Abstract
Autophagy is a vital conserved recycling process where eukaryotic cells remove unwanted proteins and organelles via lysosomal degradation and in turn, generate nutrients for the cells. The special feature of autophagy process is the formation of double-membrane vesicles called autophagosomes that engulf cellular cargo and deliver them to the vacuole or lysosomes for degradation. Inspite of more than 40 AuTophaGy (ATG) proteins and several organelles as known membrane source, autophagosome biogenesis is not entirely understood. We recently have discovered that septins contribute to autophagosome biogenesis. Septins are GTP-binding proteins, usually localized at the bud neck region and are involved in cytokinesis. Here, we show that during autophagy prevalent conditions, septins traffic between different cellular compartments such as Golgi, mitochondria, endosomes, plasma membrane, and vacuolar membranes.



Similar content being viewed by others
References
An Z, Tassa A, Thomas C, Zhong R, Xiao G, Fotedar R, Tu BP, Klionsky DJ, Levine B (2014) Autophagy is required for G(1)/G(0) quiescence in response to nitrogen starvation in Saccharomyces cerevisiae. Autophagy 10:1702–1711. https://doi.org/10.4161/auto.32122
Au Yong JY, Wang YM, Wang Y (2016) The Nim1 kinase Gin4 has distinct domains crucial for septin assembly, phospholipid binding and mitotic exit. J Cell Sci 129:2744–2756. https://doi.org/10.1242/jcs.183160
Barral Y, Mermall V, Mooseker MS, Snyder M (2000) Compartmentalization of the cell cortex by septins is required for maintenance of cell polarity in yeast. Mol Cell 5:841–851
Barve G, Sridhar S, Aher A, Sahani MH, Chinchwadkar S, Singh S, K NL, McMurray MA, Manjithaya R (2018) Septins are involved at the early stages of macroautophagy in S. cerevisiae. J Cell Sci 13110.1242/jcs.209098
Bridges AA, Jentzsch MS, Oakes PW, Occhipinti P, Gladfelter AS (2016) Micron-scale plasma membrane curvature is recognized by the septin cytoskeleton. J Cell Biol 213:23–32. https://doi.org/10.1083/jcb.201512029
De Virgilio C, DeMarini DJ, Pringle JR (1996) SPR28, a sixth member of the septin gene family in Saccharomyces cerevisiae that is expressed specifically in sporulating cells. Microbiology 142(Pt 10):2897–2905. https://doi.org/10.1099/13500872-142-10-2897
Dobbelaere J, Barral Y (2004) Spatial coordination of cytokinetic events by compartmentalization of the cell cortex. Science 305:393–396. https://doi.org/10.1126/science.1099892
Feng Y, He D, Yao Z, Klionsky DJ (2014) The machinery of macroautophagy. Cell Res 24:24–41. https://doi.org/10.1038/cr.2013.168
Garcia G, Finnigan GC, Heasley LR, Sterling SM, Aggarwal A, Pearson CG, Nogales E, McMurray MA, Thorner J (2016) Assembly, molecular organization, and membrane-binding properties of development-specific septins. J Cell Biol 212:515–529. https://doi.org/10.1083/jcb.201511029
Glomb O, Gronemeyer T (2016) Septin organization and functions in budding yeast. Front Cell Dev Biol 410.3389/fcell.2016.00123
Imai K, Hao F, Fujita N, Tsuji Y, Oe Y, Araki Y, Hamasaki M, Noda T, Yoshimori T (2016) Atg9A trafficking through the recycling endosomes is required for autophagosome formation. J Cell Sci 129:3781–3791. https://doi.org/10.1242/jcs.196196
Johnson ES, Gupta AA (2001) An E3-like factor that promotes SUMO conjugation to the yeast septins. Cell 106:735–744
Kabeya Y, Kamada Y, Baba M, Takikawa H, Sasaki M, Ohsumi Y (2005) Atg17 functions in cooperation with Atg1 and Atg13 in yeast autophagy. Mol Biol Cell 16:2544–2553. https://doi.org/10.1091/mbc.E04-08-0669
Kadota J, Yamamoto T, Yoshiuchi S, Bi E, Tanaka K (2004) Septin ring assembly requires concerted action of polarisome components, a PAK kinase Cla4p, and the actin cytoskeleton in Saccharomyces cerevisiae. Mol Biol Cell 15:5329–5345. https://doi.org/10.1091/mbc.E04-03-0254
Kamada Y, Funakoshi T, Shintani T, Nagano K, Ohsumi M, Ohsumi Y (2000) Tor-mediated induction of autophagy via an Apg1 protein kinase complex. J Cell Biol 150:1507–1513
Kang H, Lew DJ (2017) How do cells know what shape they are? Curr Genet 63:75–77. https://doi.org/10.1007/s00294-016-0623-1
Legakis JE, Yen WL, Klionsky DJ (2007) A cycling protein complex required for selective autophagy. Autophagy 3:422–432
Longatti A, Tooze SA (2012) Recycling endosomes contribute to autophagosome formation. Autophagy 8:1682–1683. https://doi.org/10.4161/auto.21486
Longtine MS, Fares H, Pringle JR (1998) Role of the yeast Gin4p protein kinase in septin assembly and the relationship between septin assembly and septin function. J Cell Biol 143:719–736
Mari M, Griffith J, Rieter E, Krishnappa L, Klionsky DJ, Reggiori F (2010) An Atg9-containing compartment that functions in the early steps of autophagosome biogenesis. J Cell Biol 190:1005–1022. https://doi.org/10.1083/jcb.200912089
McMurray MA, Thorner J (2008) Septin stability and recycling during dynamic structural transitions in cell division and development. Curr Biol 18:1203–1208. https://doi.org/10.1016/j.cub.2008.07.020
Mitchell L, Lau A, Lambert JP, Zhou H, Fong Y, Couture JF, Figeys D, Baetz K (2011) Regulation of septin dynamics by the Saccharomyces cerevisiae lysine acetyltransferase NuA4. PLoS One 6:e25336. https://doi.org/10.1371/journal.pone.0025336
Mizushima N, Yoshimori T, Ohsumi Y (2011) The role of Atg proteins in autophagosome formation. Annu Rev Cell Dev Biol 27:107–132. https://doi.org/10.1146/annurev-cellbio-092910-154005
Oh Y, Bi E (2011) Septin structure and function in yeast and beyond. Trends Cell Biol 21:141–148. https://doi.org/10.1016/j.tcb.2010.11.006
Okada S, Leda M, Hanna J, Savage NS, Bi E, Goryachev AB (2013) Daughter cell identity emerges from the interplay of Cdc42, septins, and exocytosis. Dev Cell 26:148–161. https://doi.org/10.1016/j.devcel.2013.06.015
Orsi A, Razi M, Dooley HC, Robinson D, Weston AE, Collinson LM, Tooze SA (2012) Dynamic and transient interactions of Atg9 with autophagosomes, but not membrane integration, are required for autophagy. Mol Biol Cell 23:1860–1873. https://doi.org/10.1091/mbc.E11-09-0746
Ozsarac N, Bhattacharyya M, Dawes IW, Clancy MJ (1995) The SPR3 gene encodes a sporulation-specific homologue of the yeast CDC3/10/11/12 family of bud neck microfilaments and is regulated by ABFI. Gene 164:157–162
Papinski D, Kraft C (2014) Atg1 kinase organizes autophagosome formation by phosphorylating Atg9. Autophagy 10:1338–1340. https://doi.org/10.4161/auto.28971
Puri C, Renna M, Bento CF, Moreau K, Rubinsztein DC (2014) ATG16L1 meets ATG9 in recycling endosomes: additional roles for the plasma membrane and endocytosis in autophagosome biogenesis. Autophagy 10:182–184. https://doi.org/10.4161/auto.27174
Ravikumar B, Moreau K, Jahreiss L, Puri C, Rubinsztein DC (2010) Plasma membrane contributes to the formation of pre-autophagosomal structures. Nat Cell Biol 12:747–757. https://doi.org/10.1038/ncb2078
Reggiori F, Tucker KA, Stromhaug PE, Klionsky DJ (2004) The Atg1-Atg13 complex regulates Atg9 and Atg23 retrieval transport from the pre-autophagosomal structure. Dev Cell 6:79–90
Reggiori F, Shintani T, Nair U, Klionsky DJ (2005) Atg9 cycles between mitochondria and the pre-autophagosomal structure in yeasts. Autophagy 1:101–109
Sekito T, Kawamata T, Ichikawa R, Suzuki K, Ohsumi Y (2009) Atg17 recruits Atg9 to organize the pre-autophagosomal structure. Genes Cells 14:525–538. https://doi.org/10.1111/j.1365-2443.2009.01299.x
Setty SR, Shin ME, Yoshino A, Marks MS, Burd CG (2003) Golgi recruitment of GRIP domain proteins by Arf-like GTPase 1 is regulated by Arf-like GTPase 3. Curr Biol 13:401–404
Song K, Russo G, Krauss M (2016) Septins As Modulators of Endo-Lysosomal Membrane Traffic. Front Cell Dev Biol 4:124. https://doi.org/10.3389/fcell.2016.00124
Suzuki K, Kirisako T, Kamada Y, Mizushima N, Noda T, Ohsumi Y (2001) The pre-autophagosomal structure organized by concerted functions of APG genes is essential for autophagosome formation. EMBO J 20:5971–5981. https://doi.org/10.1093/emboj/20.21.5971
Takahashi Y, Iwase M, Konishi M, Tanaka M, Toh-e A, Kikuchi Y (1999) Smt3, a SUMO-1 homolog, is conjugated to Cdc3, a component of septin rings at the mother-bud neck in budding yeast. Biochem Biophys Res Commun 259:582–587. https://doi.org/10.1006/bbrc.1999.0821
Tang CS, Reed SI (2002) Phosphorylation of the septin cdc3 in g1 by the cdc28 kinase is essential for efficient septin ring disassembly. Cell Cycle 1:42–49
Tucker KA, Reggiori F, Dunn WA Jr, Klionsky DJ (2003) Atg23 is essential for the cytoplasm to vacuole targeting pathway and efficient autophagy but not pexophagy. J Biol Chem 278:48445–48452. https://doi.org/10.1074/jbc.M309238200
Versele M, Thorner J (2004) Septin collar formation in budding yeast requires GTP binding and direct phosphorylation by the PAK, Cla4. J Cell Biol 164:701–715. https://doi.org/10.1083/jcb.200312070
Zander S, Baumann S, Weidtkamp-Peters S, Feldbrugge M (2016) Endosomal assembly and transport of heteromeric septin complexes promote septin cytoskeleton formation. J Cell Sci 129:2778–2792. https://doi.org/10.1242/jcs.182824
Acknowledgements
We would like to thank Prof. Yoshinori Ohsumi, Tokyo Institute of Technology, Tokyo, for generously sharing 2xmCherry-Atg8 plasmid, Prof. Subba Rao, IISc, Bangalore, India, for the Golgi marker plasmid RFP-Imh1p-GRIP-pGS396 and Prof. Kausik Chakraborty, IGIB, India, for sharing Septin-GFP strains. This work was supported by Wellcome Trust/DBT India Alliance Intermediate Fellowship (509159-Z-09-Z) and JNCASR intramural funds to RM.
Author information
Authors and Affiliations
Contributions
GB and PS have performed the experiments. GB and RM wrote the manuscript.
Corresponding author
Additional information
Communicated by M. Kupiec.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Movie 1
: Cdc11-GFP cells were grown in YPD until it reaches to 0.6 to 0.8 OD600 and the SD-N medium was added (1OD600/ml). After 2h incubation in starvation medium cells were centrifuged at 20817g for 2 min and were mounted on agarose pads (made in SD-N medium). Time-lapse was carried out for 50 seconds with an interval of 10 seconds at RT. Scale bar 1µm (MOV 123 KB)
Movie 2
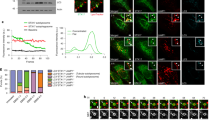
Cdc11-GFP cells were grown as mentioned in Fig 1. Cells were centrifuged at 20817g for 2 min and were mounted on agarose pads (made in SD-N medium). Time-lapse was carried out at a single Z plane for 300 seconds with an interval of 10 seconds at RT. Here first 150 seconds are shown. Arrows indicate Cdc11-GFP and FM4-64 stained endosomes simultaneously emerging from the plasma membrane. Scale bar 1µm (AVI 333 KB)
Rights and permissions
About this article
Cite this article
Barve, G., Sanyal, P. & Manjithaya, R. Septin localization and function during autophagy. Curr Genet 64, 1037–1041 (2018). https://doi.org/10.1007/s00294-018-0834-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-018-0834-8