Abstract
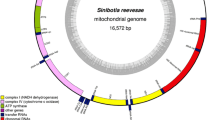
Gnathopogon herzensteini is a small-sized fish endemic to the upper reaches of the Jialing River and the Han River. Wild populations of G. herzensteini have declined sharply during the past few decades and the phylogeny of this species has not yet fully resolved. Notably, the relationship between Gnathopogon and Gobiocypris genera still remains controversial. Scientific progress is hampered by the scarcity of molecular data for G. herzensteini. To address this problem, the complete mitochondrial genome of G. herzensteini were sequenced and characterised, and used to analyse its phylogenetic position within the subfamily Gobioninae. The genome is a circular molecule of 16,597 bases, consisting of 13 protein-coding genes, 2 rRNA genes, 22 tRNA genes, and a control region (D-loop). The nucleotide composition exhibited A/T bias: A 30.0%, T 27.0%, G 17.2%, and C 25.8%. Phylogenetic tree revealed that Gobiocypris clustered within the genus Gnathopogon, thus rendering it paraphyletic. The clade of genus Gnathopogon was closely related to Coreoleuciscus, with strong statistical support. These data are a stepping stone towards the understanding of genetics and phylogeny of G. herzensteini.



Similar content being viewed by others
Abbreviations
- AIC:
-
Akaike information criterion
- ATP6:
-
ATP synthase F0 subunit 6
- ATP8:
-
ATP synthase F0 subunit 8
- bp:
-
base pair(s)
- BI:
-
Bayesian Inference
- cyt b:
-
cytochrome b
- COX1:
-
cytochrome c oxidase subunit 1
- COX2:
-
cytochrome c oxidase subunit 2
- COX3:
-
cytochrome c oxidase subunit 3
- H-strand:
-
heavy strand
- ML:
-
Maximum Likelihood
- MP:
-
Maximum Parsimony
- ND1:
-
NADH Dehydrogenase Subunit 1
- ND2:
-
NADH Dehydrogenase Subunit 2
- ND3:
-
NADH Dehydrogenase Subunit 3
- ND4:
-
NADH Dehydrogenase Subunit 4
- ND4L:
-
NADH dehydrogenase subunit 4L
- ND5:
-
NADH Dehydrogenase Subunit 5
- ND6:
-
NADH Dehydrogenase Subunit 6
- NUC:
-
13 PCGs and 2 rRNA genes (12 s RNA and 16 s RNA)
- PCG:
-
protein-coding gene
- rRNA:
-
ribosomal RNA
- RAG1:
-
exon 3 of recombination activating gene 1
- tRNA:
-
transfer RNA
References
Bankevich A, Nurk S et al (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477. https://doi.org/10.1089/cmb.2012.0021
Chen A (2014) Studies on molecular phylogeny of the Gobioninae (Teleostei:Cyprinidae). Dissertation, Fudan University
Dai M, Thompson RC, Maher C et al (2010) NGSQC: cross-platform quality analysis pipeline for deep sequencing data. BMC Genom 11(Suppl 4):S7. https://doi.org/10.1186/1471-2164-11-S4-S7
Ding R (1994) The Fishes of Sichuan. Science and Technology Press, Chengdu
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98. https://doi.org/10.1021/bk-1999-0734.ch008
He A, Luo Y, Yang H et al (2011) Complete mitochondrial DNA sequences of the Nile tilapia (Oreochromis niloticus) and Blue tilapia (Oreochromis aureus): genome characterization and phylogeny applications. Mol Biol Rep 38:2015–2021. https://doi.org/10.1007/s11033-010-0324-7
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kumar S, Stecher G, Li M et al (2018) MEGA X: molecular evolutionary genetics analysis across co -mputing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Lanfear R, Calcott B, Ho SYW et al (2012) PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol 29:1695–1701. https://doi.org/10.1093/molbev/mss020
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964. https://doi.org/10.1093/nar/25.5.955
Nylander J (2004) MrModeltest v2. Program distributed by the author. Evolutionary Biology Center, Uppsala University, Uppsala
Ojala D, Montoya J, Attardi G (1981) TRNA punctuation model of RNA processing in human mitochondria. Nature 290:470–474. https://doi.org/10.1038/290470a0
Ronquist F, Teslenko M, van der Mark P et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Rota-Stabelli O, Kayal E, Gleeson D et al (2010) Ecdysozoan mitogenomics: evidence for a common origin of the legged invertebrates, the Panarthropoda. Genome Biol Evol 2:425–440. https://doi.org/10.1093/gbe/evq030
Rubinoff D, Holland BS, Savolainen V (2005) Between two extremes: mitochondrial DNA is neither the panacea nor the nemesis of phylogenetic and taxonomic inference. Syst Biol 54:952–961. https://doi.org/10.1080/10635150500234674
Tang KL, Agnew MK, Chen WJ (2011) Phylogeny of the gudgeons (Teleostei:Cyprinidae: Gobioninae). Mol Phylogenet Evol 61:103–124. https://doi.org/10.1016/j.ympev.2011.05.022
Tao W, Zhao H (2016) The complete mitogenome of Gnathopogon polytaenia (Cypriniformes; Cyprinidae). Mitochondrial DNA 27:1307–1308. https://doi.org/10.3109/19401736.2014.945569
Trifinopoulos J, Nguyen LT, von Haeseler A et al (2016) W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res 44:W232–W235. https://doi.org/10.1093/nar/gkw256
Wang J, Tang Q, Wang Z et al (2012) The complete mitogenome sequence of a cave loach Triplophysa rosa (Teleostei, Balitoridae, Nemacheilinae). Mitochondrial DNA 23: 366–368. https://doi.org/10.3109/19401736.2012.696628
Xiang H (1999) The fishery resources of Xiangxi River and the influence of the Three Gorges Project. Bull Biol 05:44
Yang J, He S, Freyhof J et al (2006) The phylogenetic relationships of the Gobioninae (Teleostei: Cyprinidae) inferred from mitochondrial cytochrome b gene sequences. Hydrobiologia 553:255–266. https://doi.org/10.1007/s10750-005-1301-3
Zeng Y, Chen Y, Li Z (2014) Utilization and protection status of fish resources in Jialing River. Tianjin Agric Sci 20:60–62. https://doi.org/10.3969/j.issn.1006-6500.2014.02.014
Zhang F, Mi Z (1998) Advance in molecular biology of animal mitochondrial DNA. Prog Biotechnol 18:25–32. https://doi.org/10.13523/j.cb.19980305
Zhang F, Shen Y (2018) Characterization of the complete mitochondrial genome of Rhinogobius leavelli (Perciformes: Gobiidae: Gobionellinae) and its phylogenetic analysis for Gobionellinae. Biologia 74:493–499. https://doi.org/10.2478/s11756-018-00189-5
Zhang D, Zou H, Hua CJ et al (2019) Mitochondrial architecture rearrangements produce asymmetrical nonadaptive mutational pressures that subvert the phylogenetic reconstruction in Isopoda. Genome Biol Evol 11:1797–1812. https://doi.org/10.1101/607960
Zhang D, Gao F, Jakovlić I et al (2020) PhyloSuite: An integrated and scalable desktop platform for streamlined molecular sequence data management and evolutionary phylogenetics studies. Mol Ecol Resour 20:348–355. https://doi.org/10.1111/1755-0998.13096
Zhao S, Ye S, Xie S et al (2015) The current situation of fishery resources in the Xiangxi River of the Three Gorges Reservoir and advices on the management. Acta Hydrobiol Sin 39:973–982. https://doi.org/10.7541/2015.127
Acknowledgements
We thank Zhenyu Lv for his help in sample collection and identification.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
This research was supported by the National Natural Science Foundation of China (No. 51,779,210), and the Fundamental Research Funds of China West Normal University (No. 19C007, No. CXTD2016-3). The authors report no conflicts of interest. The authors alone are responsible for the content and writing of the article.
Rights and permissions
About this article
Cite this article
Mao, L., Zeng, Y., Li, J. et al. The complete mitochondrial genome sequence and phylogenetic analysis of Gnathopogon herzensteini(Cypriniformes, Cyprinidae, Gobioninae). Biologia 76, 1087–1094 (2021). https://doi.org/10.2478/s11756-020-00667-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.2478/s11756-020-00667-9