Summary
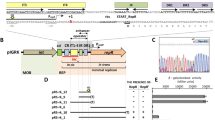
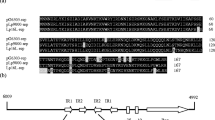
Comparative analyses were made between plasmid pSa17, a deletion derivative of pSa that is capable of replicating efficiently in Escherichia coli and plasmid pSa3, a derivative that is defective for replication. By comparing the restriction maps of these two derivatives, the regions essential for replication and for stable maintenance of the plasmid were determined. A 2.5 kb DNA segment bearing the origin of DNA replication of pSa17 was sequenced. A 36 kDa RepA protein was encoded in the region essential for replication. Downstream of the RepA coding region was a characteristic sequence including six 17 bp direct repeats, the possible binding sites of RepA protein, followed by AT-rich and GC-rich sequences. Furthermore, an 8 bp incomplete copy of the 17 bp repeat was found in the promoter region of the repA gene. Based on the hypothesis that RepA protein binds to this partial sequence as well as to intact 17 bp sequences, an autoregulatory system for the synthesis of RepA protein may be operative. Another open reading frame (ORF) was found in the region required for the stability of the plasmid. The putative protein encoded in this ORF showed significant homology to several site-specific recombination proteins. A possible role of this putative protein in stable maintenance of the plasmid is discussed.
Similar content being viewed by others
References
Birnboim HC, Doly J (1979) A rapid alkaline extraction procedure for screening recombinant plasmids. Nucleic Acids Res 7:1513–1523
Boyer HW, Roulland-Dussoix D (1969) A complementation analysis of the restriction and modification of DNA in Escherichia coli. J Mol Biol 41:459–472
Churchward G, Linder P, Caro L (1983) The nucleotide sequence of replication and maintenance function encoded by plasmid pSC101. Nucleic Acids Res 11:5645–5659
Close SM, Kado CI (1992) A gene near the plasmid pSa origin of replication encodes a nuclease. Mol Microbiol 6:521–527
Close SM, Kado CI (1991) The osa gene of pSa encodes a 21.1-kilodalton protein that suppresses Agrobacterium tumefaciens oncogenicity. J Bacteriol 173:5449–5456
Cooley MB, D'Souza MR, Kado CI (1991) The virC and virD operons of the Agrobacterium Ti plasmid are regulated by the ros chromosomal gene: analysis of the cloned ros gene. J Bacteriol 173:2608–2616
Datta N (1975) Epidemiology and classification of plasmids. In: Schlessinger D (ed) Microbiology — 1974. American Society for Microbiology, Washington, DC, pp 9–15
Diver WP, Grinsted J, Fritzinger DC, Brown NL, Altenbuchner J, Rogowsky P, Schmitt R (1983) DNA sequences of and complementation by the tnpR genes of Tn21, Tn501 and Tn1721. Mol Gen Genet 191:189–193
Dodd HM, Bennett PM (1987) The R46 site-specific recombination system is a homologue of the Tn3 and γδ (Tn1000) cointegrate resolution system. J Gen Microbiol 133:2031–2039
Gerlitz M, Hrabak O, Schwab H (1990) Partitioning of broad-host-range plasmid RP4 is a complex system involving site-specific recombination. J Bacteriol 172:6194–6203
Germino J, Bastia D (1982) Primary structure of the replication initiation protein of plasmid RK6. Proc Natl Acad Sci USA 79:5475–5479
Gorai AP, Heffron F, Falkow S, Hedges RW, Datta N (1979) Electron microscope heteroduplex studies of sequence relationships among plasmids of the W incompatibility group. Plasmid 2:489–492
Grosschedl R, Hobom G (1979) DNA sequences and structural homologies of the replication origins of lambdoid bacteriophages. Nature 277:621–627
Hakkaart MJJ, Uan den Elzen PJM (1984) Maintenance of multicopy plasmid CloDF13 in E. coli cells: Evidence for site-specific recombination at parB. Cell 36:203–209
Haring V, Scholz P, Scherzinger E, Frey J, Derbyshire K, Hatfull G, Willetts NS, Bagdasarian M (1985) Protein RepC is involved in copy number control of the broad-host-range plasmid RSF1010. Proc Natl Acad Sci USA 82:6090–6094
Henikoff S (1984) Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene 28:351–359
Hyde DR, Tu CD (1985) tnpM: A novel regulatory gene that enhances Tn21 transposition and suppresses cointegrate resolution. Cell 42:629–638
Kado CI, Heskett MG, Langley RA (1972) Studies on Agrobacterium tumefaciens. Characterization of strains 1D135 and B6, and analysis of the bacterial chromosome, transfer RNA and ribosomes for tumor-inducing ability. Physiol Plant Pathol 2:47–57
Kelley W, Bastia D (1985) Replication initiator protein of plasmid R6K autoregulates its own synthesis at the transcriptional step. Proc Natl Acad Sci USA 82:2574–2578
Klapwijk P, Van Beelen P, Schilperoort RA (1979) Isolation of a recombinant-deficient Agrobacterium tumefaciens mutant. Mol Gen Genet 173:171–175
Krishman BR, Iyer VN (1990) IncN plasmid replicon: A deletion and subcloning analysis. J Mol Biol 213:777–788
Maniatis T, Fritsch EF, Sambrook JH (1982) Molecular cloning: A laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Murotsu T, Matsubara K, Sugisaki H, Takanami M (1981) Nine unique repeating sequences in a region essential for replication and incompatibility of the mini-F plasmid. Gene 15:257–271
Newman BJ, Grindley NDF (1984) Mutants of the γδ resolvase: a genetic analysis of the recombination function. Cell 38:463–469
Pinkney M, Diaz R, Lanka E, Thomas CM (1988) Replication of mini-RK2 plasmid in extract of Escherichia coli requires plasmid-encoded protein TrfA and host-encoded proteins DnaA, B, G, DNA gyrase and DNA polymerase III. J Mol Biol 203:927–938
Plasterk RHA, Brinkman A, Van de Putte P (1983) DNA inversions in the chromosome of Escherichia coli and in bacteriophage Mu: Relationship to other site-specific recombination systems. Proc Natl Acad Sci USA 80:5355–5358
Reed RR, Shibuya GI, Steiz JA (1982) Nucleotide sequence of γδ resolvase gene and demonstration that its gene product acts as a repressor of transcription. Nature 300:381–383
Roberts RC, Burioni R, Helinski DR (1990) Genetic characterization of the stabilizing function of a region of broad-host-range plasmid RK2 J Bact 172:6204–6216
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467
Scherer G (1978) Nucleotide sequence of the 0 gene and of the origin of replication in bacteriophage λ DNA. Nucleic Acids Res 5:3141–3156
Schmidhauser TJ, Ditta G, Helinski DR (1988) Broad-host-range plasmid cloning vectors for gram-negative bacteria. In: Rodriguez RL, Denhardt DT (eds) Vectors. A survey of molecular cloning vectors and their uses. Butterworth, Stoneham, nlass., pp 287–382
Shingler V, Thomas CM (1984) Analysis of the trfA region of broad-host-range plasmid RK2 by transposon mutagenesis and identification of polypeptide products. J Mol Biol 175-229–250
Smith CA, Thomas CM (1984) Nucleotide sequences of the TrfA gene of broad-host range plasmid RK2. J Mol Biol 175:251–262
Sobell DW, Lozanski ED, Lessen M (1978) Structural and energetic considerations of wave propagation in DNA. Cold Spring Harbor Symp Quant Biol 43:11–20
Soberon X, Covarrubias L, Bolivar F (1980) Construction and characterization of new cloning vehicles. IV. Deletion derivatives of pBR322 and pBR325. Gene 9:287–305
Stalker DM, Kolter R, Helinski DR (1979) Nucleotide sequence of the region of replication of the antibiotic resistance plasmid R6K. Proc Natl Acad Sci USA 767:11150–1154
Stalker DM, Thomas CM, Helinski DR (1981) Nucleotide sequence of the region of the origin of replication of the broad host range plasmid RK2. Mol Gen Genet 181:8–12
Stalker DM, Koller R, Helinski DR (1982) Plasmid R6K DNA replication. I. Complete nucleotide sequence analysis of an autonomously replicating segment. J Mol Biol 161:33–43
Tait RC, Lundquist RC, Kado CI (1982a) Genetic map of the crown gall suppressive IncW plasmid pSa. Mol Gen Genet 186:10–15
Tait RC, Close TJ, Rodriguez RL, Kado CI (1982b) Isolation of the region of replication of the IncW-group plasmid pSa. Gene 20:39–49
Tait RC, Close TJ, Lundquist RC, Hagiya M, Rodriguez RL, Kado CI (1983) Construction and characterization of a versatile broad host range DNA cloning system for Gram-negative bacteria. Biotechnology 1:269–275
Thomas CM, Stalker DM, Helinski DR (1981) Replication and incompatibility properties of the origin region of replication of broad-host-range plasmid RK2. Mol Gen Genet 181:1–7
Veltkamp E, Stuitje AR (1981) Replication and structure of the bacteriocinogenic plasmids CloDF-13 and Col E1. Plasmid 5:76–99
Ward JM, Grinsted J (1982) Physical and genetic analysis of the IncW group plasmids 8388, Sa, and R7K. Plasmid 7:239–250
Yanisch-Perron C, Vieira J Messing J (1983) Improved M13 phage cloning vector and host strains: nucleotide sequence of the M13mp18 and pUC19 vectors. Gene 33:103–119
Author information
Authors and Affiliations
Additional information
Communicated by W. Goebel
Rights and permissions
About this article
Cite this article
Okumura, M.S., Kado, C.I. The region essential for efficient autonomous replication of pSa in Escherichia coli . Molec. Gen. Genet. 235, 55–63 (1992). https://doi.org/10.1007/BF00286181
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00286181