Abstract
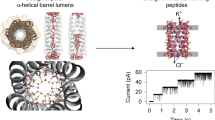
The zervamicins (Zrv) are a family of 16 residue peptaibol channel formers, related to the 20 residue peptaibol alamethicin (Alm), but containing a higher proportion of polar sidechains. Zrv-1113 forms multi-level channels in planar lipid (diphytanoyl phosphatidylcholine) bilayers in response to cis positive voltages. Analysis of the voltage and concentration dependence of macroscopic conductances induced by Zrv-IIB suggests that, on average, channels contain ca. 13 peptide monomers. Analysis of single channel conductance levels suggests a similar value. The pattern of successive conductance levels is consistent with a modified helix bundle model in which the higher order bundle are distorted within the plane of the bilayer towards a “torpedo” shaped cross-section. The kinetics of intro-burst switching between adjacent conductance levels are shown to be approximately an order of magnitude faster for Zrv-IIB than for Alm. The channel forming properties of the related naturally occurring peptaibols, Zrv-Leu and Zrv-IC, have also been demonstrated, as have those of the synthetic apolar analogue Zrv-Al-16. The experimental studies on channel formation are combined with the known crystallographic structures of Zrv-Al-16 and Zrv-Leu to develop a molecular model of Zrv-II3 channels.
Similar content being viewed by others
Abbreviations
- Alm:
-
Alamethicin
- Zrv:
-
Zervamicin
- CFP:
-
Channel forming peptide
- Aib:
-
α-aminoisobutyric acid
References
Agarwalla S, Mellor IR, Sansom MSP, Karle IL, Flippen-Anderson JL, Uma K, Krishna K, Sukumar M, Balaram P (1992) Zervamicins, a structurally characterised peptide model for membrane ion channels. Biochem Biophys Res Comm (submitted for publication)
Ball FG, Sansom MSP (1989) Single channel gating mechanisms: model identification and parameter estimation. Proc R Soc Lond B 236:385–416
Betz H (1990) Homology and analogy in transmembrane channel design: lessons from synaptic membrane proteins. Biochemistry 29:3591–3599
Boheim G (1974) Statistical analysis of alamethicin channels in black lipid membranes. J Membr Biol 19:277–303
Boheim G, Kolb HA (1978) Analysis of the multi-pore system of alamethicin in a lipid membrane: I — Voltage-jump current relaxation measurements. J Membr Biol 38:99–150
Boheim G, Hanke W, Jung G (1983) Alamethicin pore formation: voltage-dependent flip-flop of α-helix dipoles. Biophys Struct Mech 9:181–191
Boheim G, Gellert S, Jung G, Menestrina G (1987) α-Helical ion channels reconstituted into planar bilayers. In: Yagi K, Pullman B (eds) Ion transport through membranes. Academic Press, Tokyo, pp 131–145
Brandl CJ, Deber CM (1986) Hypothesis about the function of membrane-buried proline residues in transport proteins. Proc Natl Acad Sci USA 83:917–921
Coronado R, Latorre R (1983) Phospholipid bilayers made from monolayers on patch-clamp pipettes. Biophys J 43:231–236
Fox OR, Richards FM (1982) A voltage-gated ion channel model inferred from the crystal structure of alamethicin at 1.5 Å resolution. Nature 300:325–330
Fraternali F (1990) Restrained and unrestrained molecular dynamics simulations in the NVT ensemble of alamethicin. Biopolymers 30:1083–1099
Galzi JL, Revah F, Bessis A, Changeux JP (1991) Functional architecture of the nicotinic acethylholine receptor: from electric organ to brain. Ann Rev Pharmacol Toxicol 31:37–72
Hall JE, Vodyanoy I, Balasubramanian TM, Marshall GR (1984) Alamethicin: a rich model for channel behaviour. Biophys J 45:233–247
Hille B (1984) Ionic channels of excitable membranes, Sinauer, Sunderland, Mass
Hol WG, von Duijen PT, Berendsen HJC (1978) The α-helix dipole and the properties of proteins. Nature 273:443–446
Huang HW, Wu Y (1991) Lipid-alamethicin interactions influence alamethicin orientation. Biophys J 60:1079–1087
Karle IL, Flippen-Andersen J, Sukumar M, Balaram P (1987) Conformation of a 16-residue zervamicin IIA analog peptide containing 3 different structural features: 310-helix, α-helix and β-bend ribbon. Proc Natl Acad Sci USA 84:5087–5091
Karle IL, Flippen-Andersen J, Agarwalla S, Balaram P (1991) Crystal structure of Leu-zervamicin, a membrane ion channel peptide. Implications for gating mechanisms. Proc Natl Acad Sci USA 88:5307–5311
Krishna K, Sukumar M, Balaram P (1990) Structural chemistry and membrane modifying activity of the fungal polypeptides zervamicins, antiamoebins and efrapeptins. Pure Appl Chem 62:1417–1420
Lear JD, Wasserman ZR, DeGrado WF (1988) Synthetic amphiphilic peptide models for protein ion channels. Science 240:1177–1181
Mathew MK, Balaram P (1983) A helix dipole model for alamethicin and related transmembrane channels. FEBS Lett 157:1–5
Mellor IR, Thomas DH, Sansom MSP (1988) Properties of ion channels formed by Staphylococcus aureus δ-toxin. Biochim Biophys Acta 942:280–294
Mellor IR, Sansom MSP (1990) Ion channel properties of mastoparan, a 14 residue peptide from wasp venom, and of MP3, a 12 residue analogue. Proc R Soc Lond B 239:383–400
Menestrina G, Voges KP, Jung G, Boheim G (1986) Voltage-dependent channel formation by rods of helical peptides. J Membr Biol 93:111–132
Mitra AK, McCarthy MP, Stroud RM (1989) Three-dimensional structure of the nicotinic acetylcholine receptor and location of the major associated 43-kD cytoskeletal protein, determined at 22 Å by low dose electron microscopy and X-ray diffraction to 12.5 Å. J Cell Biol 109:755–774
Molle G, Dugast JY, Duclohier H, Spach G (1988) Conductance properties of des-Aib-Leu-des-Pheol-Phe-alamethicin in planar lipid bilayers. Biochim Biophys Acta 938:310–314
Molle G, Duclohier H, Julien S, Spach G (1991) Synthetic analogues of alamethicin: effect of C-terminal residue substitutions and chain length on the ion channel lifetime. Biochim Biophys Acta 1064:365–369
Mouritsen OG, Bloom M (1984) Mattress model of lipid-protein interactions in membranes. Biophys J 46:141–153
Oiki S, Madison V, Montal M (1990) Bundles of amphipathic transmembrane α-helices as a structural motif for ion-conducting channel proteins: studies on sodium channels and acetylcholine receptors. Proteins: Structure Funct Genet 8:226–236
Rinehart KL, Gaudioso LA, Moore ML, Pandey RC, Cok JC, Brber M, Sedgwick D, Bordoli RS, Tyler AN, Green BN (1981) Structures of eleven zervamicin and two emerimicin peptide antibiotics studied by fast atom bombardment mass spectroscopy. J Am Chem Soc 103:6517–6520
Rizzo V, Stankowski S, Schwarz G (1987) Alamethicin incorporation in lipid bilayers: a thermodynamic analysis. Biochemistry 26:2751–2759
Robinson AA, Stokes RH (1965) Electrolyte solutions. Butterworth, London
Sansom MSP, Usherwood PNR (1990) Single channel studies of glutamate receptors. Int Rev Neurobiol 32:51–106
Sansom MSP (1991) The biophysics of peptide models of ion channels. Prog Biophys Mol Biol 55:139–236
Sansom MSP (1992a) Proline residues in transmembrane helices of channel and transport proteins: a molecular modelling study. Prot Engineering 5:53–60
Sansom MSP (1992b) Sidechain-ion interactions in ion channels: a molecular modelling study. Biochem Soc Transac 20:2545
Sansom MSP, Kerr ID, Mellor IR (1991) Ion channels formed by amphipathic helical peptides — a molecular modelling study. Eur Biophys J 20:229–240
Sansom MSP, Balaram P, Karle IL et al. (1992) Molecular modelling of ion channels formed by zervamicin-IIB (in preparation)
Schwarz G, Savko P (1982) Structural and dipolar properties of the voltage-dependent pore former alamethicin in octanol/dioxane. Biophys J 39:211–219
Stankowski S, Schwarz UD, Schwarz G (1988) Voltage-dependent pore activity of the peptide alamethicin correlated with incorporation in the membrane: salt and cholesterol effects. Biochim Biophys Acta 941:11–18
Stroud RM, McCarthy MP, Shuster M (1990) Nicotinic acetylcholine receptor superfamiliy of ligand-gated ion channels. Biochemistry 29:11009–11023
Sukumar M (1987) PhD Thesis, Indian Institute of Science, Bangalore
Taylor RJ, de Levie R (1991) “Reversed” alamethicin conductance in bilayers. Biophys J 59:873–879
Toyoshima C, Unwin N (1988) Ion channel of acetylcholine receptor reconstructed from images of post-synaptic membranes. Nature 336:247–250
Unwin N (1989) The structure of ion channels in membranes of excitable cells. Neuron 3:665–676
Vodyanoy I, Hall JE, Balasubramanian TM (1983) Alamethicin-induced current-voltage curve asymmetry in lipid bilayers. Biophys J 42:71–82
Vodyanoy I, Hall JE, Vodyanoy V (1988) Alamethicin adsorption to a planar lipid bilayer. Biophys J 53:649–658
von Heijnc G (1991) Proline kinks in transmembrane α-helices. J Mot Biol 218:499–503
Williams KA, Deber CM (1991) Proline residues in transmembrane helices: structural or dynamic role? Biochemistry 30:8919–8923
Woolfson DN, Mortishire-Smith RJ, Williams DH (1991) Conserved positioning of proline residues in membrane-spanning helices of ion-channel proteins. Biochem Biophys Res Comm 175:733–737
Woolfson DN, Williams DH (1990) The influence of proline residues on α-helical structure. FEBS Lett 277:185–188
Author information
Authors and Affiliations
Additional information
Correspondence to: M. S. P. Sansom
Rights and permissions
About this article
Cite this article
Balaram, P., Krishna, K., Sukumar, M. et al. The properties of ion channels formed by zervamicins. Eur Biophys J 21, 117–128 (1992). https://doi.org/10.1007/BF00185426
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00185426