Abstract
Soil microbial community structure is altered by petroleum contamination in response to compound toxicity and degradation. Understanding the relation between petroleum contamination and soil microbial community structure is crucial to determine the amenability of contaminated soils to bacterial- and fungal-aided remediation. To understand how petroleum contamination and soil physicochemical properties jointly shaped the microbial structure of soils from different oilfields, high-throughput sequencing of 16S and ITS amplicons were used to evaluate the shifts of microbial communities in the petroleum-contaminated soils in Ughelli East (UE), Utorogu (UT), and Ughelli West (UW) oilfields located in Delta State, Nigeria. The results showed 1515 bacteria and 919 fungal average OTU number, and community richness and diversity, trending as AL > UT > UW > UE and AL > UW > UT > UE for bacteria, and AL > UW > UT > UE and UW > UT > AL > UE for fungi, respectively. The bacterial taxa KCM-B-112, unclassified Saccharibacteria, unclassified Rhizobiales, Desulfurellaceae, and Acidobacteriaceae and fungal Trichocomaceae, unclassified Ascomycota, unclassified Sporidiobolales, and unclassified Fungi were found to be the dominant families in petroleum-contaminated soils. Redundancy analysis (RDA) and Spearman’s correlation analysis revealed that total carbon (TC), electric conductivity (EC), pH, and moisture content (MO) were the major drivers of bacterial and fungal communities, respectively. Gas chromatography-mass spectrophotometer (GC-MS) analysis exhibited that the differences in C7–C10, C11–C16, and C12–C29 compounds in the crude oil composition and soil MO content jointly constituted the microbial community variance among the contaminated soils. This study revealed the bacterial and fungal communities responsible for the biodegradation of petroleum contamination from these oilfields, which could serve as biomarkers to monitor oil spill site restoration within these areas. Further studies on these contaminated sites could offer useful insights into other contributing factors such as heavy metals.
Similar content being viewed by others
Introduction
Soil petroleum contamination is hugely prevalent in and around oilfields in Nigeria. It poses a severe danger to the environment when it is not adequately degraded, hence the need for explicit knowledge of microbial communities in contaminated sites and their dynamic roles in transforming the surrounding ecosystem (Mafiana et al. 2020). The most common causes of land oil spills are operational failures in and around oilfields, rupture of transit pipelines and storage facilities, and theft (Babatunde et al. 2018). A few of the numerous challenging ordeals suffered by host communities from poorly remediated spill sites are hampered farming activities (Albert et al. 2018) and spill extension into streams and groundwater (Adesipo et al. 2020).
In an earlier investigation by the United Nations Environmental Programme (UNEP) in 2011, over 69 contaminated sites in 200 locations of Ogoniland in the Niger Delta region were observed to have toxic compounds of benzene, toluene, ethylbenzene, xylene (BTEX), methyl tert-butyl ether (MTBE), and other hydrocarbons of lower molecular weight (LMW). These toxic compounds from poorly remediated sites were leaching into surface and groundwater, vegetation, and sediments (UNEP 2011). According to reports, failed remediation sites that had adopted the Remediation by Natural Attenuation (RENA) approach relied on the degradation capabilities of naturally inhabiting microorganisms by tilling and windrow stimulation. Besides, 10 out of 15 remediated sites still had contaminant residues leaching into groundwater (UNEP 2011; Sam and Zabbey 2018). Studies on the ecological hazards from spilled areas and poorly remediated sites have recorded bioaccumulation in fish (Nematopalaemon hastatus) amounting to 2585.62 mg kg−1 (Akinola et al. 2019) and an increase in neonatal and infant mortality by 38.3 deaths per 1000 live births (Bruederle and Hodler 2019) among other numerous health risks associated with soil petroleum contamination (Chinedu and Chukwuemeka 2018; Iwegbue et al. 2020).
The microbial community structure and function are products of its surrounding ecosystem, altered by the hazardous epoxides and toxic effect of petroleum on soil microorganisms (Sam and Zabbey 2018). Several studies on spill site remediation approaches have observed the predominance of different microbial phyla such as Proteobacteria, Acidobacteria, Firmicutes, Fusobacteria, Ascomycota, and Basidiomycota that play significant roles in hydrocarbon degradation in petroleum-contaminated soils in Nigeria (Obi et al. 2016; Ezekoye et al. 2018a, b; Obieze et al. 2019a; Okafor et al. 2021). While some petroleum compounds are easily degradable, most of their other components are toxic to microorganisms and could cause sporulation or total dormancy of a non-tolerant group and significantly alter the soil’s microbial community structure (Oyetibo et al. 2017; Ventorino et al. 2018). The relative abundance of hydrocarbon-degrading microorganisms often differs from one contaminated site to another within the Niger Delta region (Mafiana et al. 2020), hence the crucial need to understand the microbial community structure specific to contaminated sites in and around oilfields, the soil physicochemical properties, and the nature of contaminant petroleum hydrocarbon.
Environmental factors such as temperature, soil moisture, pH, nutrients, organic matter, and nature of contaminants directly affect the microbial population (Brockett et al. 2012; Siles and Margesin 2018; Jiang et al. 2019; Wang et al. 2020). Significant microbial correlation to soil moisture, total organic carbon (TC), carbon/nitrogen ratio (C/N), and pH in petroleum-contaminated soils have been reported at varying degrees (Cui et al. 2020; Trejos-Delgado et al. 2020), as well as successive transitions linked to long-term soil complexity (Zegeye et al. 2019; Zhao et al. 2019) and different spatial or temporal distributions relative to soil nutrients (Kivlin and Hawkes 2016). Besides biological and particularly microbiological disturbance, petroleum contamination negatively alters soil physicochemical properties mostly at the surface and decreases with increasing soil depth (Sheng et al. 2021). Some notable soil properties altered include a decreased wettability (Błońska et al. 2016; Rodríguez-Rodríguez et al. 2016), hydraulic conductivity (Devatha et al. 2019), increased hydrophobicity, and CO2 efflux (Hewelke et al. 2018). Microbial tolerance to changes in soil physicochemical properties differ, bacteria strains are better known as effective degraders of PAHs, but fungi are better at surviving harsh environments. Hence, fungi could also be potential stimulators of bacterial growth in polluted environments where bacteria are limited (Oyetibo et al. 2017).
Microbes with hydrocarbonoclastic capabilities are not easily isolated using standard nutrient media in laboratory conditions and a culture-dependent method, which could only recover approximately 0.3% of its population (Chikere et al. 2019). Because of the limitations of biochemical techniques in microbial identification and characterization (Adzitey et al. 2013), as well as bacterial Gram variabilities (Ramamurthy et al. 2014; Franco-Duarte et al. 2019), in recent decades, a significant number of studies have suggested that 16S and 18S rRNA amplicon sequencing are more reliable for exploring soil microbiomes and the prevailing geophysical and chemical properties of soil (Mukherjee et al. 2017; Abia et al. 2018; Chikere et al. 2019; Kang et al. 2020). Thus, the objectives of this study were as follows: (i) to investigate the microbial community structure and diversities in petroleum-contaminated soils of Utorogu (UT), Ughelli East (UE), and Ughelli West (UW) oilfields in Delta State using 16S and 18S rRNA amplicon sequencing and (ii) to assay the difference in component composition of crude oil from these oilfields using a gas chromatography-mass spectrometer (GC-MS) analysis. We also assessed the soil physicochemical properties and their correlation to the microbial communities as a potential guide for bioremediation indicators.
Materials and methods
Characteristics of the study sites
The contaminated oilfield sites comprised UE, UT, and UW in Delta State and the non-contaminated soil sample from Alagomeji (AL) in Lagos State, Nigeria, a non-oil-producing area. The UE oil flow station has been in operation since 1959. Its average ambient temperature is 26.4 °C and 2.768 mm of precipitation. The UT oilfield exploration began in 1964. The average ambient temperature is 26.7 °C, and it receives 2768 mm of precipitation (Climate-Data.org 2019). The UW oilfield exploration began in 1960. The average temperature of the location is 26.6 °C, and it receives 2.638 mm of precipitation. The climate conditions are classified as tropical monsoon climate, and they have loamy wetland soils (Chibuzo 2016). The average temperature of the sample site AL is 27.0 °C, and it receives 1675 mm of precipitation. We also characterize its soil as a loamy wetland area with little stones (Obiefuna et al. 2013).
Soil sample collection
Soil samples were collected in August 2018 from four different sampling locations: UE (5° 32′ 17.8″ N, 5° 53′ 46.8″ E), UT (5° 25′ 55.6″ N, 5° 53′ 02.5″ E), UW (5° 32′ 17.8″ N, 5° 53′ 46.8″ E), and AL (6° 29′ 48.4″ N, 3° 22′ 36.5″ E) in triplicates to a total of 12 samples. The distances between sampling locations are 11.89 km (UT and UW), 5.29 km (UW and UE), 10.80 km (UE and UT), and 301.21 km (AL and UT) (Fig. 1). Each sample was randomly collected using an aseptic sterile auger and 50-mL tubes at 10 cm depth, and the collection points were > 5 m apart. A portion of the collected samples was immediately measured for moisture content and DNA extraction, and the remaining portions were air-dried for further physicochemical property analysis.
Soil physicochemical analysis
The soil samples were oven-dried at 105 °C to constant weight for their moisture contents to be determined gravimetrically. The soil pH was determined using a 1:4 (w/v) soil-deionized water and analyzed with a standard calibrated pH probe. The electric conductivity (EC) value was measured using a digital conductivity instrument (Leici DDSJ-308A) in a soil-deionized water slurry of 1:5. The total carbon (TC) and total nitrogen (TN) in the soil were determined using an elemental analyzer (Elementar Vario-EL, Germany).
GC-MS analysis of crude oil samples
The crude oil samples were prepared using the dichloromethane solvent in a high-resolution Agilent GC-MS 7000 C Triple Quad machine fitted with an HP-5MS (Agilent Technologies, Inc. USA), 30 m ×250 μm, 0.25 μm chromatographic column. The oven temperature was initially kept at 30 °C for 21 min and finally increased to 310 °C at 15 °C·min−1 for 9 min. The injection volume was at 1 μL, and the detector temperature was set at 350 °C with helium carrier gas of > 99.99% purity at the flow rate of 1 mL·min−1. The detector signal was integrated and analyzed in the Mass Hunter computer program through a chromatographic server that generated corresponding gas peak heights and details of specific components. The properties of the component organic compounds detected were further characterized using the National Library of Medicine database (PubChem 2020). For sample quantification and procedural recovery, the internal and surrogate standard of 16 PAH (Ultra Scientific PM-610) solution containing anthracene (AN), acenaphthylene (ACY), acenaphthene (ACE), benzo[a]anthracene (BaA), benzo[g,h,i]perylene (BP), benzo[b]fluoranthene (BbF), benzo[k]fluoranthene (BkF), benzo[a]pyrene (BaP), fluorene (FL), fluoranthene (FLU), pyrene (PY), chrysene (CH), indeno[1,2,3-cd]pyrene (IP), dibenzo[a,h]anthracene (DA), naphthalene (NA), and phenanthrene (PH) was used as calibration standards (Mills et al. 1999).
DNA extraction and rRNA amplicon sequencing
The genomic DNA was extracted from 5 g of each soil sample using the Fast DNA® SPIN Kit for soil and MP Biomedicals Fast Prep® instrument according to the manufacturer’s protocol (MP Biomedicals 2007). The bacterial 16S rRNA genes were amplified using universal primer 338F_806R 5′-ACTCCTACGGGAGGCAGCA-3′ and 5′-GGACTACHVGGGTWTCTAAT-3′ (Bakke et al. 2011). For fungi, ITS1F_ITS2R primer 5′-CTTGGTCATTTAGAGGAAGTAA-3′ and 5′-GCTGCGTTCTTCATCGATGC-3′ were used (White et al. 1990). Finally, sequencing was performed on an Illumina MiSeq PE300 platform (Illumina, USA) in Majorbio Co., Ltd. (Shanghai, China). The processed sequences were aligned and deposited on the National Center for Biotechnology Information (NCBI) with the sequence read archive accession numbers for bacteria and fungi being PRJNA593923 and PRJNA594087.
Statistical analysis
The species annotation and genomic assessment were assigned to their specific categories of operational taxonomic units (OTUs) according to the USEARCH (version 7.0 http://drive5.com/usearch/) software platform (Fan et al. 2018). Non-repeated sequences were extracted to reduce redundancy, while single sequences without repetition and chimeras were removed during clustering to obtain the representative OTU at 97% similarity. The bacteria and fungi OTUs were classified into their exact taxonomy based on the SILVA database (Release 128 http://www.arb-silva.de) and Ribosomal Database Project (https://rdp.cme.msu.edu/) and UNITE database for environmental ITS identification (Release 8.0 http://unite.ut.ee/). The Bayesian taxonomic algorithm at 97% similarity classified the community compositions as phylum, class, order, family, genus, and species.
Based on the number and abundance of OTUs, community diversity (Shannon and Simpson) and richness (Sobs, Chao, and ACE) were estimated (Hugerth and Andersson 2017). The redundancy analysis index (RDA) was calculated, and permutational multivariate analysis of variance (PERMANOVA) and environmental correlation analysis were performed using the Majorbio I-Sanger Cloud Platform (Majorbio Co., Ltd., Shanghai, China). The sample distance matrix was clustered for hierarchical analysis to understand the similarities and differences between the sample’s OTU composition and their structural differences (Borem et al. 2014). The unweighted pair-group method with arithmetic mean (UPGMA) algorithm was used to determine the degree of microbial evolution in various samples, while a weighted UniFrac analysis further quantified the lineage variation between samples (Nye et al. 2006; Clarke et al. 2006).
Bray-Curtis distance matrix algorithm:
SA,i = number of sequences in the ith taxon in the A sample and SB in the ith taxon in the B sample.
where the number of ith OTU represents samples 1 and 2; sqrt = square root.
Results
Dissimilarities in the soil physicochemical properties
The most distinct soil pH range (6.8–7.7) across samples was presented in UE and AL, which was relatively narrow compared to water content, TN, TC, and EC, with significant differences (Table 1). The same EC value at 48.2 μS cm−1 was detected in UT and UW soils. The AL soil presented the highest EC value at 93.5 μS cm−1, while the UE soil presented the lowest EC at 24.2 μS cm−1. The highest TC at 10.4 mg g−1 was recorded in the UE soil. The moisture content (MO) in UT soil at 41.47% was higher than the UE and UW soils by 22% and 23.9%, respectively. The AL soil MO significantly differed from UT, UE, and UW soils, possibly due to the riverine terrain where the oilfields are located.
Petroleum organic components of UT, UE, and UE oil samples
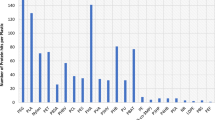
The GC-MS analysis of crude oil samples from the UE, UT, and UW oilfields showed highly complex mixtures of constituent aromatic, paraffinic, and naphthenic (cycloparaffin) hydrocarbon compounds, which were further characterized based on the National Library of Medicine database (Table S1). A total of 293 peaks (Fig. S1) detected in the crude oil samples (UT: 69, UW: 94, and UE: 130) were related to paraffin, naphthene, and aromatic hydrocarbon compounds (Table S1). Compounds of low molecular weight (LMW) (C7–C10) and complexity, such as cyclohexane, ethylbenzene, xylene, benzene, indene, octahydro-, cis-, 1-ethyl-3-methylcyclohexane, and menthane, were unique to UT. Medium molecular weight (MMW) compounds (C11–C15) were most observed in UE and UW, notably decahydro-1,1,4a,5,6-pentamethylnaphthalene highest peak areas by 80.20% and 93.80%, respectively. Among the MMW compounds, 4-(2,2,6-trimethyl-bicyclo(4.1.0) hept-1-yl)-butan-2-one (C14H24O) and 2-pentanone (C15H26O) were observed only in UE, while naphthalenone (C15H22O2) was absent in UW and 3-buten-2-one specific to UW. High molecular weight (HMW) compounds (C16–C29) were disproportionately found among the three samples but most significant in UE by 4–36% peak area. The properties of these petroleum compounds could determine their fate in the contaminated soils, which can be categorized into three principal fractions, namely the volatile (C7–C10), highly extractible (C11–C16), and low extractible (C17–C19). The LMW compounds are highly volatile, less persistent in soil, and likely to repel less soil moisture, but highly toxic due to bioavailability compared to the MMW and HMW compounds.
Variation in the number of OTUs and α-diversity of microbial communities in the soils
The clustering analysis revealed a total average of 1515 bacterial and 919 fungal OTUs from the soil samples. The highest occurring bacterial (1830) and fungal (1184) OTU numbers were found in UW soil and the lowest (1120 and 747) OTU numbers, respectively, in UE soil (Fig. S2). The bacterial richness (Chao1 index) and diversity (Shannon) exhibited the trend AL > UT > UW > UE and AL > UW > UT > UE, respectively, while the fungal richness (Chao1 index) and diversity (Shannon) was AL > UW > UT > UE and UW > UT > AL > UE, respectively (Table S2). The results suggest that the impact of petroleum toxicity reduced the richness and diversity of soil bacterial communities and the richness of fungal communities but had a lesser impact on fungal diversity.
Variation in bacterial and fungal community structure in the soils
In total, 42 bacterial phyla and 360 families were identified from the soils. The predominant bacterial phyla account for 82% of the total community abundance. These include Proteobacteria, Actinobacteria, Acidobacteria, and Chloroflexi by 44%, 17%, 12%, and 9%, respectively (Fig. 2a). Proteobacteria was the most abundant phylum (55.4–42.2%) in the contaminated soils due to its high aromatic degradative activity compared with the non-contaminated AL soil where Proteobacteria (27.2%) and Acidobacteria (8.4%) were markedly decreased, suggesting that petroleum contamination was significantly a driving factor. Bacteroidetes, Firmicutes, Fusobacteria, and Planctomycetes were also highly abundant in the UT soil and twofold higher than in UE and UW soils. The relative abundance differences suggest that these phyla could have been increased by higher moisture content and unique petroleum compounds associated with alkanes, benzene, ethylbenzene, hexane, toluene, and xylene in UT oilfields.
The predominant bacterial families in the contaminated soils were unclassified Rhizobiales (14.4%), KCM-B-112 (12.8%), and Acidobacteriaceae (8.5%) in UT, UE, and UW soils, respectively (Fig. 2b). Other dominant petroleum-degrading families notably present across contaminated soils were Desulfurellaceae, Rhodospirillaceae, and Anaerolineaceae except KCM-B-112, a hydrocarbon-degrading family whose absence in UT is not entirely apparent but could be associated with BTEX toxicity or excess MO content in the soil. Additionally, Rhodospirillaceae and Rhodocyclaceae are medium- and long-chain hydrocarbon degraders predominant in UE and distributed across UT and UW. The above distribution suggests that the relative abundance of some predominant oil-degrading biomarker families across soils were in response to petroleum contamination and soil physicochemical properties.
In total, 8 fungal phyla, 196 families, and 419 genera were identified in the soil samples. Ascomycota (92.3%) predominated the AL non-contaminated soil but was reduced to 41.5%, 55.9%, and 54.2% in UE, UT, and UW soils, respectively (Fig. 3a). Unclassified Fungi phylum predominated UE soil but was significantly decreased in UT and UW soils, where the abundance of Basidiomycota, Rozellomycota, and Chytridiomycota was notably increased. Trichocomaceae (38%) dominated AL soil at the family level, while unclassified Fungi dominated accounted for 56.2%, 27.9%, and 40.4% in UE, UT, and UW, respectively. Chaetomiaceae, Nectriaceae, unclassified Hypocreales, and Sporormiaceae were markedly decreased in the contaminated soils compared to AL. Notable petroleum-degrading fungal families in the contaminated soils include unclassified Sporidiobolales (10.6%) and Montagnulaceae (5.2%) in UT, unclassified Chaetothyriales (7%) and Pleosporaceae (6%) in UW, and unclassified Hypocreales (5%) in UE.
Aspergillus predominated the non-contaminated AL soil at the genus level while unclassified Fungi predominated the contaminated soils. The relative abundance of Aspergillus, Talaromyces, unclassified Sporidiobolales, and Paraconiothyrium was markedly more in UT than other contaminated soils (Fig. 3b). Among the dominant hydrocarbon-degrading fungi, Aspergillus is one of the well-reported genera known to survive a wide range of environmental conditions; hence, its relatively low abundance in other contaminated soils is unclear but could be linked to other inhibiting factors such as toxic heavy metals associated with crude oil which was not tested in this study. However, high soil MO and TN content in UT seem to have influenced most other dominant genera, except unclassified Fungi, whose abundance does not correlate with any of the above factors.
The responses of microbial communities to soil properties
Principal coordinate analysis (PCoA) at the genus level was used to visually evaluate microbial community responses to soil petroleum contamination and physicochemical properties based on the weighted normalized UniFrac distance statistics. The contaminated soils’ bacterial communities showed a tight eclipse matching distinct from the clustered non-contaminated AL soil (Fig. 4a). Among the contaminated soils, a certain degree of similarity among UE1, UE2, and UW3 on the positive axis is most notable. Overall, both coordinates explain 59.1% of the total bacterial community variations. The community similarities in response to petroleum contamination shown by the distance matrix are most significant between UE and UW soil by a cross-connection, suggesting a correlation with some MMW and HMW compound degrading bacterial genera. The fungal communities also showed similar tightly cluttered AL soils on the negative PC2 axis with 15.49% total variation distinct from the contaminated soils on the positive PC1 axis, representing 64.3% of the total variations. The contaminated soils formed a crossed eclipse formation of UE2, UE1, UT2, and UW1 on the positive PC1 axis. This clustered formation shows a stronger interconnected relationship within the communities in contrast with UT1, UW3, and UT3 (Fig. 4b). Overall, both principal components explain 79.79% of the total fungal genera community variations. Interestingly, the UW distribution pattern covers a narrowly extensive range across UT and UE, while UT was widely dispersed. Comparatively, the fungal genera coordinates are slightly more interconnected than those of the bacterial genera coordinates, notably a wider spread of UT and UW soils, suggesting a distinct response to petroleum contamination between the bacterial and fungal communities.
Principal coordinate analysis (PCoA) at the genus level. a Bacteria. b Fungi. The X-axis and Y-axis represent the two selected main coordinate axes, and the percentage value represents the main coordinate axis’ interpretation of the difference in sample composition. The closer the two sample points are, the more similar the genera composition
Correlation between bacterial and fungal community relationship to soil physicochemical properties
To evaluate the relationship between the relative abundance of community and environmental parameters, RDA was performed. In Fig. 5a, the bacterial class RDA by 62.56% (RDA2: 20.70% and RDA1: 41.86%) explained emerging acute angles between MO, TN, and TC positive correlation with Alpha-, Delta-, Beta-, and Gamma-proteobacteria, Acidobacteria, Anaerolineae, and unclassified Saccharibacteria, indicating a significant synergetic effect of these variables on the relative abundance of the bacterial community, while EC and pH jointly impacted on Actinobacteria, Thermomicrobia, and KD4-96 communities. The fungal family by 73.3% (RDA2: 11.52% and RDA1: 61.78%) (Fig. 5b) community variances explained the emerging acute angles between MO, TN, and TC positive correlation with unclassified Sporidiobolales, Pleosporaceae, Montagnulaceae, and unclassified Fungi, while EC most impacted on Trichomaceae, Chaetomiaceae, and Nectriaceae. These bacterial and fungal community abundance variances across soils correspond to 41.5% MO in UT and 10.4 mg·g−1 TC in UE, 93.47 μS cm−1 in AL, and 0.33 mg·g−1 in UT. Besides, the most abundant phylum Proteobacteria was positively correlated with all parameters except EC. In contrast, soil pH representing a crucial soil parameter was not strongly correlated positively with any fungal family.
Redundancy analysis (RDA) between bacterial communities at the class level (a) and fungal communities at the family level (b) and soil physicochemical parameters. Different shapes or colors represent sample groups under different environmental conditions. The red color arrow depicts the physicochemical parameters, and the green arrow depicts microbial taxa. The arrow length represents the degree of influence, while the angle of deviation between the parameters represents a positive or negative correlation. Sharp angle: positive correlation; obtuse angle: negative correlation; right angle: no correlation
A Spearman heatmap correlation analysis was used to reveal further 30 bacterial and fungal genera responses to specific soil physicochemical parameters. The top cluster of positive color gradient revealed 47% and 43% positively correlated genera with pH and EC, respectively (Fig. 6a). The most significantly positively correlated genera were unclassified Acidobacteria and unclassified Acidimicrobiales to pH and EC, respectively, while Anaerolinea was most negatively correlated with EC. The bottom-positive color gradient cluster revealed 53%, 50%, and 43% genera positively correlated with TC, TN, and MO, respectively, which are most statistically correlated with Anaerolinea (TC and TN) and unclassified Rhizobiales (MO). In the fungal genera, a cluster of color gradients at the bottom represents positively correlated genera with MO, TN, and TC by 40%, 33%, and 40%, respectively (Fig. 6b). Paraconiothyrium and Pyrenochaetopsis were the most statistically positively correlated genera with all three parameters. The top cluster of positively correlated fungal genera was 37% pH and 57% EC. Unclassified Chaetothyriales and Aspergillus most statistically correlated with EC, but none of the parameters was statistically significant with pH. In contrast, TC and EC were the most significant parameters to bacteria, but dissimilarly, fungi showed a relatively higher positive correlation with MO content than bacteria.
Hierarchical clustering heatmap analysis of 30 genera of bacteria (a) and fungi (b) and their correlation significance with physicochemical parameters of pH, EC, TN, TC, and MO. Hierarchical clustering on the top shows a classified relationship between the parameters, while on the side depicts the relationship with microbial taxa. The color intensity depicts the genera relative abundance using the legend indicator, which ranges from negative blue color coefficients to red color ones. *0.01 < P ≤ 0.05, **< P ≤ 0.001, ***P ≤ 0.001
Discussion
Extensive researches have focused on petroleum bioremediation using the RENA approach and various isolated consortia from oil spilled sites in Nigeria. However, no previous study has reported a comparative analysis of microbial community structure in petroleum-contaminated soils across these three different oilfields in Nigeria. This study analyzed the impact of soil petroleum contamination on bacterial and fungal communities across Ughelli East (UE), Ughelli West (UW), and Utorogu (UT) oilfields. The results characterized the similarities and differences in petroleum component compounds of these oilfields and soil physicochemical properties that jointly shaped the soil microbial communities.
Although these oilfields have existed since the 1960s, the soil samples were severely re-contaminated periodically with fresh crude oil within 1–12 months before sampling. Previous studies on freshly contaminated soils in operational oilfield sites have shown that indigenous microbes within a short time are adapted to petroleum contamination and activated for biodegradation rapidly (Kaufmann et al. 2004; Liu et al. 2020). Microbial statistical correlation to soil MO, pH, EC, TC, TN, and petroleum compounds revealed that differences in MO, TN, and the contaminant crude oil most jointly shaped the soil microbial communities. A previous study on petroleum-contaminated soils has reported that crude oil contamination decreases soil moisture content due to its water repellency effect (Liang et al. 2012). In the present study, a similar decrease in soil moisture content was observed in the UE soil with the most contaminant MMW and HMW compounds by GC-MS analysis. Compared to other contaminated soils, the higher concentration of these compounds could be responsible for the lower MO content and consequently decreased microbial richness and diversity in UE soil. This effect of petroleum concentration also agrees with earlier studies that reported a strong correlation between soil moisture and oil contamination due to water repellency, hydrophobicity, and decreased soil porosity and permeability (Wang et al. 2013; Hewelke et al. 2018; Devatha et al. 2019), although previous studies have shown that petroleum contamination increases soil pH up to 8.0 (Wang et al. 2013) and could be associated with increasing soil nitrogen from fertilizer application in and around agricultural land use areas (Wang et al. 2018; Shen et al. 2019). Considering the high level of nitrogen input from agricultural NPK and gas flaring in the Niger Delta region (Osuji et al. 2006; Giwa et al. 2019), it might be challenging to ascertain the level of petroleum contamination impact on the soil’s pH. However, in this study, the UW soil with the highest soil pH (7.6) and the lowest TN (0.08) level had higher microbial diversity and richness than other contaminated soils, which is in agreement with an earlier study by Obieze et al. (2019b) that a near-neutral pH in artisanal crude oil refining sites in the Niger Delta region is suitable to improve the diversity of microbial communities (Obieze et al. 2019b). A study on soil TN correlation with microbial diversity by Roesch et al. (2007) has also reported a higher microbial diversity in forest soil with a relatively lower pH than the counterpart agricultural soils with higher pH values. This factor was attributed to commercial NPK fertilizer use (Staley et al. 2018), increase in soil NH4+, which positively correlates with soil acidity and C:N ratio leading to a buildup of organic C (Zhalnina et al. 2014).
Our present results showed that microbial community linage across contaminated soils is slightly varied at the family and genus level. Beta- and Gamma-proteobacteria dominated the contaminated population by 30% abundance. Consequently, their predominance decreased the population of Acidobacteria, Gemmatimonadetes, Actinobacteria, Planctomycetes, and Chloroflexi phyla, and Delta-proteobacteria class in UW and UT. These lower phyla have been reported as mainly composed of slow-growing organisms and are thus dominated by Alpha-, Beta-, and Gamma-proteobacteria, which are regarded as fast-growing r-strategists (Maron et al. 2018; Kurm et al. 2019). They are also regarded as obligate anaerobes of sulfate reducers and early-stage hydrocarbon degraders (Viñas et al. 2005; Prince et al. 2018). Like most previous studies, Ascomycota was the predominant known petroleum-degrading fungi phylum. Ezekoye et al. (2018b) recorded > 90% relative abundance in Ikarama community spill site remediation in Bayelsa state, Nigeria. Liu et al. (2019) recorded 58.96–99.67% in Tianjin Dagang oilfield in eastern China. Some similarities and differences with previous studies include the absence of Nocardioidaceae, Sphingomonadaceae, Flavobacteriaceae, and Methylobacteriaceae families, which were significantly dominated in some petroleum-contaminated soils (Mukherjee et al. 2017; Feng et al. 2020). Conversely, these families were absent or not significantly present in this study; however, the reason remains unclear. Obligate anaerobe genera, such as Anaeromyxobacter, Ruminiclostridium, and Geobacter, have been reported with low survival tendencies in hydrocarbon-contaminated soils (Brzeszcz et al. 2020) but were significantly represented across all contaminated soils in this study. These differences could be attributed to a long-term adaptation to these oilfields or autochthonously site-specific. Yang et al. (2019) reported the presence of Nocardioidaceae in a petroleum-contaminated soil in Qinghai-Tibetan Plateau with a high petroleum degradation capability and was attributed to adaptation by gene transformation through crude oil contamination. Likewise, in the present study, Fusobacteriaceae, Prevotellaceae, and Bacteroidales S24-7 in UT are human and animal gastrointestinal tract or fecal bacteria families that have not been previously reported in petroleum-contaminated soil. Therefore, speculate that these intestinal microbes may have gained resistance to toxic hydrocarbons due to the presence of animals or indiscriminate sewage disposal associated with poor human hygiene (Adserias-Garriga et al. 2017). The presence of unclassified Acidimicrobiales in contaminated soils in this study has been reported earlier by Kämpfer (2010), who noted that some genera belonging to family Acidimicrobiales have not been reported to perform hydrocarbon degradation but should be strictly observed. Furthermore, Peng et al. (2015) identified this family in an oil-contaminated soil through a high-throughput sequencing and linked its abundance to soil saline alkalinity.
From studies, not all bacteria and fungi taxon can be affiliated to a specific hydrocarbon compound or soil property; however, relationships between microbial communities and pollutant load can be drawn (Spini et al. 2018). A previous study on petroleum-contaminated soil in Shanxi province oilfields has reported that moisture increase by 15% from the initial 5% increased the relative abundance of Firmicutes from 4.6 to 29.7% while decreased Proteobacteria phylum (Liu et al. 2020). Similar population dynamics in hydrocarbon-degrading phyla were observed in this present study. The UT soil with extreme MO content supported the growth of rare taxa in the absence of microbial competition and increased bacteria richness, notably the abundance of Firmicutes in UT. In contrast, the UW soil with intermediate soil MO promoted microbial diversity, which can be attributed to a competitively driven population. This result is in line with previous studies (Erlandson et al. 2016; Bickel and Or 2020). TC and TN were also essential factors limiting microbial population in petroleum-contaminated soils. In a previous study, Feng et al. (2020) noted that total petroleum hydrocarbon (TPH), TC, and TN were the most significant factors affecting the bacterial communities in petroleum-contaminated Dagang oilfield soils but varied at different soil levels. In the present study, TC most significantly correlated positively and negatively with bacterial genera compared to fungal genera (Fig. 6). This factor could be related to Kaufmann et al. (2004), which reported that the optimal C:N for bacteria cell growth in a petroleum-contaminated soil is between 20:1 and 50:1. The influence of soil pH (6.8–7.7) on the fungi community variation was not statistically significant with any genus, unlike EC, TC, and MO. The reason for this result may be that the soil pH range is relatively narrow. Nevertheless, most fungi could be well adapted to a wide range of pH values sufficient to obtain optimal growth (Padmavathi et al. 2003; Rousk et al. 2010; Borowik et al. 2017). For example, a survival test on a wide pH range showed that some zoosporic fungi (Blastocladiomycota and Chytridiomycota) isolated from soil in Australia could grow in a pH range between 2.9 and 11.2 (Gleason et al. 2010).
The present study showed that soil pH most statistically correlated strongly with unclassified Acidobacteria. Similarly, Acidobacteria has been reported to have a significant positive correlation with soil pH (Conradie and Jacobs 2020) and survive under a broad range of TC sources (De Castro et al. 2013) and depleted oxygen (Pankratov et al. 2012). This broad survival range implies the reason for its significant relative abundance across all soils. Soil EC is closely related to salinity and strongly correlates with Actinobacteria abundance (O’Brien et al. 2019), which affirms its significantly higher abundance in non-contaminated soils in this study. This result is also similar to Peng et al. (2015), which reported that Actinobacteria was most abundant in contaminated soils across the three oil-mined wells in the 1960s, 1970s, and 1990s Daqing oilfields.
The GS-MS analysis indicated that the oilfields situated within the same region produced slightly different petroleum compounds. The observed differences in hydrocarbon components support a previous study that closely situated oilfields in an area of the same stratum, or oil-bearing sands may produce dissimilar crude oils (International Agency for Research on Cancer 1989). While complex high molecular compounds are ultimately recalcitrant in contaminated soils (Feyzi et al. 2020), studies have shown that straight-chain alkanes are more easily oxidized than branched-chain and cycloalkanes, so also the degradation rates of medium-chain alkanes (C5–C15) significantly vary from those > C20 (Jia et al. 2017). Rhodocyclaceae have a high correlation with the degradation of mid-chain C14, C16, and C18 hydrocarbons, while Rhodospirillaceae most correlated with only C14 mid-chain alkanes (Liu et al. 2018). In the present study, both families are more abundant in UE, while Rhodospirillaceae in UT and Rhodocyclaceae in UW. A toxic compound similar to 4H-cyclopropa[5′,6′]benz[1′,2′:7,8]azuleno[5,6-b]oxiren-4-one,8-acetyloxy was most significant in UE and UT, accounting for 53% and 74%, respectively. This compound has been reported as a phytochemical component extracted from a plant with an antimicrobial effect on the Bacillus, Pseudomonas, Escherichia bacterial genera, and Candida, Trichoderma, and Aspergillus fungal genera (Abdul-Hafeez et al. 2016; Fierascu et al. 2017). Speculatively, it could have a similar toxic effect on these soil microbial communities and should be investigated in further studies. The highest degradation percentage of benzene, ethylbenzene, and xylene compounds have been observed with Aspergillus (Asemoloye et al. 2020; Usman et al. 2020a, b), which suggest its relative abundance by 7% more in UT than other contaminated soils. However, its relatively low abundance in other contaminated soils could only be associated with possible heavy metal contamination (Zeng et al. 2020). Penicillium was significantly abundant in contaminated soils but showed no significant correlation to any environmental factor. Hence, we suggest its strong affinity for long-chain n-alkane (C20–C29) hydrocarbon degradation by adaptation to petroleum contamination as reported by Barnes et al. (2018) and Yamada-Onodera et al. (2002).
Our study shows a shift in the relative abundance of some bacterial families and fungal genera, which might be useful indicators to assess soil petroleum contamination. Moreover, the distribution patterns of Proteobacteria, Firmicutes, Ascomycota, and Aspergillus communities might be useful microbial parameters in the evaluation of petroleum-contaminated soil in and around oilfields. Further studies on these oilfields would be required to determine specific roles of some newly identified genera (Acidimicrobiales, Fusobacteriaceae, Prevotellaceae, and Bacteroidales S24-7) in petroleum hydrocarbon degradation. Furthermore, our study implies that petroleum-contaminated soils did not support more bacterial and fungal community richness but only fungal diversity.
Conclusion
This study characterized bacterial and fungal diversity in petroleum-contaminated soils from three different oilfields in Nigeria. The impact of petroleum contamination on the microbial communities was observed to have caused a decline in microbial richness and diversity distinct from a non-contaminated site, except fungal diversity. Petroleum contamination favored fast-growing autochthonous petroleum-degrading class, such as Alpha-, Beta-, and Gamma-proteobacteria, while some obligate genera were observed to have undergone long-term adaptation to hydrocarbon toxicity. Proteobacteria and Ascomycota phyla played a significant role in the degradation of hydrocarbons in these contaminated sites. Excess soil MO content in contaminated soil enriched rare bacteria OTUs in the absence of competition, but intermediate soil MO increased OTU diversity. In contrast, bacteria and fungi respond differently to excess soil MO and petroleum compounds. The LMW compounds in UT different from UE and UW crude oil are partly responsible for the significant disproportion in some vital bacterial phylotypes. These compounds include cyclohexane, ethylbenzene, p-xylene, benzene, 1H-indene, and m-menthane with low complexity and biodegradation half-life values. Conversely, benzene, toluene, ethylbenzene, and xylene (BTEX) compounds are known for groundwater contamination due to easier penetration through soil and bioavailability. Thus, it should be considered a potential threat to human health when leached into the ground and nearby streams and rivers. This analysis of the microbial community richness and diversity impacted by oil pollution is limited to and around these oilfields. At the same time, it provides useful insight for an in-depth study on microbial isolation and biodegradation capabilities to improve spill site remediation application.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding authors on reasonable request.
References
Abdul-hafeez EY, Mahmoud A, Ibrahim O (2016) Antibacterial activities and phytochemical screening of Alhagi pseudalhagi. Assiut J Agric Sci 46:33–47. https://doi.org/10.21608/ajas.2016.530
Abia ALK, Alisoltani A, Keshri J, Ubomba-Jaswa E (2018) Metagenomic analysis of the bacterial communities and their functional profiles in water and sediments of the Apies River, South Africa, as a function of land use. Sci Total Environ 616–617:326–334. https://doi.org/10.1016/j.scitotenv.2017.10.322
Adesipo AA, Freese D, Nwadinigwe AO (2020) Prospects of in-situ remediation of crude oil contaminated lands in Nigeria. Sci African 8:e00403. https://doi.org/10.1016/j.sciaf.2020.e00403
Adserias-Garriga J, Hernández M, Quijada NM, Rodríguez Lázaro D, Steadman D, Garcia-Gil J (2017) Daily thanatomicrobiome changes in soil as an approach of postmortem interval estimation: an ecological perspective. Forensic Sci Int 278:388–395. https://doi.org/10.1016/j.forsciint.2017.07.017
Adzitey F, Huda N, Ali GRR (2013) Molecular techniques for detecting and typing of bacteria, advantages and application to foodborne pathogens isolated from ducks. 3 Biotech 3:97–107. https://doi.org/10.1007/s13205-012-0074-4
Akinola JO, Olawusi-Peters OO, Akpambang VOE (2019) Ecological hazards of total petroleum hydrocarbon in brackish water white shrimp Nematopalaemon hastatus (Aurivillus 1898). Egypt J Aquat Res 45:205–210. https://doi.org/10.1016/j.ejar.2019.07.004
Albert ON, Amaratunga D, Haigh RP (2018) Evaluation of the impacts of oil spill disaster on communities and its influence on restiveness in Niger Delta, Nigeria. Procedia Eng 212:1054–1061. https://doi.org/10.1016/J.PROENG.2018.01.136
Asemoloye MD, Tosi S, Daccò C, Wang X, Xu S, Marchisio MA, Gao W, Jonathan SG, Pecoraro L (2020) Hydrocarbon degradation and enzyme activities of Aspergillus oryzae and Mucor irregularis isolated from nigerian crude oil-polluted sites. Microorganisms 8:1–19. https://doi.org/10.3390/microorganisms8121912
Babatunde BB, Zabbey N, Vincent-Akpu IF, Mekuleyi GO (2018) Bunkering activities in Nigerian waters and their eco-economic consequences. Polit Ecol Oil Gas Act Niger Aquat Ecosyst:439–446. https://doi.org/10.1016/B978-0-12-809399-3.00026-4
Bakke I, De Schryver P, Boon N, Vadstein O (2011) PCR-based community structure studies of bacteria associated with eukaryotic organisms: a simple PCR strategy to avoid co-amplification of eukaryotic DNA. J Microbiol Methods 84:349–351. https://doi.org/10.1016/j.mimet.2010.12.015
Barnes NM, Khodse VB, Lotlikar NP, Meena RM, Damare SR (2018) Bioremediation potential of hydrocarbon-utilizing fungi from select marine niches of India. 3. Biotech 8:1–10. https://doi.org/10.1007/s13205-017-1043-8
Bickel S, Or D (2020) Soil bacterial diversity mediated by microscale aqueous-phase processes across biomes. Nat Commun 11:1–9. https://doi.org/10.1038/s41467-019-13966-w
Błońska E, Lasota J, Szuszkiewicz M, Łukasik A, Klamerus-Iwan A (2016) Assessment of forest soil contamination in Krakow surroundings in relation to the type of stand. Environ Earth Sci 75. https://doi.org/10.1007/s12665-016-6005-7
Borem A, Fritsche-Neto R, Cruz CD et al (2014) Biometrics applied to molecular analysis in genetic diversity. Biotechnol Plant Breed. https://doi.org/10.1016/B978-0-12-418672-9.00003-9
Borowik A, Wyszkowska J, Oszust K (2017) Functional diversity of fungal communities in soil contaminated with diesel oil. Front Microbiol 8:1–11. https://doi.org/10.3389/fmicb.2017.01862
Brockett BFT, Prescott CE, Grayston SJ (2012) Soil moisture is the major factor influencing microbial community structure and enzyme activities across seven biogeoclimatic zones in western Canada. Soil Biol Biochem 44:9–20. https://doi.org/10.1016/J.SOILBIO.2011.09.003
Bruederle A, Hodler R (2019) Effect of oil spills on infant mortality in Nigeria. Proc Natl Acad Sci U S A 116:5467–5471. https://doi.org/10.1073/pnas.1818303116
Brzeszcz J, Kapusta P, Steliga T (2020) Hydrocarbon removal by two differently developed microbial inoculants and comparing their actions. Molecules 25:1–23. https://doi.org/10.3390/molecules25030661
Chibuzo EB (2016) Alteration of organic matter by gas flaring activity: a case study of Utorogu Community in Niger-Delta , Nigeria. In: J. Environ. Earth Sci. Vol 6, No 9 https://www.iiste.org/Journals/index.php/JEES/article/view/32922. Accessed 17 Jan 2020
Chikere CB, Mordi IJ, Chikere BO, Selvarajan R, Ashafa TO, Obieze CC (2019) Comparative metagenomics and functional profiling of crude oil-polluted soils in Bodo West Community, Ogoni, with other sites of varying pollution history. Ann Microbiol 69:495–513. https://doi.org/10.1007/s13213-019-1438-3
Chinedu E, Chukwuemeka CK (2018) Oil spillage and heavy metals toxicity risk in the Niger Delta, Nigeria. J Heal Pollut 8:180905. https://doi.org/10.5696/2156-9614-8.19.180905
Clarke KR, Somerfield PJ, Chapman MG (2006) On resemblance measures for ecological studies, including taxonomic dissimilarities and a zero-adjusted Bray–Curtis coefficient for denuded assemblages. J Exp Mar Biol Ecol 330:55–80. https://doi.org/10.1016/J.JEMBE.2005.12.017
Climate-Data.org (2019) Warri Climate-Data. In: Climate-Data.org. https://en.climate-data.org/location/24741/. Accessed 19 Sep 2018
Conradie T, Jacobs K (2020) Seasonal and agricultural response of Acidobacteria present in two fynbos rhizosphere soils. Diversity 12. https://doi.org/10.3390/d12070277
Cui JQ, He QS, Liu MH, Chen H, Sun MB, Wen JP (2020) Comparative study on different remediation strategies applied in petroleum-contaminated soils. Int J Environ Res Public Health 17:1–17. https://doi.org/10.3390/ijerph17051606
De Castro VHL, Schroeder LF, Quirino BF et al (2013) Acidobacteria from oligotrophic soil from the Cerrado can grow in a wide range of carbon source concentrations. Can J Microbiol 59:746–753. https://doi.org/10.1139/cjm-2013-0331
Devatha CP, Vishnu Vishal A, Purna Chandra Rao J (2019) Investigation of physical and chemical characteristics on soil due to crude oil contamination and its remediation. Appl Water Sci 9:1–10. https://doi.org/10.1007/s13201-019-0970-4
Erlandson SR, Savage JA, Cavender-Bares JM, Peay KG (2016) Soil moisture and chemistry influence diversity of ectomycorrhizal fungal communities associating with willow along an hydrologic gradient. FEMS Microbiol Ecol 92:1–9. https://doi.org/10.1093/femsec/fiv148
Ezekoye C, Chikere G, Okpokwasili C (2018a) Field metagenomics of bacterial community involved in bioremediation of crude cil-polluted soil. J Bioremediat Biodegrad 09:09. https://doi.org/10.4172/2155-6199.1000449
Ezekoye CC, Chikere CB, Okpokwasili GC (2018b) Fungal diversity associated with crude oil-impacted soil undergoing in-situ bioremediation. Sustain Chem Pharm 10:148–152. https://doi.org/10.1016/j.scp.2018.11.003
Fan XY, Gao JF, Pan KL, Li DC, Dai HH, Li X (2018) Functional genera, potential pathogens and predicted antibiotic resistance genes in 16 full-scale wastewater treatment plants treating different types of wastewater. Bioresour Technol 268:97–106. https://doi.org/10.1016/j.biortech.2018.07.118
Feng X, Liu Z, Jia X, Lu W (2020) Distribution of bacterial communities in petroleum-contaminated soils from the Dagang oilfield, China. Trans Tianjin Univ 26:22–32. https://doi.org/10.1007/s12209-019-00226-7
Feyzi H, Chorom M, Bagheri G (2020) Urease activity and microbial biomass of carbon in hydrocarbon contaminated soils. A case study of cheshmeh-khosh oil field, Iran. Ecotoxicol Environ Saf 199:110664. https://doi.org/10.1016/j.ecoenv.2020.110664
Fierascu I, Georgiev MI, Ortan A, Fierascu RC, Avramescu SM, Ionescu D, Sutan A, Brinzan A, Ditu LM (2017) Phyto-mediated metallic nano-architectures via Melissa officinalis L.: synthesis, characterization and biological properties. Sci Rep 7:1–3. https://doi.org/10.1038/s41598-017-12804-7
Franco-Duarte R, Černáková L, Kadam S, S. Kaushik K, Salehi B, Bevilacqua A, Corbo MR, Antolak H, Dybka-Stępień K, Leszczewicz M, Relison Tintino S, Alexandrino de Souza VC, Sharifi-Rad J, Melo Coutinho HD, Martins N, Rodrigues CF (2019) Advances in chemical and biological methods to identify microorganisms—from past to present. Microorganisms 7:130. https://doi.org/10.3390/microorganisms7050130
Giwa SO, Nwaokocha CN, Kuye SI, Adama KO (2019) Gas flaring attendant impacts of criteria and particulate pollutants: a case of Niger Delta region of Nigeria. J King Saud Univ - Eng Sci 31:209–217. https://doi.org/10.1016/J.JKSUES.2017.04.003
Gleason FH, Daynes CN, McGee PA (2010) Some zoosporic fungi can grow and survive within a wide pH range. Fungal Ecol 3:31–37. https://doi.org/10.1016/j.funeco.2009.05.004
Hewelke E, Szatyłowicz J, Hewelke P, Gnatowski T, Aghalarov R (2018) The impact of diesel oil pollution on the hydrophobicity and CO2 efflux of forest soils. Water Air Soil Pollut 229. https://doi.org/10.1007/s11270-018-3720-6
Hugerth LW, Andersson AF (2017) Analysing microbial community composition through amplicon sequencing: from sampling to hypothesis testing. Front Microbiol 8:1–22. https://doi.org/10.3389/fmicb.2017.01561
International Agency for Research on Cancer (1989) International agency for research on cancer iarc monographs on the evaluation of carcinogenic risks to humans. Iarc Monogr Eval Carcinog Risks To Humansarc Monogr Eval Carcinog Risks To Humans 45:1–390. https://doi.org/10.1002/food.19940380335
Iwegbue CMA, Bebenimibo E, Tesi GO, Egobueze FE, Martincigh BS (2020) Spatial characteristics and risk assessment of polychlorinated biphenyls in surficial sediments around crude oil production facilities in the Escravos River Basin, Niger Delta, Nigeria. Mar Pollut Bull 159:111462. https://doi.org/10.1016/j.marpolbul.2020.111462
Jia J, Zong S, Hu L, Shi S, Zhai X, Wang B, Li G, Zhang D (2017) The dynamic change of microbial communities in crude oil-contaminated soils from oil fields in China. Soil Sediment Contam 26:171–183. https://doi.org/10.1080/15320383.2017.1264923
Jiang B, Adebayo A, Jia J, Xing Y, Deng S, Guo L, Liang Y, Zhang D (2019) Impacts of heavy metals and soil properties at a Nigerian e-waste site on soil microbial community. J Hazard Mater 362:187–195. https://doi.org/10.1016/j.jhazmat.2018.08.060
Kämpfer P (2010) Actinobacteria. Handb Hydrocarb Lipid Microbiol. https://doi.org/10.1007/978-3-540-77587-4
Kang XH, Leng Y, Macdonald OM et al (2020) The seasonal changes of core bacterial community decide sewage purification in sub-plateau municipal sewage treatment plants. Bioprocess Biosyst Eng 43:1609–1617. https://doi.org/10.1007/s00449-020-02352-2
Kaufmann K, Christophersen M, Buttler A, Harms H, Höhener P (2004) Microbial community response to petroleum hydrocarbon contamination in the unsaturated zone at the experimental field site Værløse, Denmark. FEMS Microbiol Ecol 48:387–399. https://doi.org/10.1016/j.femsec.2004.02.011
Kivlin SN, Hawkes CV (2016) Temporal and spatial variation of soil bacteria richness, composition, and function in a neotropical rainforest. PLoS One 11:1–17. https://doi.org/10.1371/journal.pone.0159131
Kurm V, Van Der Putten WH, Hol WHG (2019) Cultivation-success of rare soil bacteria is not influenced by incubation time and growth medium. PLoS One 14:1–14. https://doi.org/10.1371/journal.pone.0210073
Liang Y, Zhang X, Wang J, Li G (2012) Spatial variations of hydrocarbon contamination and soil properties in oil exploring fields across China. J Hazard Mater 241–242:371–378. https://doi.org/10.1016/j.jhazmat.2012.09.055
Liu Y, Ding A, Sun Y, Xia X, Zhang D (2018) Impacts of n-alkane concentration on soil bacterial community structure and alkane monooxygenase genes abundance during bioremediation processes. Front Environ Sci Eng 12:1–13. https://doi.org/10.1007/s11783-018-1064-5
Liu Q, Tang J, Liu X, Song B, Zhen M, Ashbolt NJ (2019) Vertical response of microbial community and degrading genes to petroleum hydrocarbon contamination in saline alkaline soil. J Environ Sci (China) 81:80–92. https://doi.org/10.1016/j.jes.2019.02.001
Liu H, Gao H, Wu M, Ma C, Wu J, Ye X (2020) Distribution characteristics of bacterial communities and hydrocarbon degradation dynamics during the remediation of petroleum-contaminated soil by enhancing moisture content. Microb Ecol 80:202–211. https://doi.org/10.1007/s00248-019-01476-7
Mafiana MO, Bashiru MD, Erhunmwunsee F, Dirisu CG, Li SW (2020) An insight into the current oil spills and on-site bioremediation approaches to contaminated sites in Nigeria. Environ Sci Pollut Res 28:4073–4094. https://doi.org/10.1007/s11356-020-11533-1
Maron PA, Sarr A, Kaisermann A, Lévêque J, Mathieu O, Guigue J, Karimi B, Bernard L, Dequiedt S, Terrat S, Chabbi A, Ranjard L (2018) High microbial diversity promotes soil ecosystem functioning. Appl Environ Microbiol 84:1–13. https://doi.org/10.1128/AEM.02738-17
Mills MA, McDonald TJ, Bonner JS et al (1999) Method for quantifying the fate of petroleum in the environment. Chemosphere 39:2563–2582. https://doi.org/10.1016/S0045-6535(99)00163-0
Mukherjee A, Chettri B, Langpoklakpam JS, Basak P, Prasad A, Mukherjee AK, Bhattacharyya M, Singh AK, Chattopadhyay D (2017) Bioinformatic approaches including predictive metagenomic profiling reveal characteristics of bacterial response to petroleum hydrocarbon contamination in diverse environments. Sci Rep 7:1108. https://doi.org/10.1038/s41598-017-01126-3
Nye TMW, Liò P, Gilks WR (2006) A novel algorithm and web-based tool for comparing two alternative phylogenetic trees. Bioinformatics 22:117–119. https://doi.org/10.1093/bioinformatics/bti720
O’Brien FJM, Almaraz M, Foster MA, Hill AF, Huber DP, King EK, Langford H, Lowe MA, Mickan BS, Miller VS, Moore OW, Mathes F, Gleeson D, Leopold M (2019) Soil salinity and pH drive soil bacterial community composition and diversity along a lateritic slope in the Avon river critical zone observatory, Western Australia. Front Microbiol 10:1486. https://doi.org/10.3389/fmicb.2019.01486
Obi CC, Adebusoye SA, Ugoji EO, Ilori MO, Amund OO, Hickey WJ (2016) Microbial communities in sediments of Lagos Lagoon, Nigeria: elucidation of community structure and potential impacts of contamination by municipal and industrial wastes. Front Microbiol 7:1213. https://doi.org/10.3389/fmicb.2016.01213
Obiefuna JN, Nwilo PC, Atagbaza AO, Okolie CJ (2013) Spatial changes in the wetlands of Lagos/Lekki Lagoons of Lagos, Nigeria. J Sustain Dev 6. https://doi.org/10.5539/jsd.v6n7p123
Obieze CC, Chikere CB, Adeleke R, Akaranta O (2019a) Formulation and evaluation of slow-release fertilizer from agricultural and industrial wastes for remediation of crude oil-polluted soils. Soc Pet Eng - SPE Niger Annu Int Conf Exhib 2019, NAIC 2019. https://doi.org/10.2118/198815-MS
Obieze CC, Chikere CB, Adeleke R, Akaranta O (2019b) Formulation and evaluation of slow-release fertilizer from agricultural and industrial wastes for remediation of crude oil-polluted soils. In: Society of Petroleum Engineers - SPE Nigeria Annual International Conference and Exhibition 2019, NAIC 2019. OnePetro
Okafor CP, Udemang NL, Chikere CB, Akaranta O, Ntushelo K (2021) Indigenous microbial strains as bioresource for remediation of chronically polluted Niger Delta soils. Sci African 11:e00682. https://doi.org/10.1016/j.sciaf.2020.e00682
Osuji LC, Egbuson EJ, Ojinnaka CM (2006) Assessment and treatment of hydrocarbon inundated soils using inorganic nutrient (N-P-K) supplements: II. A case study of Eneka oil spillage in Niger Delta, Nigeria. Environ Monit Assess 115:265–278. https://doi.org/10.1007/s10661-006-6552-6
Oyetibo GO, Miyauchi K, Huang Y, Chien MF, Ilori MO, Amund OO, Endo G (2017) Biotechnological remedies for the estuarine environment polluted with heavy metals and persistent organic pollutants. Int Biodeterior Biodegradation 119:614–625. https://doi.org/10.1016/J.IBIOD.2016.10.005
Padmavathi J, Uma Devi K, Uma Maheswara Rao C (2003) The optimum and tolerance pH range is correlated to colonial morphology in isolates of the entomopathogenic fungus Beauveria bassiana—a potential biopesticide. World J Microbiol Biotechnol 19:469–477. https://doi.org/10.1023/A:1025151000398
Pankratov TA, Kirsanova LA, Kaparullina EN, Kevbrin VV, Dedysh SN (2012) Telmatobacter bradus gen. nov., sp. nov., a cellulolytic facultative anaerobe from subdivision 1 of the Acidobacteria, and emended description of Acidobacterium capsulatum Kishimoto et al. 1991. Int J Syst Evol Microbiol 62:430–437. https://doi.org/10.1099/ijs.0.029629-0
Peng M, Zi X, Wang Q (2015) Bacterial community diversity of oil-contaminated soils assessed by high throughput sequencing of 16S rRNA genes. Int J Environ Res Public Health 12:12002–12015. https://doi.org/10.3390/ijerph121012002
Prince RC, Amande TJ, McGenity TJ (2018) Prokaryotic hydrocarbon degraders. Taxon Genomics Ecophysiol Hydrocarb Microbes 1–41. https://doi.org/10.1007/978-3-319-60053-6_15-1
PubChem US National Library of Medicine. In: Natl. Cent. Biotechnol. Inf. https://pubchem.ncbi.nlm.nih.gov/. Accessed 14 Sep 2020
Ramamurthy T, Ghosh A, Pazhani GP, Shinoda S (2014) Current perspectives on viable but non-culturable (VBNC) pathogenic bacteria. Front Public Health 2:1–9. https://doi.org/10.3389/fpubh.2014.00103
Rodríguez-Rodríguez N, Rivera-Cruz MC, Trujillo-Narcía A, Almaráz-Suárez JJ, Salgado-García S (2016) Spatial distribution of oil and biostimulation through the rhizosphere of Leersia hexandra in degraded soil. Water Air Soil Pollut 227:227. https://doi.org/10.1007/s11270-016-3030-9
Roesch LFW, Fulthorpe RR, Riva A, Casella G, Hadwin AKM, Kent AD, Daroub SH, Camargo FAO, Farmerie WG, Triplett EW (2007) Pyrosequencing enumerates and contrasts soil microbial diversity. Int Soc Microb Ecol 1:283–290. https://doi.org/10.1038/ismej.2007.53
Rousk J, Bååth E, Brookes PC, Lauber CL, Lozupone C, Caporaso JG, Knight R, Fierer N (2010) Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J 4:1340–1351. https://doi.org/10.1038/ismej.2010.58
Sam K, Zabbey N (2018) Contaminated land and wetland remediation in Nigeria: opportunities for sustainable livelihood creation. Sci Total Environ 639:1560–1573. https://doi.org/10.1016/j.scitotenv.2018.05.266
Shen C, Shi Y, Fan K, He JS, Adams JM, Ge Y, Chu H (2019) Soil pH dominates elevational diversity pattern for bacteria in high elevation alkaline soils on the Tibetan Plateau. FEMS Microbiol Ecol 95:1–9. https://doi.org/10.1093/femsec/fiz003
Sheng Y, Liu Y, Yang J, Dong H, Liu B, Zhang H, Li A, Wei Y, Li G, Zhang D (2021) History of petroleum disturbance triggering the depth-resolved assembly process of microbial communities in the vadose zone. J Hazard Mater 402:124060. https://doi.org/10.1016/j.jhazmat.2020.124060
Siles JA, Margesin R (2018) Insights into microbial communities mediating the bioremediation of hydrocarbon-contaminated soil from an Alpine former military site. Appl Microbiol Biotechnol 102:4409–4421. https://doi.org/10.1007/s00253-018-8932-6
Spini G, Spina F, Poli A, Blieux AL, Regnier T, Gramellini C, Varese GC, Puglisi E (2018) Molecular and microbiological insights on the enrichment procedures for the isolation of petroleum degrading bacteria and fungi. Front Microbiol 9:2543. https://doi.org/10.3389/fmicb.2018.02543
Staley C, Breuillin-Sessoms F, Wang P, Kaiser T, Venterea RT, Sadowsky MJ (2018) Urea amendment decreases microbial diversity and selects for specific nitrifying strains in eight contrasting agricultural soils. Front Microbiol 9:1–13. https://doi.org/10.3389/fmicb.2018.00634
Trejos-Delgado C, Cadavid-Restrepo GE, Hormaza-Anaguano A et al (2020) Oil bioremediation in a tropical contaminated soil using a reactor. An Acad Bras Cienc 92:e20181396. https://doi.org/10.1590/0001-3765202020181396
UNEP (2011) Environmental assessment of Ogoniland site-specific fact sheets. In: United Nations Environ. Prot. Agency. https://wedocs.unep.org/handle/20.500.11822/23016. Accessed 16 Sep 2020
Usman N, Atta HI, Tijjani MB (2020a) Biodegradation studies of benzene, toluene, ethylbenzene and xylene ( BTEX ) compounds by Gliocladium sp. and Aspergillus terreus. J Appl Sci Environ Manag 24:1063–1069. https://doi.org/10.4314/jasem.v24i6.19
Usman N, Tijjani MB, Atta HI (2020b) Isolation and identification of BTEX-utilizing Fungi from soil polluted with petroleum hydrocarbons. In: Biosci. Res. J. http://ojs.klobexjournals.com/index.php/brj/article/view/458. Accessed 14 Sep 2020
Ventorino V, Pascale A, Adamo P, Rocco C, Fiorentino N, Mori M, Faraco V, Pepe O, Fagnano M (2018) Comparative assessment of autochthonous bacterial and fungal communities and microbial biomarkers of polluted agricultural soils of the Terra dei Fuochi. Sci Rep 8:1–13. https://doi.org/10.1038/s41598-018-32688-5
Viñas M, Sabaté J, Espuny MJ, Solanas AM (2005) Bacterial community dynamics and polycyclic aromatic hydrocarbon degradation during bioremediation of heavily creosote-contaminated soil. Appl Environ Microbiol 71:7008–7018. https://doi.org/10.1128/AEM.71.11.7008-7018.2005
Wang X, Guan X, Zhang X, Xiang S, Zhang R, Liu M (2020) Microbial communities in petroleum-contaminated seasonally frozen soil and their response to temperature changes. Chemosphere 258:127375. https://doi.org/10.1016/j.chemosphere.2020.127375
Wang Y, Feng J, Lin Q, Lyu X, Wang X, Wang G (2013) Effects of crude oil contamination on soil physical and chemical properties in momoge wetland of China. Chin Geogr Sci 23:708–715. https://doi.org/10.1007/s11769-013-0641-6
Wang C, Liu D, Bai E (2018) Decreasing soil microbial diversity is associated with decreasing microbial biomass under nitrogen addition. Soil Biol Biochem 120:126–133. https://doi.org/10.1016/j.soilbio.2018.02.003
Zegeye EK, Brislawn CJ, Farris Y, Fansler SJ, Hofmockel KS, Jansson JK, Wright AT, Graham EB, Naylor D, McClure RS, Bernstein HC (2019) Selection, succession, and stabilization of soil microbial consortia. Am Soc Microbiol 4:1–13. https://doi.org/10.1128/msystems.00055-19
White TJ, Bruns TD, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal genes form phylogenetics. In: PCR Protoc. A Guid. to methods Appl. https://nature.berkeley.edu/brunslab/papers/white1990.pdfAccessed 16 Oct 2020
Yamada-Onodera K, Mukumoto H, Katsuyama Y, Tani Y (2002) Degradation of long-chain alkanes by a polyethylene-degrading fungus, Penicillium simplicissimum YK. Enzym Microb Technol 30:828–831. https://doi.org/10.1016/S0141-0229(02)00065-0
Yang R, Liu G, Chen T, Li S, An L, Zhang G, Li G, Chang S, Zhang W, Chen X, Wu X, Zhang B (2019) Characterization of the genome of a Nocardia strain isolated from soils in the Qinghai-Tibetan Plateau that specifically degrades crude oil and of this biodegradation. Genomics 111:356–366. https://doi.org/10.1016/j.ygeno.2018.02.010
Zhao C, Long J, Liao H, Zheng C, Li J, Liu L, Zhang M (2019) Dynamics of soil microbial communities following vegetation succession in a karst mountain ecosystem, Southwest China. Sci Rep 9:1–10. https://doi.org/10.1038/s41598-018-36886-z
Zeng XY, Li SW, Leng Y, Kang XH (2020) Structural and functional responses of bacterial and fungal communities to multiple heavy metal exposure in arid loess. Sci Total Environ 723:138081. https://doi.org/10.1016/j.scitotenv.2020.138081
Zhalnina K, Dias R, de Quadros PD, Davis-Richardson A, Camargo FAO, Clark IM, McGrath SP, Hirsch PR, Triplett EW (2014) Soil pH determines microbial diversity and composition in the park grass experiment. Microb Ecol 69:395–406. https://doi.org/10.1007/s00248-014-0530-2
Acknowledgements
The authors are thankful to the Chinese Scholarship Council (CSC) of China and the School of International Studies, Lanzhou Jiaotong University: The Nigerian Petroleum Development Corporation (NPDC) OML 34 team of sample collection assistance.
Funding
This work was supported through funding from the National Natural Science Foundation of China (31760110 and 31560121).
Author information
Authors and Affiliations
Contributions
MOM: conceived and designed the analysis; collecting of the data and analysis tools; experimental procedure; writing of the paper; XHK: collecting the data and analysis tools; analysis of data; YL: experimental procedure; LFH: experimental procedure; SWL: supervision, conceptualization, revision of manuscript
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible editor: Robert Duran
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(PDF 481 kb)
Rights and permissions
About this article
Cite this article
Mafiana, M.O., Kang, XH., Leng, Y. et al. Petroleum contamination significantly changes soil microbial communities in three oilfield locations in Delta State, Nigeria. Environ Sci Pollut Res 28, 31447–31461 (2021). https://doi.org/10.1007/s11356-021-12955-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-021-12955-1