Abstract
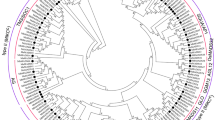
The BAHD family is involved in different biological roles in plants, including secondary metabolite synthesis, improving abiotic/biotic stress resistance, and influencing fruit quality. However, the knowledge about BAHD in banana, an important fruit crop, is limited. In this study, 46 banana BAHD genes (MaBAHDs) were identified and divided into four groups according to phylogenetic analysis. Most of the MaBAHD genes in the same group presented similar conserved motifs and genetic structures. MaBAHD genes have similar expression patterns in two banana varieties, and more genes showed high expressions in the roots. The comprehensive MaBAHD gene expression patterns obtained from two varieties of banana showed valuable information regarding their participation in fruit development, ripening, and response to abiotic/biotic stresses, suggesting that they play key roles in these processes. The systematic analysis of MaBAHD genes offered basic insight for further gene functional assays and potential applications in genetically improving banana cultivars.






Similar content being viewed by others
References
Pichersky E, Gang DR (2000) Genetics and biochemistry of secondary metabolites in plants: an evolutionary perspective. Trends Plant Sci 5(10):439–445. https://doi.org/10.1016/S1360-1385(00)01741-6
Pichersky E, Noel JP, Dudareva N (2006) Biosynthesis of plant volatiles: Nature’s diversity and ingenuity. Science 311(5762):808–811. https://doi.org/10.1126/science.1118510
St-Pierre B, De Luca V (2000) Chapter Nine – Evolution of acyltransferase genes: origin and diversification of the BAHD superfamily of acyltransferases involved in secondary metabolism. In: Romeo JT, Ibrahim R, Varin L, De Luca V (eds) Recent advances in phytochemistry, vol 34. Elsevier, pp 285–315. https://doi.org/10.1016/S0079-9920(00)80010-6
D’Auria JC (2006) Acyltransferases in plants: a good time to be BAHD. Curr Opin Plant Biol 9(3):331–340. https://doi.org/10.1016/j.pbi.2006.03.016
Ma X, Koepke J, Panjikar S, Fritzsch G, Stöckigt J (2005) Crystal structure of vinorine synthase, the first representative of the BAHD superfamily. J Biol Chem 280(14):13576–13583. https://doi.org/10.1074/jbc.M414508200
Tuominen LK, Johnson VE, Tsai C (2011) Differential phylogenetic expansions in BAHD acyltransferases across five angiosperm taxa and evidence of divergent expression among Populus paralogues. BMC Genomics 12(1):236. https://doi.org/10.1186/1471-2164-12-236
Liu C, Qiao X, Li Q, Zeng W, Wei S, Wang X, Chen Y, Wu X, Wu J, Yin H, Zhang S (2020) Genome-wide comparative analysis of the BAHD superfamily in seven Rosaceae species and expression analysis in pear (Pyrus bretschneideri). BMC Plant Biol 20(1):14. https://doi.org/10.1186/s12870-019-2230-z
Cheng A, Gou J, Yu X, Yang H, Fang X, Chen X, Liu C (2013) Characterization and ectopic expression of a Populus Hydroxyacid Hydroxycinnamoyltransferase. Mol Plant 6(6):1889–1903. https://doi.org/10.1093/mp/sst085
Schreiber L, Franke R, Hartmann K (2005) Wax and suberin development of native and wound periderm of potato (Solanum tuberosum L.) and its relation to peridermal transpiration. Planta 220(4):520–530. https://doi.org/10.1007/s00425-004-1364-9
Aggelis A, John I, Karvouni Z, Grierson D (1997) Characterization of two cDNA clones for mRNAs expressed during ripening of melon (Cucumis melo L.) fruits. Plant Mol Biol 33(2):313–322. https://doi.org/10.1023/A:1005701730598
Degenhardt B, Gimmler H (2000) Cell wall adaptations to multiple environmental stresses in maize roots. J Exp Bot 51(344):595–603. https://doi.org/10.1093/jexbot/51.344.595
Liu Q, Luo L, Zheng L (2018) Lignins: biosynthesis and biological functions in plants. Int J Mol Sci 19(2). https://doi.org/10.3390/ijms19020335
Panikashvili D, Shi JX, Schreiber L, Aharoni A (2009) The Arabidopsis DCR encoding a soluble BAHD acyltransferase is required for cutin polyester formation and seed hydration properties. Plant Physiol 151(4):1773–1789. https://doi.org/10.1104/pp.109.143388
Peng B, Yu M, Zhang B, Xu J, Ma R (2020) Differences in PpAAT1 activity in high- and low-aroma peach varieties affect γ-Decalactone production. Plant Physiol 182(4):2065–2080. https://doi.org/10.1104/pp.19.00964
Gou J, Yu X, Liu C (2009) A hydroxycinnamoyltransferase responsible for synthesizing suberin aromatics in Arabidopsis. Proc Natl Acad Sci 106(44):18855–18860. https://doi.org/10.1073/pnas.0905555106
Molina I, Bonaventure G, Ohlrogge J, Pollard M (2006) The lipid polyester composition of Arabidopsis thaliana and Brassica napus seeds. Phytochemistry 67(23):2597–2610. https://doi.org/10.1016/j.phytochem.2006.09.011
Molina I, Kosma D (2015) Role of HXXXD-motif/BAHD acyltransferases in the biosynthesis of extracellular lipids. Plant Cell Rep 34(4):587–601. https://doi.org/10.1007/s00299-014-1721-5
Molina I, Li-Beisson Y, Beisson F, Ohlrogge JB, Pollard M (2009) Identification of an Arabidopsis feruloyl-coenzyme A transferase required for suberin synthesis. Plant Physiol 151(3):1317–1328. https://doi.org/10.1104/pp.109.144907
Kosma DK, Molina I, Ohlrogge JB, Pollard M (2012) Identification of an Arabidopsis fatty alcohol:caffeoyl-Coenzyme A acyltransferase required for the synthesis of alkyl hydroxycinnamates in root waxes. Plant Physiol 160(1):237–248. https://doi.org/10.1104/pp.112.201822
Roh H, Jeong CW, Fujioka S, Kim YK, Lee S, Ahn JH, Choi YD, Lee JS (2012) Genetic evidence for the reduction of brassinosteroid levels by a BAHD acyltransferase-like protein in Arabidopsis. Plant Physiol 159(2):696–709. https://doi.org/10.1104/pp.112.197202
Wang M, Liu X, Wang R, Li W, Rodermel S, Yu F (2012) Overexpression of a putative Arabidopsis BAHD acyltransferase causes dwarfism that can be rescued by brassinosteroid. J Exp Bot 63(16):5787–5801. https://doi.org/10.1093/jxb/ers227
Zhang Z, Xu L (2018) Arabidopsis BRASSINOSTEROID INACTIVATOR2 is a typical BAHD acyltransferase involved in brassinosteroid homeostasis. J Exp Bot 69(8):1925–1941. https://doi.org/10.1093/jxb/ery057
Hu W, Wang L, Tie W, Yan Y, Ding Z, Liu J, Li M, Peng M, Xu B, Jin Z (2016) Genome-wide analyses of the bZIP family reveal their involvement in the development, ripening and abiotic stress response in banana. Sci Rep 6(1):30203. https://doi.org/10.1038/srep30203
Zheng Z, Qualley A, Fan B, Dudareva N, Chen Z (2009) An important role of a BAHD acyl transferase-like protein in plant innate immunity. Plant J 57(6):1040–1053. https://doi.org/10.1111/j.1365-313X.2008.03747.x
Chowdhury E, Choi B, Park SU, Lim HS, Bae H (2012) Transcriptional analysis of hydroxycinnamoyl transferase (HCT) in various tissues of Hibiscus cannabinus in response to abiotic stress conditions. Plant OMICS 5:305–313
Wei X, Lu W, Mao L, Han X, Wei X, Zhao X, Xia M, Xu C (2020) ABF2 and MYB transcription factors regulate feruloyl transferase FHT involved in ABA-mediated wound suberization of kiwifruit. J Exp Bot 71(1):305–317. https://doi.org/10.1093/jxb/erz430
Yu J, Hu SN, Wang J, Wong GKS, Li SG, Liu B, Deng YJ, Dai L, Zhou Y, Zhang XQ, Cao ML, Liu J, Sun JD, Tang JB, Chen YJ, Huang XB, Lin W, Ye C, Tong W, Cong LJ, Geng JN, Han YJ, Li L, Li W, Hu GQ, Huang XG, Li WJ, Li J, Liu ZW, Li L, Liu JP, Qi QH, Liu JS, Li L, Li T, Wang XG, Lu H, Wu TT, Zhu M, Ni PX, Han H, Dong W, Ren XY, Feng XL, Cui P, Li XR, Wang H, Xu X, Zhai WX, Xu Z, Zhang JS, He SJ, Zhang JG, Xu JC, Zhang KL, Zheng XW, Dong JH, Zeng WY, Tao L, Ye J, Tan J, Ren XD, Chen XW, He J, Liu DF, Tian W, Tian CG, Xia HG, Bao QY, Li G, Gao H, Cao T, Wang J, Zhao WM, Li P, Chen W, Wang XD, Zhang Y, Hu JF, Wang J, Liu S, Yang J, Zhang GY, Xiong YQ, Li ZJ, Mao L, Zhou CS, Zhu Z, Chen RS, Hao BL, Zheng WM, Chen SY, Guo W, Li GJ, Liu SQ, Tao M, Wang J, Zhu LH, Yuan LP, Yang HM (2002) A draft sequence of the rice genome (Oryza sativa L. ssp indica). Science 296(5565):79–92. https://doi.org/10.1126/science.1068037
Goff SA, Ricke D, Lan TH, Presting G, Wang RL, Dunn M, Glazebrook J, Sessions A, Oeller P, Varma H, Hadley D, Hutchinson D, Martin C, Katagiri F, Lange BM, Moughamer T, Xia Y, Budworth P, Zhong JP, Miguel T, Paszkowski U, Zhang SP, Colbert M, Sun WL, Chen LL, Cooper B, Park S, Wood TC, Mao L, Quail P, Wing R, Dean R, Yu YS, Zharkikh A, Shen R, Sahasrabudhe S, Thomas A, Cannings R, Gutin A, Pruss D, Reid J, Tavtigian S, Mitchell J, Eldredge G, Scholl T, Miller RM, Bhatnagar S, Adey N, Rubano T, Tusneem N, Robinson R, Feldhaus J, Macalma T, Oliphant A, Briggs S (2002) A draft sequence of the rice genome (Oryza sativa L. ssp japonica). Science 296(5565):92–100. https://doi.org/10.1126/science.1068275
Arabidopsis Genome I (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature (London) 408(6814):796–815
Tuskan G, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313(5793):1596–1604
Wang Z, Miao H, Liu J, Xu B, Yao X, Xu C, Zhao S, Fang X, Jia C, Wang J, Zhang J, Li J, Xu Y, Wang J, Ma W, Wu Z, Yu L, Yang Y, Liu C, Guo Y, Sun S, Baurens F, Martin G, Salmon F, Garsmeur O, Yahiaoui N, Hervouet C, Rouard M, Laboureau N, Habas R, Ricci S, Peng M, Guo A, Xie J, Li Y, Ding Z, Yan Y, Tie W, D’Hont A, Hu W, Jin Z (2019) Musa balbisiana genome reveals subgenome evolution and functional divergence. Nature Plants 5(8):810–821. https://doi.org/10.1038/s41477-019-0452-6
Hu W, Zuo J, Hou X, Yan Y, Wei Y, Liu J, Li M, Xu B, Jin Z (2015) The auxin response factor gene family in banana: genome-wide identification and expression analyses during development, ripening, and abiotic stress. Front Plant Sci 6:742–742. https://doi.org/10.3389/fpls.2015.00742
Günther CS, Chervin C, Marsh KB, Newcomb RD, Souleyre EJF (2011) Characterisation of two alcohol acyltransferases from kiwifruit (Actinidia spp.) reveals distinct substrate preferences. Phytochemistry 72(8):700–710. https://doi.org/10.1016/j.phytochem.2011.02.026
Schilmiller AL, Charbonneau AL, Las RL (2012) Identification of a BAHD acetyltransferase that produces protective acyl sugars in tomato trichomes. Proc Natl Acad Sci U S A 109(40):16377–16382. https://doi.org/10.1073/pnas.1207906109
Wang JQ, Yi zhong HE, Kun yang XU, Luo Y, Sheng L, Luo T, Liu H, Cheng YJ (2016) Characterization of mature fruit surface waxes of three cultivated Citrus species. Sci Agric Sin 49(10):1936–1945
Tzafrir I, Pena-Muralla R, Dickerman A, Berg M, Rogers R, Hutchens S, Sweeney TC, McElver J, Aux G, Patton D, Meinke D (2004) Identification of genes required for embryo development in Arabidopsis. Plant Physiol 135(3):1206–1220. https://doi.org/10.1104/pp.104.045179
Xi W, Zhang B, Shen J, Sun C, Xu C, Chen K (2012) Intermittent warming alleviated the loss of peach fruit aroma-related esters by regulation of AAT during cold storage. Postharvest Biol Technol 74:42–48. https://doi.org/10.1016/j.postharvbio.2012.07.003
Cao X, Xie K, Duan W, Zhu Y, Liu M, Chen K, Klee H, Zhang B (2019) Peach carboxylesterase PpCXE1 is associated with catabolism of volatile esters. J Agric Food Chem 67(18):5189–5196. https://doi.org/10.1021/acs.jafc.9b01166
Li D, Shen J, Wu T, Xu Y, Zong X, Li D, Shu H (2008) Overexpression of the apple alcohol acyltransferase gene alters the profile of volatile blends in transgenic tobacco leaves. Physiol Plant 134(3):394–402. https://doi.org/10.1111/j.1399-3054.2008.01152.x
Balbontín C, Gaete-Eastman C, Fuentes L, Figueroa CR, Herrera R, Manriquez D, Latché A, Pech JC, Moya-León MA (2010) VpAAT1, a gene encoding an alcohol acyltransferase, is involved in ester biosynthesis during ripening of mountain papaya fruit. J Agric Food Chem 58(8):5114–5121. https://doi.org/10.1021/jf904296c
Beekwilder J, Alvarez-Huerta M, Neef E, Verstappen FWA, Bouwmeester HJ, Aharoni A (2004) Functional characterization of enzymes forming volatile esters from strawberry and banana. Plant Physiol 135(4):1865–1878. https://doi.org/10.1104/pp.104.042580
Luo J, Nishiyama Y, Fuell C, Taguchi G, Elliott K, Hill L, Tanaka Y, Kitayama M, Yamazaki M, Bailey P, Parr A, Michael AJ, Saito K, Martin C (2007) Convergent evolution in the BAHD family of acyl transferases: identification and characterization of anthocyanin acyl transferases from Arabidopsis thaliana. Plant J 50(4):678–695. https://doi.org/10.1111/j.1365-313X.2007.03079.x
Sun C, Yang C, Tzen JTC (2018) Molecular identification and characterization of Hydroxycinnamoyl transferase in tea plants (Camellia sinensis L.). Int J Mol Sci 19(12):3938
Nadakuduti SS, Uebler JB, Liu X, Jones AD, Barry CS (2017) Characterization of Trichome-expressed BAHD acyltransferases in Petunia axillaris reveals distinct Acylsugar assembly mechanisms within the Solanaceae. Plant Physiol 175(1):36–50. https://doi.org/10.1104/pp.17.00538
Haslam TM, Gerelle WK, Graham SW, Kunst L (2017) The unique role of the ECERIFERUM2-LIKE clade of the BAHD acyltransferase superfamily in Cuticular wax metabolism. Plants (Basel, Switzerland) 6(2). https://doi.org/10.3390/plants6020023
Wang X, Guan Y, Zhang D, Dong X, Tian L, Qu LQ (2017) A β-Ketoacyl-CoA synthase is involved in Rice leaf Cuticular wax synthesis and requires a CER2-LIKE protein as a cofactor. Plant Physiol 173(2):944–955. https://doi.org/10.1104/pp.16.01527
Serra O, Figueras M, Franke R, Prat S, Molinas M (2010) Unraveling ferulate role in suberin and periderm biology by reverse genetics. Plant Signal Behav 5(8):953–958. https://doi.org/10.4161/psb.5.8.12405
Serra O, Hohn C, Franke R, Prat S, Molinas M, Figueras M (2010) A feruloyl transferase involved in the biosynthesis of suberin and suberin-associated wax is required for maturation and sealing properties of potato periderm. Plant J 62(2):277–290. https://doi.org/10.1111/j.1365-313X.2010.04144.x
Dwivany FM, Esyanti RR, Pratiwi AS, Zaskia H (2016) Expression study of Banana pathogenic resistance genes. HAYATI J Biosci 23(4):196–199. https://doi.org/10.1016/j.hjb.2016.06.007
Nicholson RL, Hammerschmidt R (1992) Phenolic compounds and their role in disease resistance. Annu Rev Phytopathol 30(1):369–389. https://doi.org/10.1146/annurev.py.30.090192.002101
Acknowledgements
This research was supported by the 2020 Research Program of Sanya Yazhou Bay Science and Technology City (SKJC-2020-02-002), the Central Public-Interest Scientific Institution Basal Research Fund for Chinese Academy of Tropical Agricultural Sciences (1630052016005, 1630052016006, and 1630052020006), and the Central Public-Interest Scientific Institution Basal Research Fund for Innovative Research Team Program of CATAS (17CXTD-28, 1630052017017).
Author information
Authors and Affiliations
Contributions
Jian Zeng, Wei Hu, and Zhiqiang Jin designed the research; Yun Xu and Weiwei Tie performed the research; Yan Yan, Biyu Xu, Juhua Liu, Meiying Li, and Jinghao Yang analyzed the data; Jian Zeng, Yun Xu, Weiwei Tie, and Wei Hu wrote the paper; Jian Zeng, Wei Hu, and Zhiqiang Jin reviewed the paper. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, Y., Tie, W., Yan, Y. et al. Identification and expression of the BAHD family during development, ripening, and stress response in banana. Mol Biol Rep 48, 1127–1138 (2021). https://doi.org/10.1007/s11033-020-06132-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-06132-9