Abstract
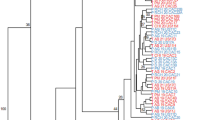
Late blight disease caused by an Oomycete Phytophthora infestans is a major constraint to potato production and is causing significant yield losses in Ethiopia. This study was conducted to characterize the genetic diversity of the pathogen population in the major potato growing regions Awi, East Hararghe, South Gondar, West Arsi, West Gojjam and West Shewa in Ethiopia. In total 138 P. infestans isolates were collected using FTA cards in 2017, genotyped using 12-plex SSR markers and characterized for the mitochondrial haplotype. The genotypic patterns were compared to those of reference isolates from the EU2_A1 and US-1 clonal lineages. The population structure analysis using discriminant analysis of principal components (DAPC) and STRUCTURE indicated that most of the Ethiopian isolates were similar to the EU2_A1, while a second cluster of isolates was formed that was clearly different from EU2_A1 as well as the US-1 reference isolates. This new genotype was characterized by private alleles in the SSR D13 locus. We named this new genotype as ET-1 lineage. All isolates had the same mitochondrial haplotype (Ia). EU2_A1 was dominant clonal lineage in all locations except West Arsi which was dominated with ET-1 lineage. The old US-1 lineage was not discovered among the Ethiopian samples which suggest that it has been displaced. The West Arsi, West Gojjam and West Shewa populations were found to contain the highest genetic diversity, with the greatest number of multi locus genotypes (MLGs) and a higher diversity index compared to the other locations. The findings of this study establish a baseline of the pathogen population diversity in Ethiopia. Continuous tracking of P. infestans population in both potato and tomato is recommended to monitor the changes and migration patterns.




Similar content being viewed by others
References
Bruvo R, Michiels NK, D’Souza TG, Schulenburg H (2004) A simple method for the calculation of microsatellite genotype distances irrespective of ploidy level. Mol Ecol 13:2101–2106
Chowdappa P, Nirmal Kumar BJ, Madhura S, Mohan Kumar SP, Myers KL, Fry WE, Cooke DEL (2015) Severe outbreaks of late blight on potato and tomato in South India caused by recent changes in the Phytophthora infestans population. Plant Pathol 64(1):191–199
Clark LV, Jasieniuk M (2011) POLYSAT: an R package for polyploid microsatellite analysis. Mol Ecol Resour 11:562–566
Cooke DEL, Lees AK (2004) Markers, old and news, for examining Phytophthora infestans diversity. Plant Pathol 53: 692-704
Dey T, Saville A, Myers K et al (2018) Large sub-clonal variation in Phytophthora infestans from recent severe late blight epidemics in India. Sci Rep 8:4429
de Vries S, von Dahlen JK, Uhlmann C, Schnake A, Kloesges T, Rose LE (2017) Signatures of selection and host-adapted gene expression of the Phytophthora infestans RNA silencing suppressor PSR2. Mol Plant Pathol 18:110–124
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Forbes GA, Gamboa S, Lindqvist-Kreuze H, Oliva RF, Perez W (2016) Identification of an A2 population of Phythophthora andina attacking tree tomato in Peru indicates a risk of sexual reproduction in this pathosystem. Plant Pathol 65:1109–1117
Fry WE, Goodwin SB, Dyer AT, Matuszak JM, Drenth A et al (1993) Historical and recent migrations of Phytophthora infestans: chronology, pathways,and implications. Plant Disease, 77:653–661
Fry WE, Birch PRJ, Judelson HS, Grünwald NJ, Danies G, Everts KL, Gevens AJ, Gugino BK, Johnson DA, Johnson SB, McGrath M (2015) Five reasons to consider Phytophthora infestans a reemerging pathogen. Phytopathology 105(7):966–981
Gebremedhin W (2013) Potato variety development strategies and methodologies in Ethiopia. In: Gebremedhin W, Schulz S, Bave B, eds. Proceedings of the National Workshop on Seed potato tuber production and disseminations: experiences, challenges and prospects. Bahir Dar, Ethiopia: Ethiopian Institute of Agricultural Research and Amhara Region Agriculture Research Institute, 45–59
Goodwin SB (1997) The population genetics of Phytophthora. Phytopathology 87: 462–473
Griffith GW, Shaw DS (1998) Polymorphisms in Phytophthora infestans: Four Mitochondrial Haplotypes Are Detected after PCR Amplification of DNA from Pure Cultures or from Host Lesions. Appl Environ Microbiol 64:4007–4014
Hedrick PW (1999) Perspective: Highly variable loci and their interpretation in evolution and conservation. International Journal of Organic Evolution 53:313–318
Hedrick PW (2005) A standardized genetic differentiation measure. Evolution 59:1633–1638
Hu C-H, Perez FG, Donahoo R et al (2012) Recent Genotypes of Phytophthora infestans in the Eastern United States Reveal Clonal Populations and Reappearance of Mefenoxam Sensitivity. Plant Dis 96:1323–1330
Hurlbert SH (1971) The Nonconcept of Species Diversity: A Critique and Alternative Parameters. Ecology 52:577–586
Hussain T (2016) Diagnostic of most important pathogens on Potato through PCR techniques: a Systematic Review. Trends in Biosciences 9(4):203–212
Hussain T (2017) Potatoes: Ensuring food for the future. Adv in Plants and Agricultural Res 3(6):00117
Imranul Haq QM, Hussain T, Kumar A (2016) Molecular markers: A tool to identify hidden science with special emphasis on agricultural crops. International Journal of Biology Research 1(5):50–56
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics (Oxford, England) 23:1801–1806
Jombart T, Ahmed I (2011) adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinformatics (Oxford, England) 27:3070–3071
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94
Kamoun S, Furzer O, Jones JDG et al (2015) The Top 10 oomycete pathogens in molecular plant pathology. Mol Plant Pathol 16:413–434
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:281
Knapova G, Tenzer I, Gessler C and Gisi U (2001) Characterization of Phytophthora infestans from potato and tomato with molecular markers. Proceedings of the 5th Congress of the European Foundation for Plant Pathology (Biodiversity in Plant Pathology). Taormina, Italy: SIVP, pp.6–9
Knapova G, Gisi U (2002) Phenotypic and genotypic structure of Phytophthora infestans populations on potato and tomato in France and Switzerland. Plant Pathol 51(5):641–53. https://doi.org/10.1046/j.1365-3059.2002.00750
Kassa B, Olanya M, Tesfaye A, Lemaga B, Woldegiorgis G (2002) Economic implications of late blight management in tropical highlands of Ethiopia. In: Lizárraga C ed. Proceedings of the Global Initiative on Late Blight Conference. Hamburg, Germany: International Potato Center, Lima, Peru, 161
Laufer B, Wilbur CM (1938) The American plant migration. Part I: the potato. Part I: the potato. Chicago: Field Museum of Natural History.
Lees AK, Wattier R, Shaw DS, Sullivan L, Williams NA and Cooke DEL (2006) Novel microsatellite markers for the analysis of Phytophthora infestans populations. Plant Pathology, 55:311-319
Li Y, Cooke DEL, Jacobsen E, van der Lee T (2013) Efficient multiplex simple sequence repeat genotyping of the oomycete plant pathogen Phytophthora infestans. J Microbiol Methods 92:316–322. https://doi.org/10.1016/j.mimet.2012.11.021
Martin FN, Zhang Y, Cooke DEL, Coffey MD, Grünwald NJ, Fry WE (2019) Insights into evolving global populations of Phytophthora infestans via new complementary mtDNA haplotype markers and nuclear SSRs. PLoS ONE 14:0208606
Mihretu E, Mohammod W, Kassa B, Lindqvist-Kreuze H (2020) Virulence Spectrum of Phytophthora infestans and Spatial Distribution of Physiological Races in Northwestern Ethiopia. Ethiopian Journal of Agricultural Sciences 30(1):69–85
Njoroge AW, Andersson B, Lees AK et al (2019a) Genotyping of Phytophthora infestans in Eastern Africa Reveals a Dominating Invasive European Lineage. Phytopathology® 109, 670–680
Njoroge AW, Andersson B, Yuen JE, Forbes GA (2019) Greater aggressiveness in the 2_A1 lineage of Phytophthora infestans may partially explain its rapid displacement of the US-1 lineage in east Africa. Plant Pathol 68:566–575
Nnadi NE, Datiri AM, Pam DB, Ngene AC, Okonkwo FO, Sullivan L, Cooke DEL (2019) First report of the EU_33_A2 clonal lineage of Phytophthora infestans causing late blight disease of potato in Nigeria. New Disease Reports 40, 20. https://doi.org/10.5197/j.2044-0588.2019.040.020
Pankhurst R (1964) Notes on the history of Ethiopian agriculture. Observer 7:210–240
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
R Core Team (2018) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria
Richard W, Simko I (2005) Resistance to Late Blight and Other Fungi. In: Razdan MK,, Matto AK, eds. Genetic Improvement of Solanaceous crops. Science publisher Inc, 451
Rosenberg NA (2003) distruct: a program for the graphical display of population structure: PROGRAM NOTE. Mol Ecol Notes 4:137–138
Saville AC, Martin MD, Ristaino JB (2016) Historic Late Blight Outbreaks Caused by a Widespread Dominant Lineage of Phytophthora infestans (Mont.) de Bary. PLoS One 11
Schiessendoppler E, Molnar O (2002) Characterization of Phytophthora infestans populations in sub-Saharan Africa as a basis for simulation modeling and integrated disease management. In: Lizárraga C ed. Proceedings of the Global Initiative on Late Blight Conference. Hamburg, Germany: International Potato Center, Lima, Peru, 140
Shannon CE (2001) A mathematical theory of communication. ACM SIGMOBILE Mob. Comput. Commun. Rev. 5:3-55
Shimelash D, Hussien T, Fininsa C, Forbes G, Yuen J (2016) Mitochondrial DNA assessment of Phytophthora infestans isolates from potato and tomato in Ethiopia reveals unexpected diversity. Curr Genet 62:657–667
Simpson EH (1949) Measurement of diversity. Nature, 163:688
Tesfahun F, Tsedeke A, Boris A (1985) A review of potato diseases research in Ethiopia. Tsedeke, A. A review of crop protection research in Ethiopia. Addis Ababa, IAR, Addis Ababa, pp 445–465
Yoshida K, Schuenemann VJ, Cano LM, Pais M, Mishra B et al (2013) The rise and fall of the Phytophthora infestans lineage that triggered the Irish potato famine. Life, 2:-00731
Acknowledgments
We thank David Cooke for providing the SSR marker profiles of US-1 and 2_A1 isolates. Amhara Agricultural Research Institute and Agricultural Growth Program II funded this work. The research done at CIP was undertaken as part of, and funded by, the CGIAR Research Program on Roots, Tubers and Bananas (RTB) and supported by CGIAR Trust Fund contributors https://www.cgiar.org/funders/.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mihretu, E., Izarra, M., Lindqvist-Kreuze, H. et al. Population structure of Phytophthora infestans (Mont.) de Bary in Ethiopia. J Plant Pathol 103, 759–767 (2021). https://doi.org/10.1007/s42161-021-00820-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42161-021-00820-6