Abstract
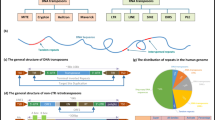
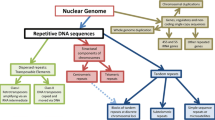
Although discovered decades ago, the molecular identification, the diversity and versatility of functions, and the evolutionary origin of repeat DNA sequences (REPs) containing palindromic units in prokaryotes are now bringing attention to a wide range of biological scientists. A brief account of the current state of the repeat DNA sequences is presented here.




Similar content being viewed by others
References
Aranda-Olmedo I, Tobes R, Manzanera M, Ramos JL, Marques S (2002) Species-specific repetitive extragenic palindromic (REP) sequences in Pseudomonas putida. Nucl Acids Res 30:1826–1833
Arthanari H, Wojtuszewski K, Mukerji I, Bolton PH (2004) Effects of HU binding on the equilibrium cyclization of mismatched, curved, and normal DNA. Biophys J 86:1625–1631
Bachellier S, Perrin D, Hofnung M, Gilson E (1993) Bacterial interspersed mosaic elements (BIMEs) are present in the genome of Klebsiella. Mol Microbiol 7:537–544
Bachellier S, Saurin W, Perrin D, Hofnung M, Gilson E (1994) Structural and functional diversity among bacterial interspersed mosaic elements (BIMEs). Mol Microbiol 12:61–70
Bachellier S, Clement JM, Hofnung M (1999) Short palindromic repetitive DNA elements in enterobacteria: a survey. Res Microbiol 150:627–639
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712
Boccard F, Prentki P (1993) Specific interaction of IHF with RIBs, a class of bacterial repetitive DNA elements located at the 3′ end of transcription units. EMBO J 12:5019–5027
Bondy-Denomy J, Garcia B, Strum S, Du M, Rollins MF, Hidalgo-Reyes Y, Wiedenheft B, Maxwell KL, Davidson AR (2015) Multiple mechanisms for CRISPR-Cas inhibition by anti-CRISPR proteins. Nature 526:136–139
Chandler M, de la Cruz F, Dyda F, Hickman AB, Moncalian G, Ton-Hoang B (2013) Breaking and joining single-stranded DNA: the HUH endonuclease superfamily. Nat Rev Microbiol 11:525–538
Dimri GP, Rudd KE, Morgan MK, Bayat H, Ames GF (1992) Physical mapping of repetitive extragenic palindromic sequences in Escherichia coli and phylogenetic distribution among Escherichia coli strains and other enteric bacteria. J Bacteriol 174:4583–4593
Espeli O, Boccard F (1997) In vivo cleavage of Escherichia coli BIME-2 repeats by DNA gyrase: genetic characterization of the target and identification of the cut site. Mol Microbiol 26:767–777
Espeli O, Moulin L, Boccard F (2001) Transcription attenuation associated with bacterial repetitive extragenic BIME elements. J Mol Biol 314:375–386
Filee J, Siguier P, Chandler M (2007) Insertion sequence diversity in archaea. Microbiol Mol Biol Rev MMBR 71:121–157
Florek MC, Gilbert DP, Plague GR (2014) Insertion sequence distribution bias in Archaea. Mob Genet Elem 4:e27829
Garneau JE, Dupuis ME, Villion M, Romero DA, Barrangou R, Boyaval P, Fremaux C, Horvath P, Magadan AH, Moineau S (2010) The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468:67–71
George B, Bhatt BS, Awasthi M, George B, Singh AK (2015) Comparative analysis of microsatellites in chloroplast genomes of lower and higher plants. Curr Genet 61:665–677
Gilson E, Clement JM, Brutlag D, Hofnung M (1984) A family of dispersed repetitive extragenic palindromic DNA sequences in E. coli. EMBO J 3:1417–1421
Gilson E, Perrin D, Clement JM, Szmelcman S, Dassa E, Hofnung M (1986) Palindromic units from E. coli as binding sites for a chromoid-associated protein. FEBS Lett 206:323–328
Gilson E, Perrin D, Hofnung M (1990) DNA polymerase I and a protein complex bind specifically to E. coli palindromic unit highly repetitive DNA: implications for bacterial chromosome organization. Nucl Acids Res 18:3941–3952
Gilson E, Saurin W, Perrin D, Bachellier S, Hofnung M (1991a) The BIME family of bacterial highly repetitive sequences. Res Microbiol 142:217–222
Gilson E, Saurin W, Perrin D, Bachellier S, Hofnung M (1991b) Palindromic units are part of a new bacterial interspersed mosaic element (BIME). Nucl Acids Res 19:1375–1383
Hammel M, Amlanjyoti D, Reyes FE, Chen JH, Parpana R, Tang HY, Larabell CA, Tainer JA, Adhya S (2016) HU multimerization shift controls nucleoid compaction. Sci Adv 2:e1600650
Hanke ML, Kielian T (2011) Toll-like receptors in health and disease in the brain: mechanisms and therapeutic potential. Clin Sci 121:367–387
Hatfield GW, Benham CJ (2002) DNA topology-mediated control of global gene expression in Escherichia coli. Annu Rev Genet 36:175–203
Hemmi H, Takeuchi O, Kawai T, Kaisho T, Sato S, Sanjo H, Matsumoto M, Hoshino K, Wagner H, Takeda K, Akira S (2000) A Toll-like receptor recognizes bacterial DNA. Nature 408:740–745
Hickman AB, Dyda F (2015) Mechanisms of DNA transposition. Microbiology spectrum 3: MDNA3-0034-2014
Higgins CF, Ames GF, Barnes WM, Clement JM, Hofnung M (1982) A novel intercistronic regulatory element of prokaryotic operons. Nature 298:760–762
Horvath P, Barrangou R (2010) CRISPR/Cas, the immune system of bacteria and archaea. Science 327:167–170
Ishihama A (2009) The nucleoid: an overview. EcoSal Plus. doi: 10.1128/ecosalplus.2.6
Jorgensen R (1990) Altered gene expression in plants due to trans interactions between homologous genes. Trends Biotechnol 8:340–344
Kar S, Edgar R, Adhya S (2005) Nucleoid remodeling by an altered HU protein: reorganization of the transcription program. Proc Natl Acad Sci USA 102:16397–16402
Lam S, Roth JR (1983) IS200: a Salmonella-specific insertion sequence. Cell 34:951–960
Lee R, Feinbaum R, Ambros V (2004) A short history of a short RNA. Cell 116:S89–S92 (1 p following S96)
Liu LF, Wang JC (1987) Supercoiling of the DNA template during transcription. Proc Natl Acad Sci USA 84:7024–7027
Macvanin M, Adhya S (2012) Architectural organization in E. coli nucleoid. Biochim Biophys Acta 1819:830–835
Macvanin M, Edgar R, Cui F, Trostel A, Zhurkin V, Adhya S (2012) Noncoding RNAs binding to the nucleoid protein HU in Escherichia coli. J Bacteriol 194:6046–6055
Magnusson M, Tobes R, Sancho J, Pareja E (2007) Cutting edge: natural DNA repetitive extragenic sequences from gram-negative pathogens strongly stimulate TLR9. J Immunol 179:31–35
Maxwell A, Gellert M (1986) Mechanistic aspects of DNA topoisomerases. Adv Protein Chem 38:69–107
Messing SA, Ton-Hoang B, Hickman AB, McCubbin AJ, Peaslee GF, Ghirlando R, Chandler M, Dyda F (2012) The processing of repetitive extragenic palindromes: the structure of a repetitive extragenic palindrome bound to its associated nuclease. Nucl Acids Res 40:9964–9979
Morrison A, Cozzarelli NR (1979) Site-specific cleavage of DNA by E. coli DNA gyrase. Cell 17:175–184
Nunvar J, Huckova T, Licha I (2010) Identification and characterization of repetitive extragenic palindromes (REP)-associated tyrosine transposases: implications for REP evolution and dynamics in bacterial genomes. BMC Genom 11:44
Oppenheim AB, Rudd KE, Mendelson I, Teff D (1993) Integration host factor binds to a unique class of complex repetitive extragenic DNA sequences in Escherichia coli. Mol Microbiol 10:113–122
Parthiban P, Mahendra J (2015) Toll-like receptors: a key marker for periodontal disease and preterm birth—a contemporary review. J Clin Diagn Res JCDR 9:ZE14–ZE17
Postow L, Hardy CD, Arsuaga J, Cozzarelli NR (2004) Topological domain structure of the Escherichia coli chromosome. Genes Dev 18:1766–1779
Qian Z, Macvanin M, Dimitriadis EK, He X, Zhurkin V, Adhya S (2015) A new noncoding RNA arranges bacterial chromosome organization. mBio 6(4):e00998–15. doi:10.1128/mBio.00998-15.
Raghavan R, Groisman EA, Ochman H (2011) Genome-wide detection of novel regulatory RNAs in E. coli. Genome Res 21:1487–1497
Rocco F, De Gregorio E, Di Nocera PP (2010) A giant family of short palindromic sequences in Stenotrophomonas maltophilia. FEMS Microbiol Lett 308:185–192
Rudd KE (1998) Linkage map of Escherichia coli K-12, edition 10: the physical map. Microbiol Mol Biol Rev MMBR 62:985–1019
Sternberg SH, Richter H, Charpentier E, Qimron U (2016) Adaptation in CRISPR-Cas systems. Mol Cell 61:797–808
Tobes R, Pareja E (2005) Repetitive extragenic palindromic sequences in the Pseudomonas syringae pv. tomato DC3000 genome: extragenic signals for genome reannotation. Res Microbiol 156:424–433
Tobes R, Ramos JL (2005) REP code: defining bacterial identity in extragenic space. Environ Microbiol 7:225–228
Ton-Hoang B, Siguier P, Quentin Y, Onillon S, Marty B, Fichant G, Chandler M (2012) Structuring the bacterial genome: Y1-transposases associated with REP-BIME sequences. Nucl Acids Res 40:3596–3609
Yang Y, Ames GF (1988) DNA gyrase binds to the family of prokaryotic repetitive extragenic palindromic sequences. Proc Natl Acad Sci USA 85:8850–8854
Yang J, Li F (2016) Are all repeats created equal? Understanding DNA repeats at an individual level. Curr Genet. doi:10.1007/s00294-016-0619-x
Acknowledgments
The work on naRNA was supported by the Intramural Research Program of the National Institutes of Health, National Cancer Institute, and the Center for Cancer Research.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Qian, Z., Adhya, S. DNA repeat sequences: diversity and versatility of functions. Curr Genet 63, 411–416 (2017). https://doi.org/10.1007/s00294-016-0654-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-016-0654-7