Vecchio VG, Petroziello JM, Gress MJ, McCleskey FK, Melcher GP, Crouch HK, Lupski JR (1995) Molecular genotyping of methicillin-resistant Staphylococcus aureus via fluorophore-enhanced repetitive-sequence PCR. J Clin Microbiol 33:2141–2144
Devita MD, Wadhera RK, Theis ML, Ingham SC (2007) Assessing the potential of Streptococcus pyogenes and Staphylococcus aureus transfer to foods and customers via a survey of hands, hand-contact surfaces and food-contact surfaces at foodservice facilities. J Food Serv 18:76–79
Gharsa H, Sallem RB, Slama KB, Gómez-Sanz E, Lozano C, Jouini A, Klibi N, Zarazaga M, Boudabous A, Torres C (2012) High diversity of genetic lineages and virulence genes in nasal Staphylococcus aureus isolates from donkeys destined to food consumption in Tunisia with predominance of the ruminant associated CC133 lineage. BMC Vet Res 8:203
Giaouris E, Heir E, Hébraud M, Chorianopoulos N, Langsrud S, Møretrø T, Habimana O, Desvaux M, Renier S, Nychas GJ (2014) Attachment and biofilm formation by foodborne bacteria in meat processing environments: causes, implications, role of bacterial interactions and control by alternative novel methods. Meat Sci 97:298–309
Gillet Y, Issartel B, Vanhems P, Fournet JC, Lina G, Bes M, Vandenesch F, Piémont Y, Brousse N, Floret D, Etienne J (2002) Association between Staphylococcus aureus strains carrying gene for Panton-Valentine leukocidin and highly lethal necrotising pneumonia in young immunocompetent patients. Lancet 359:753–759
Gilot P, Van Leeuwen JW (2004) Comparative analysis of agr locus diversification and overall genetic variability among bovine and human Staphylococcus aureus isolates. Clin Microbiol 42:1265
Goerke C, Esser S, Kummel M, Wolz C (2005) Staphylococcus aureus strain designation by agr and cap polymorphism typing and delineation of agr diversification by sequence analysis. Int J Med Microbiol 295:67–75
Greig JD, Ravel A (2009) Analysis of foodborne outbreak data reported internationally for source attribution. Int J Food Microbiol 130:77–87
Gundogan N, Ataol O, Gunal S (2012) Determination of some virulence factors in staphylococci isolated from meat and milk products. Arch Lebensmittelhyg 63:182–186
Gundogan N, Ataol O, Torlak FO (2013) Determination of some virulence factors in Staphylococcus aureus, Enterococcus faecalis and Enterococcus faecium isolated from meat and milk products. J Food Safety 33:387–393
Gutiérrez D, Delgado S, Vázquez-Sánchez D, Martínez B, Cabo ML, Rodríguez A (2012) Incidence of Staphylococcus aureus and analysis of associated bacterial communities on food industry surfaces. Appl Environ Microbiol 78:8547–8554
Hanselman BA, Kruth SA, Rousseau J, Weese JS (2009) Coagulase positive staphylococcal colonization of humans and their household pets. Can Vet 50:954–958
Jarraud S, Mougel C, Thioulouse J, Lina G, Meugnier H, Forey F, Nesme X, Etienne J, Vandenesch F (2002) Relationships between Staphylococcus aureus genetic background, virulence factors, agr groups (alleles), and human disease. Infect Immun 70:631–641
Jessen B, Lammert L (2003) Biofilm and disinfection in meat processing plants. Int Biodeterior Biodegrad 51:265–279
Khan S, Rasheed F, Zahra R (2014) Genetic polymorphism of agr locus and antibiotic resistance of Staphylococcus aureus at two hospitals in Pakistan. Pak J Med Sci 30:172–176
Martins PD, De Almeida TT, Basso AP, De Moura TM, Frazzon J, Tondo EC, Frazzon APG (2013) Coagulase-positive staphylococci isolated from chicken meat: pathogenic potential and vancomycin resistance. Foodborne Pathog Dis 10:771–776
Moise-Broder PA, Sakoulas G, Eliopoulos GM, Schentag JJ, Forrest A, Moellering RCJR (2004) Accessory gene regulator group II polymorphism in methicillin-resistant Staphylococcus aureus is predictive of failure of vancomycin therapy. Clin Infect Dis 38:1700–1705
Moura TM, Campos FS, D´Azevedo PA, Van Der Sand ST, Franco AC, Frazzon J, Frazzon APG (2012) Prevalence of enterotoxin-encoding genes and antimicrobial resistance in coagulase-negative and coagulase-positive Staphylococcus isolates from black pudding in Southern Brazil. Rev Soc Bras Med Trop 45:579–585
Njage PMK, Dolci S, Jans C, Wangoh J, Lacroix C, Meile L (2013) Biodiversity and enterotoxigenic potential of staphylococci isolated from raw and spontaneously fermented camel milk. British Microb Research J 3:128–138
Novick RP, Schlievert P, Ruzin A (2001) Pathogenicity and resistance islands of staphylococci. Microbes Infect 3:585–594
Otto M (2013) Staphylococcal infections: mechanisms of biofilm maturation and detachment as critical determinants of pathogenicity. Annu Rev Med 64:175–188
Parkash M, Rajasekar K, Karmegam N (2007) Bacterial population of raw milk and their proteolytic and lipolytic activities. Res J Bas Appl Sci 3:848–851
Peacock SJ, Moore CE, Justice A, Kantzanou M, Story L, Mackie K, O’Neill G, Day NP (2002) Virulent combinations of adhesin and toxin genes in natural populations of Staphylococcus aureus. Infect Immun 70:4987–4996
Pereira V, Lopes C, Castro A, Silva J, Gibbs P, Teixeira P (2009) Characterization for enterotoxin production, virulence factors, and antibiotic susceptibility of Staphylococcus aureus isolates from various foods in Portugal. Food Microbiol 26:278–282
Pu S, Wang F, Ge B (2011) Characterization of toxin genes and antimicrobial susceptibility of Staphylococcus aureus isolates from Louisiana retail meats. Foodborne Pathog Dis 8:299–306
Reinoso EB, El-Sayedb A, Lämmlerb C, Bognia C, Zscho¨ckc M (2008) Genotyping of Staphylococcus aureus isolated from humans, bovine subclinical mastitis and food samples in Argentina. Microbiol Res 163:314–322
Reiter KC, Villa B, Da Silva Paim TG, Sambrano GE, De Oliveira CF, D'azevedo PA (2012) Enhancement of antistaphylococcal activities of six antimicrobials against sasG negative methicillin-susceptible Staphylococcus aureus: an in vitro biofilm model. Diagn Microbiol Infect Dis 74:101–105
Roche FM, Meehan M, Foster TJ (2003) The Staphylococcus aureus surface protein SasG and its homologues promote bacterial adherence to human desquamated nasal epithelial cells. Microbiology 149:2759–2767
Rodrigues LB, Dos Santos LR, Tagliari VZ, Rizzo NN, Trenhago G, De Oliveira AP, Goetz F, Nascimento VP (2010) Quantification of biofilm production on polystyrene by Listeria, Escherichia coli and Staphylococcus aureus isolated from a poultry slaughterhouse. Braz J Microbiol 41:1082–1085
Ruaro A, Andrighetto C, Torriani S, Lombardi A (2013) Biodiversity and characterization of indigenous coagulase-negative staphylococci isolated from raw milk and cheese of North Italy. Food Microbiol 34:106–111
Sadoyama G, Santos KR, Brilhante AP, Filho PP (2008) Staphylococcus aureus as source of catheter related bloodstream infection evaluated by PFGE and rep-PCR typing in a Brazilian hospital. APMIS 116:953–960
Sakoulas G, Eliopoulos GM, Moellering RCJR, Novick RP, Venkataraman L, Wennersten C, Degirolami PC, Schwaber MJ, Gold HS (2003) Staphylococcus aureus accessory gene regulator (agr) group II: is there a relationship to the development of intermediate-level glycopeptide resistance? J Infect Dis 187:929–938
Santos KRN, Fonseca LS, Teixeira LM, Gontijo Filho PP (2001) Typing of Staphylococcus aureus from surgical site infections: comparison of pulsed-field gel electrophoresis (PFGE) and PCR technique using repetitive extragenic palindromic (rep) and Tn916-Shine-Dalgarno (TnSD) target sequences. Int J Med Microbiol 291:231–236
Spanu V, Spanu C, Virdis S, Cossu F, Scarano C, De Santis EP (2012) Virulence factors and genetic variability of Staphylococcus aureus strains isolated from raw sheep’s milk cheese. Int J Food Microbiol 153:53–57
Stepanović S, Vuković D, Hola V, Bonaventura G, Djukić S, Ćirković I, Ruzicka F (2007) Quantification of biofilm in microtiter plates: overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. APMIS 115:891–899
Takeuchi S, Kinoshita T, Kaidoh T, Hashizume N (1999) Purification and characterization of protease produced by Staphylococcus aureus isolated from a diseased chicken. Vet Microbiol 67:195–202
Tang J, Chen J, Li H, Zeng P, Li J (2013) Characterization of adhesin genes, staphylococcal nuclease, hemolysis, and biofilm formation among Staphylococcus aureus strains isolated from different sources. Foodborne Pathog Dis 10:757–763
Vázquez-Sánchez D, Habimana O, Holck A (2013) Impact of food-related environmental factors on the adherence and biofilm formation of natural Staphylococcus aureus isolates. Curr Microbiol 66:110–121
Weese JS, Reid-Smith R, Rousseau J, Avery B (2010) Methicillin-resistant Staphylococcus aureus (mrsA) contamination of retail pork. Can Vet J 51:749–52
Acknowledgments
We thank the Conselho Nacional de Desenvolvimento Científico e Tecnológico do Brasil (CNPq) and the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) of the Brazilian government for the support received.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pinto, J.B., Rossatto, F.C.P., Martins, P.D. et al. Genetic relationships and virulence factors in Staphylococcus aureus isolated from raw poultry in South Brazil. Ann Microbiol 65, 1933–1940 (2015). https://doi.org/10.1007/s13213-014-1031-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-014-1031-8