Abstract
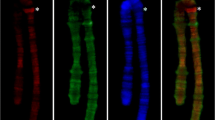
After applying proper deoxyribonucleic acid (DNA) probes, fluorescence in situ hybridization (FISH) showed that the 8/9 centromeres—one per chromatid of the male haploid complement (X0) of Pyrgomorpha conica grasshopper—colocalized at the spermatid blunt end, where the spermatozoa flagellum inserts. A bundle of aligned 4′,6-diamidino-2-phenylindole-positive chromatid scaffolds, which formed the central spermatid core, was observed after DNA breakage detection followed by FISH. Modular nature of scaffold DNA was occasionally evident. The technique also showed that in the early spermatid, the chromatid scaffolds lacked any DNA nick, whereas abundant breaks accumulated in the surrounding loops. Moreover, immunodetection showed that scaffold DNA participated in the formation of triplex DNA, while this configuration was absent from the loops. During spermatid maturation, triplex DNA disappeared from the scaffold in parallel with loop retraction, while protamines replace histones. Thus, the presence of triplex DNA in the chromatid scaffold correlates with the anchoring of expanded DNA loops to it. After loop retraction, the scaffolds of all chromatids coiled as a single unit in the spermatid head. This cooperative coiling produced enlargement and tilting of the distal telomeric signals, which were distributed along the spermatid head according to the length of each chromosome. We propose that specific DNA sequences dispersed throughout the whole chromatid fold forward and backward coaxially to chromatid length, forming individual scaffold modules whose linear assembly accounts for the minimum length of each individual chromatid. Finally, the core of the grasshopper male spermatid should be considered as a single chromosome in which the DNA scaffolds of the whole set of the nonhomologous chromosomes of the haploid complement are interconnected. This pattern of chromatin organization applies probably to other elongated spermatids.






Similar content being viewed by others
References
Burkholder GD, Latimer LJ, Lee JS (1991) Immunofluorescence localization of triplex DNA in polytene chromosomes of Chironomus and Drosophila. Chromosoma 101:11–18
Dorus S, Busby SA, Gerike U, Shabanowitz J, Hunt DF, Karr TL (2006) Genomic and functional evolution of the Drosophila melanogaster sperm proteome. Nat Genet 38:1440–1445
Earnshaw WC, Halligan B, Cooke CC, Heck MMS (1985) Topoisomerase II is a structural component of mitotic chromosome scaffolds. J Cell Biol 100:1706–1715
Felsenfeld G, Rich A (1957) Studies on the formation of two- and three-stranded polyribonucleotides. Biochim Biophys Acta 26:457–468
Fernández JL, Goyanes VJ, Ramiro-Díaz J, Gosálvez J (1998) Application of FISH for in situ detection and quantification of DNA breakage. Cytogenet Cell Genet 82:251–256
Fernández JL, Goyanes V, Gosálvez J (2002) DNA breakage detection-FISH (DBD-FISH). In: Rautenstrauss B, Liehr T (eds) FISH technology. Springer, New York, pp 282–290
Gassmann R, Henzing AJ, Earnshaw WC (2005) Novel components of human mitotic chromosomes identified by proteomic analysis of the chromosome scaffold fraction. Chromosoma 113:385–397
Goñi JR, De la Cruz X, Orozco M (2004) Triplex-forming oligonucleotide target sequences in the human genome. Nucleic Acids Res 32:354–360
Goobes R, Cohen O, Minsky A (2002) Unique condensation patterns of triplex DNA: physical aspects and physiological implications. Nucleic Acids Res 30:2154–2161
Govin J, Caron C, Lestrat C, Rousseaux S, Khochbin S (2004) The role of histones in chromatin remodeling during mammalian spermiogenesis. Eur J Biochem 271:3459–3469
Hennig W (2003) Chromosomal proteins in the spermatogenesis of Drosophila. Chromosoma 111:489–494
Jayaramaiah-Raja S, Renkawitz-Pohl R (2005) Replacement by Drosophila melanogaster protamines and Mst77F of histones during chromatin condensation in late spermatids and role of sesame in the removal of these proteins from the male pronucleus. Mol Cell Biol 25:6165–6177
Kireeva N, Lakonishok M, Kireev I, Hirano T, Belmont AS (2004) Visualization of early chromosome condensation: a hierarchical folding, axial glue model of chromosome structure. J Cell Biol 166:775–785
Kleinschmidt AK (1968) Monolayer techniques in electron microscopy of nucleic acid molecules. Method Enzymol 12B:361–377 (edited by Grossman LL, Moldave K)
Labergé R-M, Boissonneault G (2005) On the nature and origin of DNA strand breaks in elongating spermatids. Biol Reprod 73:289–296
López-Fernández C, Pradillo E, Zabal-Aguirre M, Fernández JL, García de la Vega C, Gosálvez J (2004) Telomeric and interstitial telomeric-like DNA sequences in Orthoptera genomes. Genome 47:757–763
López-Fernández C, Arroyo P, Fernández JL, Gosálvez J (2006) Interstitial telomeric sequence blocks in constitutive pericentromeric heterochromatin from Pyrgomorpha conica (Orthoptera) are enriched in constitutive alkali-labile sites. Mutat Res 599:36–44
Mullinger AM, Johnson RT (1980) Packing DNA into chromosomes. J Cell Sci 46:61–86
Ohno M, Fukagawa T, Lee JS, Ikemura T (2002) Triplex-forming DNAs in the human interphase nucleus visualized in situ by polypurine/polypirimidine DNA probes and antitriplex antibodies. Chromosoma 111:201–213
Pallota D, Tessier A (1976) Amino acid composition of sperm histones in the house cricket Acheta domestica. Can J Biochem 54:56–61
Paulson JR, Laemmli UK (1977) The structure of histone-depleted metaphase chromosomes. Cell 12:817–828
Poirier MG, Marko JF (2002) Mitotic chromosomes are chromatin networks without a mechanically continuous protein scaffold. Proc Natl Acad Sci USA 99:15393–15397
Rathke C, Baarends W, Jayaramaiah-Raja WM, Bartkuhn S, Renkawitz M, Renkawitz-Pohl R (2007) Transition for a nucleosome-based to a protamine-based chromatin configuration during spermiogenesis in Drosophila. J Cell Sci 120:1689–16700
Ren J, Bailly C, Chaires JB (2000) NB-506, an indolocarbazole topoisomerase I inhibitor, binds preferentially to triplex DNA. FEBS Lett 470:355–359
Sakkas D, Manicardi GC, Bianchi PG, Bizzaro D, Bianchi U (1995) Relationship between the presence of endogenous nicks and sperm chromatin packaging in maturing and fertilizing mouse spermatozoa. Biol Reprod 52:1149–1155
Sheval EV, Polyakov VY (2006) Visualization of the chromosome scaffold and intermediates of loop domain compaction in extracted mitotic cells. Cell Biol Int 30:1028–1040
Spitzner JR, Chung IK, Muller MT (1995) Determination of 5′ and 3′ DNA triplex interference boundaries reveals the core DNA binding sequence for topoisomerase II. J Biol Chem 270:5932–5943
Takata M, Sasaki MS, Sonoda E, Morrison C, Hashimoto M, Utsumi H, Yamaguchi-Iwai Y, Shinohara A, Takeda S (1998) Homologous recombination and non-homologous end-joining pathways of DNA double-straand break repair have overlapping roles in the maintenance of chromosomal integrity in vertebrate cells. EMBO J 17:5497–5508
Tessier A, Pallota D (1973) Analysis of basic proteins during spermatogenesis in the cricket, Acheta domestica. Exp Cell Res 82:103–110
Ward WS (1993) Deoxyribonucleic acid loop domain tertiary structure in mammalian spermatozoa. Biol Reprod 48:1193–1201
Wray W, Stubblefield E (1970) A new method for the rapid isolation of chromosomes, mitotic apparatus, or nuclei from mammalian fibroblasts at near neutral pH. Exp Cell Res 59:469–478
Wykes SM, Krawetz SA (2003) The structural organization of sperm chromatin. J Biol Chem 278:29471–29477
Acknowledgments
We thank Prof. J Lee and Prof GD Burkholder for generously providing us with the triplex DNA antibodies. The present work has been supported by the Dirección General de Investigación of the Spanish Education and Science Ministry (Projects BFU2007-66340/BFI, BFU2004-03071, BMC2003-05751, BOS2003-04263, and BOS2002-00232). AC was recipient of a fellowship from the Spanish Foreign Office (AECI, Spanish Agency for International Cooperation), which we also greatly acknowledge.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Gerbi
Rights and permissions
About this article
Cite this article
Černá, A., López-Fernández, C., Fernández, J.L. et al. Triplex configuration in the nick-free DNAs that constitute the chromosomal scaffolds in grasshopper spermatids. Chromosoma 117, 15–24 (2008). https://doi.org/10.1007/s00412-007-0121-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-007-0121-9