Abstract
Key message
Global survey of plastid gene expression during fruit ripening in kiwifruit provides cis-elements for the future engineering of the plastid genome of kiwifruit.
A limitation in the application of plastid biotechnology for molecular farming is the low-level expression of transgenes in non-green plastids compared with photosynthetically active chloroplasts. Unlike other fruits, not all chloroplasts are transformed into chromoplasts during ripening of red-fleshed kiwifruit (Actinidia chinensis cv. Hongyang) fruits, which may make kiwifruit an ideal horticultural plant for recombinant protein production by plastid engineering. To identify cis-elements potentially triggering high-level transgene expression in edible tissues of the ‘Hongyang’ kiwifruit, here we report a comprehensive analysis of kiwifruit plastid gene transcription in green leaves and fruits at three different developmental stages. While transcripts of a few photosynthesis-related genes and most genetic system genes were substantially upregulated in green fruits compared with leaves, nearly all plastid genes were significantly downregulated at the RNA level during fruit development. Expression of a few genes remained unchanged, including psbA, the gene encoding the D1 polypeptide of photosystem II. However, PsbA protein accumulation decreased continuously during chloroplast-to-chromoplast differentiation. Analysis of post-transcriptional steps in mRNA maturation, including intron splicing and RNA editing, revealed that splicing and editing may contribute to regulation of plastid gene expression. Altogether, 40 RNA editing sites were verified, and 5 of them were newly discovered. Taken together, this study has generated a valuable resource for the analysis of plastid gene expression and provides cis-elements for future efforts to engineer the plastid genome of kiwifruit.





Similar content being viewed by others
References
Ahmad N, Michoux F, Lossl AG, Nixon PJ (2016) Challenges and perspectives in commercializing plastid transformation technology. J Exp Bot 67:5945–5960
Barkan A (2011) Expression of plastid genes: organelle-specific elaborations on a prokaryotic scaffold. Plant Physiol 155:1520–1532
Bock R (2000) Sense from nonsense: How the genetic information of chloroplasts is altered by RNA editing. Biochimie 82:549–557
Bock R (2015) Engineering plastid genomes: methods, tools, and applications in basic research and biotechnology. Annu Rev Plant Biol 66:211–241
Bock R (2017) Witnessing genome evolution: experimental reconstruction of endosymbiotic and horizontal gene transfer. Annu Rev Genet 51:1–22
Börner T, Aleynikova A, Zubo Y, Kusnetsov V (2015) Chloroplast RNA polymerases: role in chloroplast biogenesis. Biochim Biophys Acta 1847:761–769
Cahoon EB, Shanklin J, Ohlrogge JB (1992) Expression of a coriander desaturase results in petroselinic acid production in transgenic tobacco. Proc Natl Acad Sci USA 89:11184–11188
Caroca R, Howell KA, Hasse C, Ruf S, Bock R (2013) Design of chimeric expression elements that confer high-level gene activity in chromoplasts. Plant J 73:368–379
Corneille S, Lutz K, Maliga P (2000) Conservation of RNA editing between rice and maize plastids: are most editing events dispensable? Mol Gen Genet 264:419–424
Daniell H, Lin CS, Yu M, Chang WJ (2016) Chloroplast genomes: diversity, evolution, and applications in genetic engineering. Genome Biol 17:134
Daniell H, Jin S, Zhu XG, Gitzendanner MA, Soltis DE, Soltis PS (2021) Green giant—a tiny chloroplast genome with mighty power to produce high-value proteins: history and phylogeny. Plant Biotechnol J 19:430–447
Eberhard S, Drapier D, Wollman F (2002) Searching limiting steps in the expression of chloroplast-encoded proteins: relations between gene copy number, transcription, transcript abundance and translation rate in the chloroplast of Chlamydomonas reinhardtii. Plant J 31:149–160
Ferguson A, Huang H (2007) Genetic resources of kiwifruit: domestication and breeding. Hortic Rev 33:1–121
Greiner S, Sobanski J, Bock R (2015) Why are most organelle genomes transmitted maternally? BioEssays 37:80–94
Greiner S, Lehwark P, Bock R (2019) OrganellarGenomeDRAW (OGDRAW) version 1.3.1: expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res 47:W59–W64
Hajdukiewicz PT, Allison LA, Maliga P (1997) The two RNA polymerases encoded by the nuclear and the plastid compartments transcribe distinct groups of genes in tobacco plastids. EMBO J 16:4041–4048
Hertel S, Zoschke R, Neumann L, Qu YJ, Axmann IM, Schmitz-Linneweber C (2013) Multiple checkpoints for the expression of the chloroplast-encoded splicing factor MatK. Plant Physiol 163:1686–1698
Huang S, Ding J, Deng D, Tang W, Sun H, Liu D, Zhang L, Niu X, Zhang X, Meng M, Yu J, Liu J, Han Y, Shi W, Zhang D, Cao S, Wei Z, Cui Y, Xia Y, Zeng H, Bao K, Lin L, Min Y, Zhang H, Miao M, Tang X, Zhu Y, Sui Y, Li G, Sun H, Yue J, Sun J, Liu F, Zhou L, Lei L, Zheng X, Liu M, Huang L, Song J, Xu C, Li J, Ye K, Zhong S, Lu BR, He G, Xiao F, Wang HL, Zheng H, Fei Z, Liu Y (2013) Draft genome of the kiwifruit Actinidia chinensis. Nat Commun 4:2640
Ichinose M, Sugita M (2017) RNA editing and its molecular mechanism in plant organelles. Genes (basel) 8:5
Kahlau S, Bock R (2008) Plastid transcriptomics and translatomics of tomato fruit development and chloroplast-to-chromoplast differentiation: chromoplast gene expression largely serves the production of a single protein. Plant Cell 20:856–874
Kahlau S, Aspinall S, Gray JC, Bock R (2006) Sequence of the tomato chloroplast DNA and evolutionary comparison of solanaceous plastid genomes. J Mol Evol 63:194–207
Karcher D, Bock R (2002) Temperature sensitivity of RNA editing and intron splicing reactions in the plastid ndhB transcript. Curr Genet 41:48–52
Kim SC, Lee JW, Baek SH, Lee MW, Kang YJ (2018) The complete chloroplast genome sequence of Actinidia Rufa (Actinidiaceae). Mitochondrial DNA B 3:564–565
Kudla J, Bock R (1999) RNA editing in an untranslated region of the Ginkgo chloroplast genome. Gene 234:81–86
Kugita M, Yamamoto Y, Fujikawa T, Matsumoto T, Yoshinaga K (2003) RNA editing in hornwort chloroplasts makes more than half the genes functional. Nucleic Acids Res 31:2417–2423
Lan Y, Cheng L, Huang W, Cao Q, Zhou Z, Luo A, Hu G (2018) The complete chloroplast genome sequence of Actinidia kolomikta from north China. Conserv Genet Resour 10:475–477
Li W, Liu Y, Zeng S, Xiao G, Wang G, Wang Y, Peng M, Huang H (2015) Gene expression profiling of development and anthocyanin accumulation in kiwifruit (Actinidia chinensis) based on transcriptome sequencing. PLoS ONE 10:e0136439
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods 25:402–408
Maier RM, Neckermann K, Igloi GL, Kössel H (1995) Complete sequence of the maize chloroplast genome: gene content, hotspots of divergence and fine tuning of genetic information by transcript editing. J Mol Biol 251:614–628
Montefiori M, McGhie TK, Costa G, Ferguson AR (2005) Pigments in the fruit of red-fleshed kiwifruit (Actinidia chinensis and Actinidia deliciosa). J Agric Food Chem 53:9526–9530
Moreno JC, Tiller N, Diez M, Karcher D, Tillich M, Schottler MA, Bock R (2017) Generation and characterization of a collection of knock-down lines for the chloroplast Clp protease complex in tobacco. J Exp Bot 68:2199–2218
Nishiyama I, Fukuda T, Oota T (2005) Genotypic differences in chlorophyll, lutein, and -carotene contents in the fruits of Actinidia species. J Agric Food Chem 53:6403–6407
Oda K, Yamato K, Ohta E, Nakamura Y, Takemura M, Nozato N, Akashi K, Kanegae T, Ogura Y, Kohchi T, Ohyama K (1992) Gene organization deduced from the complete sequence of liverwort Marchantia polymorpha mitochondrial DNA. a primitive form of plant mitochondrial genome. J Mol Biol 223:1–7
Oey M, Lohse M, Kreikemeyer B, Bock R (2009) Exhaustion of the chloroplast protein synthesis capacity by massive expression of a highly stable protein antibiotic. Plant J 57:436–445
Palmer J (2003) The symbiotic birth and spread of plastids: how many times and whodunit? J Phycol 39:4–12
Petersen K, Schottler MA, Karcher D, Thiele W, Bock R (2011) Elimination of a group II intron from a plastid gene causes a mutant phenotype. Nucleic Acids Res 39:5181–5192
Ruwe H, Castandet B, Schmitz-Linneweber C, Stern DB (2013) Arabidopsis chloroplast quantitative editotype. FEBS Lett 587:1429–1433
Sadali NM, Sowden RG, Ling Q, Jarvis RP (2019) Differentiation of chromoplasts and other plastids in plants. Plant Cell Rep 38:803–818
Scharff LB, Bock R (2014) Synthetic biology in plastids. Plant J 78:783–798
Small ID, Schallenberg-Rudinger M, Takenaka M, Mireau H, Ostersetzer-Biran O (2020) Plant organellar RNA editing: what 30 years of research has revealed. Plant J 101:1040–1056
Stegemann S, Bock R (2006) Experimental reconstruction of functional gene transfer from the tobacco plastid genome to the nucleus. Plant Cell 18:2869–2878
Stern DB, Goldschmidt-Clermont M, Hanson MR (2010) Chloroplast RNA metabolism. Annu Rev Plant Biol 61:125–155
Stoppel R, Meurer J (2013) Complex RNA metabolism in the chloroplast: an update on the psbB operon. Planta 237:441–449
Valkov VT, Scotti N, Kahlau S, Maclean D, Grillo S, Gray JC, Bock R, Cardi T (2009) Genome-wide analysis of plastid gene expression in potato leaf chloroplasts and tuber amyloplasts: transcriptional and posttranscriptional control. Plant Physiol 150:2030–2044
Valkov V, Gargano D, Manna C, Formisano G, Dix P, Gray J, Scotti N, Cardi T (2011) High efficiency plastid transformation in potato and regulation of transgene expression in leaves and tubers by alternative 5’ and 3’ regulatory sequences. Transgenic Res 20:137–151
Villarreal AJ, Turmel M, Bourgouin-Couture M, Laroche J, Salazar Allen N, Li FW, Cheng S, Renzaglia K, Lemieux C (2018) Genome-wide organellar analyses from the hornwort Leiosporoceros dussii show low frequency of RNA editing. PLoS ONE 13:e0200491
Wakasugi T, Hirose T, Horihata M, Tsudzuki T, Kössel H, M. S, (1996) Creation of a novel protein-coding region at the RNA level in black pine chloroplasts: the pattern of RNA editing in the gymnosperm chloroplast is different from that in angiosperms. Proc Natl Acad Sci USA 93:8766–8770
Wang WC, Chen SY, Zhang XZ (2016) Chloroplast genome evolution in Actinidiaceae: clpP loss, heterogenous divergence and phylogenomic practice. PLoS ONE 11:e0162324
Wu H, Li M, Wang D, Liu H, Xu X (2019a) The complete chloroplast genome sequence of Actinidia Callosa Var. Henryi Mitochondrial DNA B 4:652–653
Wu H, Ma T, Kang M, Ai F, Zhang J, Dong G, Liu J (2019b) A high-quality Actinidia chinensis (kiwifruit) genome. Hortic Res 6:117
Wu L, Lan J, Xiang X, Xiang H, Jin Z, Khan S, Liu Y (2020) Transcriptome sequencing and endogenous phytohormone analysis reveal new insights in CPPU controlling fruit development in kiwifruit (Actinidia chinensis). PLoS ONE 15:e0240355
Yagi Y, Shiina T (2014) Recent advances in the study of chloroplast gene expression and its evolution. Front Plant Sci 5:61
Yang A, Liu S, Liu T, Hu M, Zhong Y, Liu L, Yu F (2019) The complete chloroplast genome sequence of Actinidia styracifolia C. F Liang Mitochondrial DNA B 5:90–91
Yao X, Tang P, Li Z, Li D, Liu Y, Huang H (2015) The first complete chloroplast genome sequences in Actinidiaceae: genome structure and comparative analysis. PLoS ONE 10:e0129347
Zandueta-Criado A, Bock R (2004) Surprising features of plastid ndhD transcripts: addition of non-encoded nucleotides and polysome association of mRNAs with an unedited start codon. Nucleic Acids Res 32:542–550
Zhang L, Li Z, Wang Y, Jiang Z, Wang S, Huang H (2010) Vitamin C flower color and ploidy variation of hybrids from a ploidy-unbalanced Actinidia interspecific cross and SSR characterization. Euphytica 175(133):143
Zhang J, Ruf S, Hasse C, Childs L, Scharff LB, Bock R (2012) Identification of cis-elements conferring high levels of gene expression in non-green plastids. Plant J 72:115–128
Zhang AD, Wang WQ, Tong Y, Li MJ, Grierson D, Ferguson I, Chen KS, Yin XR (2018) Transcriptome analysis identifies a zinc finger protein regulating starch degradation in kiwifruit. Plant Physiol 178:850–863
Zhou F, Karcher D, Bock R (2007) Identification of a plastid intercistronic expression element (IEE) facilitating the expression of stable translatable monocistronic mRNAs from operons. Plant J 52:961–972
Zhou F, Badillo-Corona JA, Karcher D, Gonzalez-Rabade N, Piepenburg K, Borchers AMI, Maloney AP, Kavanagh TA, Gray JC, Bock R (2008) High-level expression of human immunodeficiency virus antigens from the tobacco and tomato plastid genomes. Plant Biotechnol J 6:897–913
Acknowledgements
This research was supported by grants from the National Natural Science Foundation of China (31872035, 32071477), the Science and Technology Department of Hubei Province of China (2020CFA012) and Innovation Base for Introducing Talents of Discipline of Hubei Province (2019BJH021).
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Contributions
SL and JZ: project design; QC and PS: performing the experiments; SL and JZ: analysis of the data; SL and JZ: writing; RB: review and editing. All the authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors report no declarations of interest.
Additional information
Communicated by Teodoro Cardi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
299_2022_2840_MOESM1_ESM.pptx
Supplementary file1 (PPTX 1880 KB) Incorrect gene annotations in the published Actinidia plastome map (Yao et al. 2015). (A) The trnE-UUC needs to be annotated on the outer cycle. The trnM-CAU (B) and trnT-GGU (C) are reduntant. (D) Annotation of the rps12 5′ needs to be on the inner cycle. (E) The rps12 gene in IRB region is missing.
299_2022_2840_MOESM2_ESM.pptx
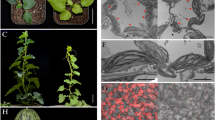
Supplementary file2 (PPTX 1880 KB) Kiwifruit samples representing different ripening stages. Cross-sections are shown for a young fruit (A), a turning fruit (B), and a mature fruit (C).
Rights and permissions
About this article
Cite this article
Chen, Q., Shen, P., Bock, R. et al. Comprehensive analysis of plastid gene expression during fruit development and ripening of kiwifruit. Plant Cell Rep 41, 1103–1114 (2022). https://doi.org/10.1007/s00299-022-02840-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-022-02840-7