Abstract
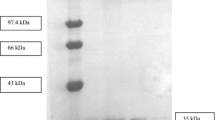
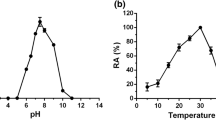
Bacterial esterases are highly versatile enzymes, currently widely used in detergents, biosurfactants, bioemulsifiers and as biocatalysts in paper and food industries. Present work describes heterologous expression, purification, and biophysical and biochemical characterization of a halotolerant esterase from Bacillus licheniformis (BlEstA). BlEstA preferentially cleaves pNP-octanoate and both activity and stability of the enzyme increased in the presence of 2 M NaCl, and also with several organic solvents (ethanol, methanol and DMSO). Furthermore, BlEstA has considerable emulsifying properties, particularly with olive oil as substrate. Our studies also show that the enzyme is monomeric in solution and its small-angle X-ray scattering low-resolution molecular envelope fits well its high-resolution homology model.









Similar content being viewed by others
References
Abramić M, Leščić I, Korica T, Vitale L, Saenger W, Pigac J (1999) Purification and properties of extracellular lipase from Streptomyces rimosus. Enzyme Microb Technol 25:522–529. https://doi.org/10.1016/S0141-0229(99)00077-0
Adrio J, Demain A (2013) Microbial enzymes: tools for biotechnological processes. Biomolecules 4:22. https://doi.org/10.3390/biom4010117
Alex D, Shainu A, Pandey A, Sukumaran RK (2014) Esterase active in polar organic solvents from the yeast Pseudozyma sp. NII 08165. Enzyme Res. https://doi.org/10.1155/2014/494682
Balaji L, Jayaraman G (2014) Metal ion activated lipase from halotolerant Bacillus Sp. VITL8 displays broader operational range. Int J Biol Macromol 67:380–386. https://doi.org/10.1016/j.ijbiomac.2014.03.050
Balan DAI (2012) Purification and Characterization of a Thermostable Lipase from Geobacillus thermodenitrificans IBRL-nra. Enzyme Res. https://doi.org/10.1155/2012/987523
Banat IM, Makkar RS, Cameotra SS (2000) Potential commercial applications of microbial surfactants. Appl Microbiol Biotechnol 53:495–508. https://doi.org/10.1007/s002530051648
Barros M, Fleuri LF, Macedo GA (2010) Seed lipases: sources, applications and properties - a review. Brazilian J Chem Eng 27:15–29. https://doi.org/10.1590/s0104-66322010000100002
Bora L, Bora M (2012) Optimization of extracellular thermophilic highly alkaline lipase from thermophilic Bacillus Sp isolated from hotspring of Arunachal pradesh, India. Brazilian J Microbiol 43:30–42. https://doi.org/10.1590/s1517-83822012000100004
Boratyn GM, Schäffer AA, Agarwala R, Altschul SF, Lipman DJ (2012) Domain enhanced lookup time accelerated BLAST. Biol Direct 7:1–14. https://doi.org/10.1186/1745-6150-7-12
Camilo CM, Polikarpov I (2014) High-throughput cloning, expression and purification of glycoside hydrolases using ligation-independent cloning (LIC). Protein Expr Purif 99:35–42. https://doi.org/10.1016/j.pep.2014.03.008
Cooper DG (1986) Surface-active agents from two Bacilllus Species. Appl Environ Microbiol 53:224–229.
de Almeida JM, Moure VR, Muller-Santos M, de Souza EM, Pedrosa FO, Mitchell DA, Krieger N (2018) Tailoring recombinant lipases: keeping the his-tag favors esterification reactions, removing it favors hydrolysis reactions. Sci Rep 8:1–11. https://doi.org/10.1038/s41598-018-27579-8
Doukyu N, Ogino H (2010) Organic solvent-tolerant enzymes. Biochem Eng J 48:270–282. https://doi.org/10.1016/j.bej.2009.09.009
Dutta S, Ray L (2009) Production and characterization of an alkaline thermostable crude lipase from an isolated strain of Bacillus cereus C7. Appl Biochem Biotechnol 159:142–154. https://doi.org/10.1007/s12010-009-8543-x
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. BMC Bioinform 19:1792–1797. https://doi.org/10.1093/nar/gkh340
Esteban-Torres M, Mancheno JM, de las Rivas B, Munoz R, (2015) Characterization of a halotolerant lipase from the lactic acid bacteria Lactobacillus plantarum useful in food fermentations LWT. Food Sci Technol 60:246–252. https://doi.org/10.1016/j.lwt.2014.05.063
Eveleigh DE (1981) The microbiological production of industrial-chemicals. Sci Am 245:155–156. https://doi.org/10.2307/24964563
Fariha H, Aamer AS, Hammed A (2006) Industrial application of microbial lipases. Enzyme Microb Technol 39:235–251. https://doi.org/10.1016/j.enzmictec.2005.10.016
Fariha H, Aamer S, Hameed A (2007) Purification and characterization of a mesophilic lipase from Bacillus subtilis FH5 stable at high temperature And pH. Acta Biol Hung. https://doi.org/10.1556/ABiol.58.2007.1.11
Ghati A, Paul G (2015) Purification and characterization of a thermo-halophilic, alkali-stable and extremely benzene tolerant esterase from a thermo-halo tolerant Bacillus cereus strain AGP-03, isolated from ‘Bakreshwar’ Hot Spring India. Process Biochem 50(5):771–781. https://doi.org/10.1016/j.procbio.2015.01.026
Glatter O, Kratky O (1982) Small angle X‐Ray scattering, Acad. Press Inc. Ltd. 10.1002/actp.1985.010360520
Gokbulut AA, Arslanoglu A (2013) Purification and biochemical characterization of an extracellular lipase from psychrotolerant Pseudomonas fluorescens KE38. Turkish J Biol 37:538–546. https://doi.org/10.3906/biy-1211-10
Guinier G (1955) A small-angle scattering of X-Rays., John Wiley Sons, Inc 10.1002/pol.1956.120199326
Hammersley A (2016) FIT2D: a multi-purpose data reduction, analysis and visualization program. J Appl Crystallogr 49:646–652. https://doi.org/10.1107/S1600576716000455
Joseph B, Ramteke PW, Thomas G (2008) Cold active microbial lipases: Some hot issues and recent developments. Biotechnol Adv 26:457–470. https://doi.org/10.1016/j.biotechadv.2008.05.003
Joshi SJ, Al-Wahaibi YM, Al-Bahry SN, Elshafie AE, Al-Bemani AS, Al-Bahri A, Al-Mandhari MS (2016) Production, characterization, and application of Bacillus licheniformis W16 biosurfactant in enhancing oil recovery. Front Microbiol 7:1–14. https://doi.org/10.3389/fmicb.2016.01853
Kanmani P, Kumaresan K, Aravind J (2015) Gene cloning, expression, and characterization of the Bacillus amyloliquefaciens PS35 lipase. Brazilian J Microbiol 46:1235–1243. https://doi.org/10.1590/s1517-838246420141068
Karpushova A, Brümmer F, Barth S, Lange S, Schmid RD (2005) Cloning, recombinant expression and biochemical characterisation of novel esterases from Bacillus Sp associated with the marine sponge Aplysina aerophoba. Appl Microbiol Biotechnol 67:59–69. https://doi.org/10.1007/s00253-004-1780-6
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJE (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10:845–858. https://doi.org/10.1038/nprot.2015.053
KhyamiHorani H (1996) Thermotolerant strain of Bacillus licheniformis producing lipase. World J Microbiol Biotechnol 12:399–401. https://doi.org/10.1007/bf00340219
Kim KK, Song HK, Shin DH, Hwang KY, Suh SW (1997) The crystal structure of a triacylglycerol lipase from Pseudomonas cepacia reveals a highly open conformation in the absence of a bound inhibitor. Structure 5(2):173–185. https://doi.org/10.1016/S0969-2126(97)00177-9
Kokal S (2005) Crude-oil emulsions: a state-of-the-art review. Spe Prod Facil 20:5–13. https://doi.org/10.2118/77497-pa
Konarev PV, Svergun DI (2015) A Posteriori determination of the useful data range for small-angle Scattering Experiments On Dilute Monodisperse Systems. IUCR J 2:352–360. https://doi.org/10.1107/s2052252515005163
Kozin MB, Svergun DI (2001) Automated matching of high- and low-resolution structural models. J Appl Crystallogr 34:33–41. https://doi.org/10.1107/s0021889800014126
Kumagai PS, Gutierrez RF, Lopes JLS, Martins JM, Jameson D, Castro AM, Martins LF, DeMarco R, Bossolan NRS, Wallace BA, Araujo APU (2018) Characterization of esterase activity from an Acetomicrobium hydrogeniformans enzyme with high structural stability in extreme conditions. Extremophiles 22:781–793. https://doi.org/10.1007/s00792-018-1038-3
Kumar STK, Stecher G (2016) MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 10.1093/molbev/msw054
Kumar S, Kikon K, Upadhyay A, Kanwar SS, Gupta R (2005) Production, purification, and characterization of lipase from thermophilic and alkaliphilic Bacillus coagulans BTS-3. Protein Expres Purific 41:38–44. https://doi.org/10.1016/j.pep.2004.12.010
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 15:680–685. https://doi.org/10.1038/227680a0
Lin Y, Cheng C, Shih C, Hwang J, Yu C, Lu C (2016) MIB: metal ion-binding site prediction and docking server. J Chem Inf Model 56(12):2287–2291. https://doi.org/10.1021/acs.jcim.6b00407
Lopes E (2014) Mendes, emulsification properties of bioemulsifiers produced by wild-type and mutant Bradyrhizobium elkanii strains. J Bioremed Biodeg 5:1–6. https://doi.org/10.4172/2155-6199.1000245
Madern D, Ebel C, Zaccai G (2000) Halophilic adaptation of enzymes. Extremophiles 4:91–98. https://doi.org/10.1007/s007920050142
Mala JGS, Takeuchi S (2008) Understanding structural features of microbial lipases—an overview. Anal Chem Insights 3:9–19. https://doi.org/10.4137/aci.s551
Nakamura AM, Seiki Kadowaki MA, Godoy A, Nascimento AS, Polikarpov I (2018) Low-resolution envelope, biophysical analysis and biochemical characterization of a short-chain specific and halotolerant carboxylesterase from Bacillus licheniformis. Int J Biol Macromol 120:1893–1905. https://doi.org/10.1016/j.ijbiomac.2018.10.003
Nardini M (2000) Crystal structure of Pseudomonas aeruginosa lipase in the open conformation. J Biol Chem 275:31219–31225. https://doi.org/10.1074/jbc.M003903200
Nthangeni MB, Patterton H-G, Tonder A, van, Vergeer W.P., Litthauer D., (2001) Over-expression and properties of a purified recombinant Bacillus licheniformis lipase: a comparative report on Bacillus Lipases. Enzyme Microb Technol 28:705–712. https://doi.org/10.1016/S0141-0229(01)00316-7
Pandey A, Benjamin S, Soccol CR, Nigam P, Krieger N, Soccol VT (1999) The realm of microbial lipases in biotechnology. Biotechnol Appl Biochem 29:119–131. https://doi.org/10.1111/j.1470-8744.1999.tb00541.x
Piiadov V, de Araujo EA, Oliveira Neto M, Craievich AF, Polikarpov I (2019) SAXSMoW 2.0 online calculator of the molecular weight of proteins in dilute solution from experimental SAXS data measured on a relative scale. Protein Sci 28:454–463. https://doi.org/10.1002/pro.3528
Pontius JWS, Richelle J (1996) Deviations from standard atomic volumes as a quality measure for protein crystal structures. J Mol Biol 264:121–136. https://doi.org/10.1006/jmbi.1996.0628
Receveur-Bréchot V, Durand D (2012) How random are intrinsically disordered proteins? a small angle scattering perspective. Curr Protein Pept Sci 13:55–75. https://doi.org/10.2174/138920312799277901
Rodríguez BH, Córdova J, Barzana E, Torres EF (2009) Effects organic solvents on activity and stability of lipases produced by thermotolerant fungi in solid-state fermentation. J Mol Catal B: Enzymatic 61:136–142. https://doi.org/10.1016/j.molcatb.2009.06.004
Sangeetha R, Geetha A, Arulpandi I (2010) Concomitant production of protease and lipase by Bacillus licheniformis Vsg1: production, purification and characterization. Brazilian J Microbiol 41:179–185. https://doi.org/10.1590/s1517-83822010000100026
Schallmey M, Singh A, Ward OP (2004) Developments in the use of Bacillus species for industrial production. Can J Microbiol 50:1–17. https://doi.org/10.1139/w03-076
Schneidman-Duhovny D, Hammel M, Tainer JA, Sali A (2016) FoXS, FoXSDock and MultiFoXS: single-state and multi-state structural modeling of proteins and their complexes based on SAXS profiles. Nucleic Acids Res. 44:424–429. https://doi.org/10.1093/nar/gkw389
Schuck P, Perugini MA, Gonzales NR, Howlett GJ, Schubert D (2002) Size-distribution analysis of proteins by analytical ultracentrifugation: strategies and application to model systems. Biophys J 82:1096–1111. https://doi.org/10.1016/s0006-3495(02)75469-6
Secundo F, Carrea G (2002) Lipase activity and conformation in neat organic solvents. J Mol Catalysis B Enzymatic 19–20:93–102. https://doi.org/10.1016/S1381-1177(02)00155-8
Sharma R, Chisti Y, Banerjee UC (2001) Production, purification, characterization, and applications of lipases. Biotechnol Adv 19:627–662. https://doi.org/10.1016/s0734-9750(01)00086-6
Sharma D, Sharma B, Shukla AK (2011) Biotechnological approach of microbial lipase: a review. Biotechnology 10:17. https://doi.org/10.3923/biotech.2011.23.40
Siglioccolo A, Paiardini A, Piscitelli M, Pascarella S (2011) Structural adaptation of extreme halophilic proteins through decrease of conserved hydrophobic contact surface. BMC Struct Biol 11:50. https://doi.org/10.1186/1472-6807-11-50
Soliman NA, Knoll M, Abdel-Fattah YR, Schmid RD, Lange S (2007) Molecular cloning and characterization of thermostable esterase and lipase from Geobacillus thermoleovorans YN isolated from desert soil in Egypt. Process Biochem 42:1090–1100. https://doi.org/10.1016/j.procbio.2007.05.005
Svergun DI (1992) Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J Appl Crystallogr 25:495–503. https://doi.org/10.1107/s0021889892001663
Svergun DI, Petoukhov MV, Koch MHJ (2001) Determination of domain structure of proteins from x-ray solution scattering. Biophys J 80:2946–2953. https://doi.org/10.1016/s0006-3495(01)76260-1
Torres S, Martínez MA, Pandey A, Castro GR (2009) An organic-solvent-tolerant esterase from thermophilic Bacillus licheniformis S-86. Biores Technol 100:896–902. https://doi.org/10.1016/j.biortech.2008.07.009
Volkov VV, Svergun DI (2003) Uniqueness of Ab initio shape determination in small-angle scattering. J Appl Crystallogr 36:860–864. https://doi.org/10.1107/s0021889803000268
Wakelin NG, Forster CF (1997) An Investigation into microbial removal of fats, oils and greases. Bioresour Technol 59:37–43. https://doi.org/10.1016/s0960-8524(96)00134-4
Wang G, Wang Q, Lin X, Bun Ng T, Yan R, Lin J, Ye X (2016) A novel cold-adapted and highly salt-tolerant esterase from Alkalibacterium Sp. SL3 from the sediment of a soda lake. Sci Rep 6:19494. https://doi.org/10.1038/srep19494F
Waterhouse T, Bertoni A, Bienert M, Studer S, Tauriello G, Gumienny G, Heer R, de Beer FT, Rempfer TAP, Bordoli C, Lepore L, Schwede R (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. https://doi.org/10.1093/nar/gky427
Willumsen PA, Karlson U (1997) Screening of bacteria, isolated from PAH-contaminated soils, for production of biosurfactants and bioemulsifiers. Biodegradation 7:415–423. https://doi.org/10.1007/bf00056425
Yao CJ, Cao Y, Wu SS, Li S, He BF (2013) An Organic solvent and thermally stable lipase from Burkholderia ambifaria YCJ01: purification, characteristics and application for chiral resolution of mandelic acid. J Mol Catal B-Enzymatic 85–86:105–110. https://doi.org/10.1016/j.molcatb.2012.08.016
Zhang Y (2008) I-TASSER server for protein 3D structure prediction. BioMed Cent. https://doi.org/10.1186/1471-2105-9-40
Zhang W, Xua H, Wu Y, Zeng J, Guo Z, Wang L, Shen C, Qiao D, Cao Y (2018) A new cold-adapted, alkali-stable and highly salt-tolerant esterase from Bacillus licheniformis. Int J Biol Macromol 111:1183–1193. https://doi.org/10.1016/j.ijbiomac.2018.01.152
Zheng H, Cooper DR, Porebski PJ, Shabalin IG, Handing KB, Minor W (2017) CheckMyMetal: a macromolecular metal-binding validation tool, Acta crystallographica. Sect D Struct Biol 73:223–233. https://doi.org/10.1107/S2059798317001061
Acknowledgements
This research was supported by Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP) via grant 2015/13684-0 and by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) via grants 303988/2016-9 and 440977/2016-9. This research used resources of the Brazilian Synchrotron Light Laboratory (LNLS), an open national facility operated by the Brazilian Centre for Research in Energy and Materials (CNPEM) for the Brazilian Ministry for Science, Technology, Innovations, and Communications (MCTIC). The beamline staffs are acknowledged for assistance during the experiments. We also would like to thank Prof. Dr. Ana Paula Ulian de Araujo and Raissa Ferreira Gutierrez, who helped to carry out CD measurements.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
The free informed consent form for specimen donation was signed by all participants in this research.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Leite, A.E.T., Briganti, L., de Araújo, E.A. et al. Low-resolution molecular shape, biochemical characterization and emulsification properties of a halotolerant esterase from Bacillus licheniformis. Eur Biophys J 49, 435–447 (2020). https://doi.org/10.1007/s00249-020-01448-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-020-01448-7