Abstract
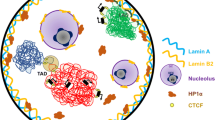
The past decades have provided remarkable insights into how the eukaryotic cell nucleus and the genome within it are organized. The combined use of imaging, biochemistry and molecular biology approaches has revealed several basic principles of nuclear architecture and function, including the existence of chromatin domains of various sizes, the presence of a large number of non-membranous intranuclear bodies, non-random positioning of genes and chromosomes in 3D space, and a prominent role of the nuclear lamina in organizing genomes. Despite this tremendous progress in elucidating the biological properties of the cell nucleus, many questions remain. Here, we highlight some of the key open areas of investigation in the field of nuclear organization and genome architecture with a particular focus on the mechanisms and principles of higher-order genome organization, the emerging role of liquid phase separation in cellular organization, and the functional role of the nuclear lamina in physiological processes.
Similar content being viewed by others
References
Adamson B, Norman TM, Jost M et al (2016) A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167:1867–1882.e21. https://doi.org/10.1016/j.cell.2016.11.048
Aguzzi A, Altmeyer M (2016) Phase separation: linking cellular compartmentalization to disease. Trends Cell Biol 26:547–558. https://doi.org/10.1016/j.tcb.2016.03.004
Ahmed S, Brickner DG, Light WH et al (2010) DNA zip codes control an ancient mechanism for gene targeting to the nuclear periphery. Nat Cell Biol 12:111–118. https://doi.org/10.1038/ncb2011
Amendola M, van Steensel B (2015) Nuclear lamins are not required for lamina-associated domain organization in mouse embryonic stem cells. EMBO Rep 16:610–617. https://doi.org/10.15252/embr.201439789
Atlasi Y, Stunnenberg HG (2017) The interplay of epigenetic marks during stem cell differentiation and development. Nat Rev Genet 18:643–658
Banani SF, Lee HO, Hyman AA, Rosen MK (2017) Biomolecular condensates: organizers of cellular biochemistry. Nat Rev Mol Cell Biol 18:285–298. https://doi.org/10.1038/nrm.2017.7
Barutcu AR, Maass PG, Lewandowski JP et al (2018) A TAD boundary is preserved upon deletion of the CTCF-rich Firre locus. Nat Commun 9:1444. https://doi.org/10.1038/s41467-018-03614-0
Beliveau BJ, Joyce EF, Apostolopoulos N et al (2012) Versatile design and synthesis platform for visualizing genomes with Oligopaint FISH probes. Proc Natl Acad Sci USA 109:21301–21306. https://doi.org/10.1073/pnas.1213818110
Beliveau BJ, Boettiger AN, Avendano MS et al (2015) Single-molecule super-resolution imaging of chromosomes and in situ haplotype visualization using Oligopaint FISH probes. Nat Commun 6:7147. https://doi.org/10.1038/ncomms8147
Berman BP, Weisenberger DJ, Aman JF et al (2011) Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina-associated domains. Nat Genet 44:40–46. https://doi.org/10.1038/ng.969
Bickmore WA (2013) The spatial organization of the human genome. Annu Rev Genomics Hum Genet 14:67–84. https://doi.org/10.1146/annurev-genom-091212-153515
Boban M, Foisner R (2016) Degradation-mediated protein quality control at the inner nuclear membrane. Nucleus 7:41–49. https://doi.org/10.1080/19491034.2016.1139273
Boettiger AN, Bintu B, Moffitt JR et al (2016) Super-resolution imaging reveals distinct chromatin folding for different epigenetic states. Nature 529:418–422. https://doi.org/10.1038/nature16496
Bonev B, Cavalli G (2016) Organization and function of the 3D genome. Nat Rev Genet 17:661–678. https://doi.org/10.1038/nrg.2016.112
Borden J, Manuelidis L (1988) Movement of the X chromosome in epilepsy. Science 242:1687–1691. https://doi.org/10.1126/science.3201257
Bouwman BAM, de Laat W (2015) Getting the genome in shape: The formation of loops, domains and compartments. Genome Biol 16:154. https://doi.org/10.1186/s13059-015-0730-1
Brangwynne CP, Mitchison TJ, Hyman AA (2011) Active liquid-like behavior of nucleoli determines their size and shape in Xenopus laevis oocytes. Proc Natl Acad Sci USA 108:4334–4339. https://doi.org/10.1073/pnas.1017150108
Briand N, Guénantin AC, Jeziorowska D et al (2018) The lipodystrophic hotspot lamin A p.R482W mutation deregulates the mesodermal inducer T/Brachyury and early vascular differentiation gene networks. Hum Mol Genet 27:1447–1459. https://doi.org/10.1093/hmg/ddy055
Brickner DG, Cajigas I, Fondufe-Mittendorf Y et al (2007) H2A.Z-mediated localization of genes at the nuclear periphery confers epigenetic memory of previous transcriptional state. PLoS Biol 5:704–716. https://doi.org/10.1371/journal.pbio.0050081
Butler JS, Koutelou E, Schibler AC, Dent SYR (2012) Histone-modifying enzymes: Regulators of developmental decisions and drivers of human disease. Epigenomics 4:163–177. https://doi.org/10.2217/epi.12.3
Cao K, Graziotto JJ, Blair CD et al (2011) Rapamycin reverses cellular phenotypes and enhances mutant protein clearance in Hutchinson-Gilford progeria syndrome cells. Sci Transl Med 3:89. https://doi.org/10.1126/scitranslmed.3002346
Casolari JM, Brown CR, Komili S et al (2004) Genome-wide localization of the nuclear transport machinery couples transcriptional status and nuclear organization. Cell 117:427–439. https://doi.org/10.1016/S0092-8674(04)00448-9
Cenni V, Capanni C, Columbaro M et al (2011) Autophagic degradation of farnesylated prelamin A as a therapeutic approach to lamin-linked progeria. Eur J Histochem 55:200–205. https://doi.org/10.4081/ejh.2011.e36
Chen H, Zheng X, Zheng Y (2014) Age-associated loss of lamin-B leads to systemic inflammation and gut hyperplasia. Cell 159:829–843. https://doi.org/10.1016/j.cell.2014.10.028
Chen H, Levo M, Barinov L et al (2018) Dynamic interplay between enhancer–promoter topology and gene activity. Nat Genet 50:1296–1303. https://doi.org/10.1038/s41588-018-0175-z
Cho WK, Spille JH, Hecht M et al (2018) Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science 361:412–415. https://doi.org/10.1126/science.aar4199
Chong S, Dugast-darzacq C, Liu Z et al (2018) Imaging dynamic and selective low-complexity domain interactions that control gene transcription. Science 361:6400. https://doi.org/10.1126/science.aar2555
Cremer T, Cremer C (2001) Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat Rev Genet 2:292–301. https://doi.org/10.1038/35066075
Cremer M, Küpper K, Wagler B et al (2003) Inheritance of gene density-related higher order chromatin arrangements in normal and tumor cell nuclei. J Cell Biol 162:809–820. https://doi.org/10.1083/jcb.200304096
Cuervo AM, Wong E (2014) Chaperone-mediated autophagy: roles in disease and aging. Cell Res 24:92–104. https://doi.org/10.1038/cr.2013.153
De Sandre-Giovannoli A, Bernard R, Cau P et al (2003) Lamin A truncation in Hutchinson-Gilford progeria. Science 300:2055. https://doi.org/10.1126/science.1084125
de Groot R, Luthi J, Lindsay H et al (2018) Large-scale image-based profiling of single-cell phenotypes in arrayed CRISPR-Cas9 gene perturbation screens. Mol Syst Biol 14:e8064
Dechat T, Shimi T, Adam SA et al (2007) Alterations in mitosis and cell cycle progression caused by a mutant lamin A known to accelerate human aging. Proc Natl Acad Sci USA 104:4955–4960. https://doi.org/10.1073/pnas.0700854104
Dekker J, Misteli T (2015) Long-range chromatin interactions. Cold Spring Harb Perspect Biol 7:10. https://doi.org/10.1101/cshperspect.a019356
Denholtz M, Bonora G, Chronis C et al (2013) Long-range chromatin contacts in embryonic stem cells reveal a role for pluripotency factors and polycomb proteins in genome organization. Cell Stem Cell 13:602–616. https://doi.org/10.1016/j.stem.2013.08.013
Dietzel S, Zolghadr K, Hepperger CBA (2004) Differential large-scale chromatin compaction and intranuclear positioning of transcribed versus non-transcribed transgene arrays containing beta-globin regulatory sequences. J Cell Sci 117:4603–4614. https://doi.org/10.1242/jcs.01330
Dittmer T, Misteli T (2011) The lamin protein family. Genome Biol 12:5. https://doi.org/10.1186/gb-2011-12-5-222
Dittmer TA, Sahni N, Kubben N et al (2014) Systematic identification of pathological lamin A interactors. Mol Biol Cell 25:1493–1510. https://doi.org/10.1091/mbc.E14-02-0733
Dixit A, Parnas O, Li B et al (2016) Perturb-seq: Dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167:1853–1866.e17. https://doi.org/10.1016/j.cell.2016.11.038
Dixon JR, Selvaraj S, Yue F et al (2012) Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485:376–380. https://doi.org/10.1038/nature11082
Dixon JR, Jung I, Selvaraj S et al (2015) Chromatin architecture reorganization during stem cell differentiation. Nature 518:331–336. https://doi.org/10.1038/nature14222
Dixon JR, Gorkin DU, Ren B (2016) Chromatin domains: The unit of chromosome organization. Mol Cell 62:668–680. https://doi.org/10.1016/j.molcel.2016.05.018
Dou Z, Xu C, Donahue G et al (2015) Autophagy mediates degradation of nuclear lamina. Nature 527:105–109. https://doi.org/10.1038/nature15548
Dundr M (2012) Nuclear bodies: Multifunctional companions of the genome. Curr Opin Cell Biol 24:415–422. https://doi.org/10.1016/j.ceb.2012.03.010
Elzeneini E, Wickström SA (2017) Lipodystrophic laminopathy: Lamin A mutation relaxes chromatin architecture to impair adipogenesis. J Cell Biol 216:2607–2610. https://doi.org/10.1083/jcb.201707090
Erdel F, Rippe K (2018) Formation of chromatin subcompartments by phase separation. Biophys J 114:2262–2270. https://doi.org/10.1016/j.bpj.2018.03.011
Eriksson M, Brown WT, Gordon LB et al (2003) Recurrent de novo point mutations in lamin A cause Hutchinson-Gilford progeria syndrome. Nature 423:293–298. https://doi.org/10.1038/nature01629
Evers B, Jastrzebski K, Heijmans JPM et al (2016) CRISPR knockout screening outperforms shRNA and CRISPRi in identifying essential genes. Nat Biotechnol 34:631–633. https://doi.org/10.1038/nbt.3536
Feric M, Vaidya N, Harmon TS et al (2016) Coexisting liquid phases underlie nucleolar subcompartments. Cell 165:1686–1697. https://doi.org/10.1016/j.cell.2016.04.047
Ferté C, André F, Soria JC (2010) Molecular circuits of solid tumors: Prognostic and predictive tools for bedside use. Nat Rev Clin Oncol 7:367–380
Finlan LE, Sproul D, Thomson I et al (2008) Recruitment to the nuclear periphery can alter expression of genes in human cells. PLoS Genet 4:3. https://doi.org/10.1371/journal.pgen.1000039
Foresti O, Rodriguez-Vaello V, Funaya C, Carvalho P (2014) Quality control of inner nuclear membrane proteins by the Asi complex. Science 751:751–756. https://doi.org/10.1126/science.1255638
Fraser J, Ferrai C, Chiariello AM et al (2015a) Hierarchical folding and reorganization of chromosomes are linked to transcriptional changes in cellular differentiation. Mol Syst Biol 11:852–852. https://doi.org/10.15252/msb.20156492
Fraser J, Williamson I, Bickmore WA, Dostie J (2015b) An overview of genome organization and how we got there: from FISH to Hi-C. Microbiol Mol Biol Rev 79:347–372. https://doi.org/10.1128/MMBR.00006-15
Freire-Pritchett P, Schoenfelder S, Várnai C et al (2017) Global reorganisation of cis-regulatory units upon lineage commitment of human embryonic stem cells. Elife 6:e21926. https://doi.org/10.7554/eLife.21926
Gabriel D, Roedl D, Gordon LB, Djabali K (2015) Sulforaphane enhances progerin clearance in Hutchinson-Gilford progeria fibroblasts. Aging Cell 14:78–91. https://doi.org/10.1111/acel.12300
Galganski L, Urbanek MO, Krzyzosiak WJ (2017) Nuclear speckles: Molecular organization, biological function and role in disease. Nucleic Acids Res 45:10350–10368. https://doi.org/10.1093/nar/gkx759
Gasperini M, Findlay GM, McKenna A et al (2017) CRISPR/Cas9-Mediated scanning for regulatory elements required for HPRT1 expression via thousands of large, programmed genomic deletions. Am J Hum Genet 101:192–205. https://doi.org/10.1016/j.ajhg.2017.06.010
Geyer PK, Vitalini MW, Wallrath LL (2011) Nuclear organization: taking a position on gene expression. Curr Opin Cell Biol 23:354–359. https://doi.org/10.1016/j.ceb.2011.03.002
Gilbert LA, Horlbeck MA, Adamson B et al (2014) Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159:647–661. https://doi.org/10.1016/j.cell.2014.09.029
Giorgetti L, Heard E (2016) Closing the loop: 3C versus DNA FISH. Genome Biol 17:215. https://doi.org/10.1186/s13059-016-1081-2
Gonzalez-Sandoval A, Towbin BD, Kalck V et al (2015) Perinuclear anchoring of H3K9-methylated chromatin stabilizes induced cell fate in C. elegans embryos. Cell 163:1333–1347. https://doi.org/10.1016/j.cell.2015.10.066
Grob S, Cavalli G (2018) Technical review: A hitchhiker’s guide to chromosome conformation capture. Methods Mol Biol 1675:233–246. https://doi.org/10.1007/978-1-4939-7318-7_14
Gruenbaum Y, Foisner R (2015) Lamins: Nuclear intermediate filament proteins with fundamental functions in nuclear mechanics and genome regulation. Annu Rev Biochem 84:131–164. https://doi.org/10.1146/annurev-biochem-060614-034115
Guelen L, Pagie L, Brasset E et al (2008) Domain organization of human chromosomes revealed by mapping of nuclear lamina interactions. Nature 453:948–951. https://doi.org/10.1038/nature06947
Guo Y, Xu Q, Canzio D et al (2015) CRISPR inversion of CTCF sites alters genome topology and enhancer/promoter function. Cell 162:900–910. https://doi.org/10.1016/j.cell.2015.07.038
Harr JC, Luperchio TR, Wong X et al (2015) Directed targeting of chromatin to the nuclear lamina is mediated by chromatin state and A-type lamins. J Cell Biol 208:33–52. https://doi.org/10.1083/jcb.201405110
Hennig S, Kong G, Mannen T et al (2015) Prion-like domains in RNA binding proteins are essential for building subnuclear paraspeckles. J Cell Biol 210:529–539. https://doi.org/10.1083/jcb.201504117
Henser-Brownhill T, Monserrat J, Scaffidi P (2017) Generation of an arrayed CRISPR-Cas9 library targeting epigenetic regulators: from high-content screens to in vivo assays. Epigenetics 12:1065–1075. https://doi.org/10.1080/15592294.2017.1395121
Hetzer MW (2010) The role of the nuclear pore complex in aging of post-mitotic cells. Aging (Albany NY) 2:74–75. https://doi.org/10.18632/aging.100125
Hewitt SL, High FA, Reiner SL et al (2004) Nuclear repositioning marks the selective exclusion of lineage-inappropriate transcription factor loci during T helper cell differentiation. Eur J Immunol 34:3604–3613. https://doi.org/10.1002/eji.200425469
Hnisz D, Shrinivas K, Young RA et al (2017) A phase separation model for transcriptional control. Cell 169:13–23. https://doi.org/10.1016/j.cell.2017.02.007
Hyman AA, Weber CA, Ulicher F (2014) Liquid-liquid phase separation in biology. Annu Rev Cell Dev Biol 30:39–58. https://doi.org/10.1146/annurev-cellbio-100913-013325
Jain A, Vale RD (2017) RNA phase transitions in repeat expansion disorders. Nature 546:243–247. https://doi.org/10.1038/nature22386
Ji X, Dadon DB, Powell BE et al (2016) 3D Chromosome regulatory landscape of human pluripotent cells. Cell Stem Cell 18:262–275. https://doi.org/10.1016/j.stem.2015.11.007
Joyce EF, Williams BR, Xie T, Wu C-T (2012) Identification of genes that promote or antagonize somatic homolog pairing using a high-throughput FISH-based screen. PLoS Genet 8:e1002667. https://doi.org/10.1371/journal.pgen.1002667
Kemeny S, Tatout C, Salaun G et al (2018) Spatial organization of chromosome territories in the interphase nucleus of trisomy 21 cells. Chromosoma 127:247–259. https://doi.org/10.1007/s00412-017-0653-6
Khanna R, Krishnamoorthy V, Parnaik VK (2018) E3 ubiquitin ligase RNF123 targets lamin B1 and lamin-binding proteins. FEBS J 285:2243–2262. https://doi.org/10.1111/febs.14477
Khmelinskii A, Blaszczak E, Pantazopoulou M et al (2014) Protein quality control at the inner nuclear membrane. Nature 516:410–413. https://doi.org/10.1038/nature14096
Kind J, van Steensel B (2010) Genome-nuclear lamina interactions and gene regulation. Curr Opin Cell Biol 22:320–325
Kind J, Pagie L, Ortabozkoyun H et al (2013) Single-cell dynamics of genome-nuclear lamina interactions. Cell 153:178–192. https://doi.org/10.1016/j.cell.2013.02.028
Kind J, Pagie L, De Vries SS et al (2015) Genome-wide maps of nuclear lamina interactions in single human cells. Cell 163:134–147. https://doi.org/10.1016/j.cell.2015.08.040
Kirby TJ, Lammerding J (2018) Emerging views of the nucleus as a cellular mechanosensor. Nat Cell Biol 20:373–381. https://doi.org/10.1038/s41556-018-0038-y
Korfali N, Wilkie GS, Swanson SK et al (2012) The nuclear envelope proteome differs notably between tissues. Nucl (United States) 3:552–564. https://doi.org/10.4161/nucl.22257
Kosak ST, Skok JA, Medina KL et al (2002) Subnuclear compartmentalization of immunoglobulin loci during lymphocyte development. Science 296:158–162. https://doi.org/10.1126/science.1068768
Krishnamoorthy V, Khanna R, Parnaik VK (2018) E3 ubiquitin ligase HECW2 targets PCNA and lamin B1. Biochim Biophys Acta Mol Cell Res 1865:1088–1104. https://doi.org/10.1016/j.bbamcr.2018.05.008
Krumm A, Duan Z (2018) Understanding the 3D genome: Emerging impacts on human disease. Semin Cell Dev Biol S 1084-9521:30592–X. https://doi.org/10.1016/j.semcdb.2018.07.004
Kubben N, Misteli T (2017) Shared molecular and cellular mechanisms of premature ageing and ageing-associated diseases. Nat Rev Mol Cell Biol 18:595–609. https://doi.org/10.1038/nrm.2017.68
Kubben N, Zhang W, Wang L et al (2016) Repression of the antioxidant NRF2 pathway in premature aging. Cell 165:1361–1374. https://doi.org/10.1016/j.cell.2016.05.017
Lakadamyali M, Cosma MP (2015) Advanced microscopy methods for visualizing chromatin structure. FEBS Lett 589:3023–3030. https://doi.org/10.1016/j.febslet.2015.04.012
Lemaître C, Bickmore WA (2015) Chromatin at the nuclear periphery and the regulation of genome functions. Histochem Cell Biol 144:111–122. https://doi.org/10.1007/s00418-015-1346-y
Lenain C, Gusyatiner O, Douma S et al (2015) Autophagy-mediated degradation of nuclear envelope proteins during oncogene-induced senescence. Carcinogenesis 36:1263–1274. https://doi.org/10.1093/carcin/bgv124
Leshner M, Devine M, Roloff GW et al (2016) Locus-specific gene repositioning in prostate cancer. Mol Biol Cell 27:236–246. https://doi.org/10.1091/mbc.E15-05-0280
Lopez AD, Mathers CD, Ezzati M et al (2006) Global and regional burden of disease and risk factors, 2001: systematic analysis of population health data. Lancet 367:1747–1757. https://doi.org/10.1016/S0140-6736(06)68770-9
López-Otín C, Blasco MA, Partridge L et al (2013) The hallmarks of aging. Cell 153:1194–1217. https://doi.org/10.1016/j.cell.2013.05.039
Lupiáñez DG, Kraft K, Heinrich V et al (2015) Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161:1012–1025. https://doi.org/10.1016/j.cell.2015.04.004
Meaburn KJ (2016) Spatial genome organization and its emerging role as a potential diagnosis tool. Front Genet 7:134. https://doi.org/10.3389/fgene.2016.00134
Meaburn KJ, Cabuy E, Bonne G et al (2007) Primary laminopathy fibroblasts display altered genome organization and apoptosis. Aging Cell 6:139–153. https://doi.org/10.1111/j.1474-9726.2007.00270.x
Meaburn KJ, Gudla PR, Khan S et al (2009) Disease-specific gene repositioning in breast cancer. J Cell Biol 187:801–812. https://doi.org/10.1083/jcb.200909127
Meaburn KJ, Agunloye O, Devine M et al (2016a) Tissue-of-origin-specific gene repositioning in breast and prostate cancer. Histochem Cell Biol 145:433–446. https://doi.org/10.1007/s00418-015-1401-8
Meaburn KJ, Burman B, Misteli T (2016b) Spatial Genome Organization and Disease. In: Bazett-Jones DP, Dellaire G (eds) The Functional Nucleus. Springer International Publishing, Cham, pp 101–125
Mewborn SK, Puckelwartz MJ, Abuisneineh F et al (2010) Altered chromosomal positioning, compaction, and gene expression with a lamin A/C gene mutation. PLoS One 5:e14342. https://doi.org/10.1371/journal.pone.0014342
Mishra A, Hawkins RD (2017) Three-dimensional genome architecture and emerging technologies: Looping in disease. Genome Med 9:87. https://doi.org/10.1186/s13073-017-0477-2
Misteli T (2007) Beyond the sequence: Cellular organization of genome function. Cell 128:787–800. https://doi.org/10.1016/j.cell.2007.01.028
Morgan SL, Mariano NC, Bermudez A et al (2017) Manipulation of nuclear architecture through CRISPR-mediated chromosomal looping. Nat Commun 8:15993. https://doi.org/10.1038/ncomms15993
Morgens DW, Deans RM, Li A, Bassik MC (2016) Systematic comparison of CRISPR/Cas9 and RNAi screens for essential genes. Nat Biotechnol 34:634–636. https://doi.org/10.1038/nbt.3567
Nagano T, Lubling Y, Stevens TJ et al (2013) Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502:59–64. https://doi.org/10.1038/nature12593
Nagano T, Lubling Y, Várnai C et al (2017) Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature 547:61–67. https://doi.org/10.1038/nature23001
Nakamura N (2011) The role of the transmembrane RING finger proteins in cellular and organelle function. Membranes (Basel) 1:354–393. https://doi.org/10.3390/membranes1040354
Narlikar GJ, Larson AG, Elnatan D et al (2017) Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 547:236–240. https://doi.org/10.1038/nature22822
Nielsen JA (2002) Nuclear organization in differentiating oligodendrocytes. J Cell Sci 115:4071–4079. https://doi.org/10.1242/jcs.00103
Nora EP, Lajoie BR, Schulz EG et al (2012) Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485:381–385. https://doi.org/10.1038/nature11049
Olive M, Harten I, Mitchell R et al (2010) Cardiovascular pathology in Hutchinson-Gilford progeria: Correlation with the vascular pathology of aging. Arterioscler Thromb Vasc Biol 30:2301–2309. https://doi.org/10.1161/ATVBAHA.110.209460
Ou HD, Phan S, Deerinck TJ et al (2017) ChromEMT: Visualizing 3D chromatin structure and compaction in interphase and mitotic cells. Science 357:6349. https://doi.org/10.1126/science.aag0025
Pajares M, Rojo AI, Arias E et al (2018) Transcription factor NFE2L2/NRF2 modulates chaperone-mediated autophagy through the regulation of LAMP2A. Autophagy 14:1310–1322. https://doi.org/10.1080/15548627.2018.1474992
Parfrey LW, Lahr DJG, Katz LA (2008) The dynamic nature of eukaryotic genomes. Mol Biol Evol 25:787–794. https://doi.org/10.1093/molbev/msn032
Park C, Suh Y, Cuervo AM (2015) Regulated degradation of Chk1 by chaperone-mediated autophagy in response to DNA damage. Nat Commun 6:6823. https://doi.org/10.1038/ncomms7823
Paz N, Zabala A, Royo F et al (2013) Combined Fluorescent-Chromogenic In Situ Hybridization for Identification and Laser Microdissection of Interphase Chromosomes. PLoS One 8:e60238. https://doi.org/10.1371/journal.pone.0060238
Paz N, Felipe-Blanco I, Royo F et al (2015) Expression of the DYRK1A gene correlates with its 3D positioning in the interphase nucleus of Down syndrome cells. Chromosom Res 23:285–298. https://doi.org/10.1007/s10577-015-9467-7
Pegoraro G, Misteli T (2017) High-throughput imaging for the discovery of cellular mechanisms of disease. Trends Genet 33:604–615. https://doi.org/10.1016/j.tig.2017.06.005
Peric-Hupkes D, Meuleman W, Pagie L et al (2010) Molecular maps of the reorganization of genome-nuclear lamina interactions during differentiation. Mol Cell 38:603–613. https://doi.org/10.1016/j.molcel.2010.03.016
Poudyal RR, Pir Cakmak F, Keating CD, Bevilacqua PC (2018) Physical principles and extant biology reveal roles for RNA-containing membraneless compartments in origins of life chemistry. Biochemistry 57:2509–2519. https://doi.org/10.1021/acs.biochem.8b00081
Quinodoz SA, Ollikainen N, Tabak B et al (2018) Higher-order inter-chromosomal hubs shape 3D genome organization in the nucleus. Cell 174:744–757.e24. https://doi.org/10.1016/j.cell.2018.05.024
Ramani V, Deng X, Qiu R et al (2017) Massively multiplex single-cell Hi-C. Nat Methods 14:263–266. https://doi.org/10.1038/nmeth.4155
Rao SSP, Huntley MH, Durand NC et al (2014) A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159:1665–1680. https://doi.org/10.1016/j.cell.2014.11.021
Reddy S, Comai L (2012) Lamin A, farnesylation and aging. Exp Cell Res 318:1–7. https://doi.org/10.1016/j.yexcr.2011.08.009
Reddy KL, Zullo JM, Bertolino E, Singh H (2008) Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature 452:243–247. https://doi.org/10.1038/nature06727
Ricci MA, Manzo C, García-Parajo MF et al (2015) Chromatin fibers are formed by heterogeneous groups of nucleosomes in vivo. Cell 160:1145–1158. https://doi.org/10.1016/j.cell.2015.01.054
Robson MI, de las Heras JI, Czapiewski R et al (2016) Tissue-specific gene repositioning by muscle nuclear membrane proteins enhances repression of critical developmental genes during myogenesis. Mol Cell 62:834–847. https://doi.org/10.1016/j.molcel.2016.04.035
Rodríguez-Carballo E, Lopez-Delisle L, Zhan Y et al (2017) The HoxD cluster is a dynamic and resilient TAD boundary controlling the segregation of antagonistic regulatory landscapes. Genes Dev 31:2264–2281. https://doi.org/10.1101/gad.307769.117
Ron G, Globerson Y, Moran D, Kaplan T (2017) Promoter-enhancer interactions identified from Hi-C data using probabilistic models and hierarchical topological domains. Nat Commun 8:2237. https://doi.org/10.1038/s41467-017-02386-3
Sabari BR, Dall’agnese A, Boija A et al (2018) Coactivator condensation at super-enhancers links phase separation and gene control. Science 361:6400. https://doi.org/10.1126/science.aar3958
Sanborn AL, Rao SSP, Huang S-C et al (2015) Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc Natl Acad Sci USA 112:E6456–E6465. https://doi.org/10.1073/pnas.1518552112
Scaffidi P, Misteli T (2006) Lamin A-dependent nuclear defects in human aging. Science 312:1059–1063. https://doi.org/10.1126/science.1127168
Schoenfelder S, Furlan-Magaril M, Mifsud B et al (2015) The pluripotent regulatory circuitry connecting promoters to their long-range interacting elements. Genome Res 25:582–597. https://doi.org/10.1101/gr.185272.114
Serebryannyy L, Misteli T (2018) Protein sequestration at the nuclear periphery as a potential regulatory mechanism in premature aging. J Cell Biol 217:21–38. https://doi.org/10.1083/jcb.201706061
Shachar S, Misteli T (2017) Causes and consequences of nuclear gene positioning. J Cell Sci 130:1501–1508. https://doi.org/10.1242/jcs.199786
Shachar S, Voss TC, Pegoraro G et al (2015) Identification of gene positioning factors using high-throughput imaging mapping. Cell 162:911–923. https://doi.org/10.1016/j.cell.2015.07.035
Shalem O, Sanjana NE, Hartenian E et al (2014) Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343:84–87. https://doi.org/10.1126/science.1247005
Shin Y, Brangwynne CP (2017) Liquid phase condensation in cell physiology and disease. Science 357:6357. https://doi.org/10.1126/science.aaf4382
Simon DN, Wilson KL (2013) Partners and post-translational modifications of nuclear lamins. Chromosoma 122:13–31. https://doi.org/10.1007/s00412-013-0399-8
Smith EM, Lajoie BR, Jain G, Dekker J (2016) Invariant TAD boundaries constrain cell-type-specific looping interactions between promoters and distal elements around the CFTR Locus. Am J Hum Genet 98:185–201. https://doi.org/10.1016/j.ajhg.2015.12.002
Solovei I, Wang AS, Thanisch K et al (2013) LBR and lamin A/C sequentially tether peripheral heterochromatin and inversely regulate differentiation. Cell 152:584–598. https://doi.org/10.1016/j.cell.2013.01.009
Stanton BZ, Chory EJ, Crabtree GR (2018) Chemically induced proximity in biology and medicine. Science 359:6380. https://doi.org/10.1126/science.aao5902
Steglich B, Sazer S, Ekwall K (2013) Transcriptional regulation at the yeast nuclear envelope. Nucleus 4:379–389. https://doi.org/10.4161/nucl.26394
Stevens TJ, Lando D, Basu S et al (2017) 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 544:59–68. https://doi.org/10.1038/nature21429
Stewart CL, Kozlov S, Fong LG, Young SG (2007) Mouse models of the laminopathies. Exp Cell Res 313:2144–2156. https://doi.org/10.1016/j.yexcr.2007.03.026
Strickfaden H, Zunhammer A, van Koningsbruggen S et al (2010) 4D Chromatin dynamics in cycling cells: theodor Boveri’s hypotheses revisited. Nucleus 1:284–297. https://doi.org/10.4161/nucl.1.3.11969
Strom AR, Emelyanov AV, Mir M et al (2017) Phase separation drives heterochromatin domain formation. Nature 547:241–245. https://doi.org/10.1038/nature22989
Swift J, Ivanovska IL, Buxboim A et al (2013) Nuclear lamin-A scales with tissue stiffness and enhances matrix-directed differentiation. Science 341:6149. https://doi.org/10.1126/science.1240104
Taberlay PC, Achinger-Kawecka J, Lun ATL et al (2016) Three-dimensional disorganization of the cancer genome occurs coincident with long-range genetic and epigenetic alterations. Genome Res 26:719–731. https://doi.org/10.1101/gr.201517.115
Taimen P, Pfleghaar K, Shimi T et al (2009) A progeria mutation reveals functions for lamin A in nuclear assembly, architecture, and chromosome organization. Proc Natl Acad Sci USA 106:20788–20793. https://doi.org/10.1073/pnas.0911895106
Takizawa T, Meaburn KJ, Misteli T (2008) The meaning of gene positioning. Cell 135:9–13. https://doi.org/10.1016/j.cell.2008.09.026
Tan J, Martin SE (2016) Validation of synthetic CRISPR reagents as a tool for arrayed functional genomic screening. PLoS One 11:e0168968. https://doi.org/10.1371/journal.pone.0168968
Tang Z, Luo OJ, Li X et al (2015) CTCF-mediated human 3D genome architecture reveals chromatin topology for transcription. Cell 163:1611–1627. https://doi.org/10.1016/j.cell.2015.11.024
Tekirdag K, Cuervo AM (2018) Chaperone-mediated autophagy and endosomal microautophagy: Joint by a chaperone. J Biol Chem 293:5414–5424. https://doi.org/10.1074/jbc.R117.818237
Thaller DJ, Patrick Lusk C (2018) Fantastic nuclear envelope herniations and where to find them. Biochem Soc Trans 46:877–889. https://doi.org/10.1042/BST20170442
Therizols P, Illingworth RS, Courilleau C et al (2014) Chromatin decondensation is sufficient to alter nuclear organization in embryonic stem cells. Science 346:1238–1242. https://doi.org/10.1126/science.1259587
Thomson I, Gilchrist S, Bickmore WA, Chubb JR (2004) The radial positioning of chromatin is not inherited through mitosis but is established de novo in early G1. Curr Biol 14:166–172. https://doi.org/10.1016/j.cub.2003.12.024
Timp W, Feinberg AP (2013) Cancer as a dysregulated epigenome allowing cellular growth advantage at the expense of the host. Nat Rev Cancer 13:497–510. https://doi.org/10.1038/nrc3486
Torre LA, Bray F, Siegel RL et al (2015) Global cancer statistics, 2012. CA Cancer J Clin 65:87–108. https://doi.org/10.3322/caac.21262
Towbin BD, González-Aguilera C, Sack R et al (2012) Step-wise methylation of histone H3K9 positions heterochromatin at the nuclear periphery. Cell 150:934–947. https://doi.org/10.1016/j.cell.2012.06.051
Uhler C, Shivashankar GV (2017) Regulation of genome organization and gene expression by nuclear mechanotransduction. Nat Rev Mol Cell Biol 18:717–727. https://doi.org/10.1038/nrm.2017.101
Ulianov SV, Khrameeva EE, Gavrilov AA et al (2016) Active chromatin and transcription play a key role in chromosome partitioning into topologically associating domains. Genome Res 26:70–84. https://doi.org/10.1101/gr.196006.115
Valton AL, Dekker J (2016) TAD disruption as oncogenic driver. Curr Opin Genet Dev 36:34–40. https://doi.org/10.1016/j.gde.2016.03.008
van Steensel B, Belmont AS (2017) Lamina-associated domains: links with chromosome architecture, heterochromatin, and gene repression. Cell 169:780–791. https://doi.org/10.1016/j.cell.2017.04.022
Van Treeck B, Parker R (2018) Emerging roles for intermolecular RNA–RNA interactions in RNP assemblies. Cell 174:791–802. https://doi.org/10.1016/j.cell.2018.07.023
Vazquez J, Belmont AS, Sedat JW (2001) Multiple regimes of constrained chromosome motion are regulated in the interphase Drosophila nucleus. Curr Biol 11:1227–1239. https://doi.org/10.1016/S0960-9822(01)00390-6
Vian L, Pękowska A, Rao SSP et al (2018) The energetics and physiological impact of cohesin extrusion. Cell 173:1165–1178.e20. https://doi.org/10.1016/j.cell.2018.03.072
Vidak S, Foisner R (2016) Molecular insights into the premature aging disease progeria. Histochem Cell Biol 145:401–417. https://doi.org/10.1007/s00418-016-1411-1
Walczak A, Szczepankiewicz AA, Ruszczycki B et al (2013) Novel higher-order epigenetic regulation of the Bdnf gene upon seizures. J Neurosci 33:2507–2511. https://doi.org/10.1523/JNEUROSCI.1085-12.2013
Walter J, Schermelleh L, Cremer M et al (2003) Chromosome order in HeLa cells changes during mitosis and early G1, but is stably maintained during subsequent interphase stages. J Cell Biol 160:685–697. https://doi.org/10.1083/jcb.200211103
Wang T, Wei JJ, Sabatini DM, Lander ES (2014) Genetic screens in human cells using the CRISPR-Cas9 system. Science 343:80–84. https://doi.org/10.1126/science.1246981
Wang J, Choi JM, Holehouse AS et al (2018) A molecular grammar governing the driving forces for phase separation of prion-like RNA binding proteins. Cell 174:688–699.e16. https://doi.org/10.1016/j.cell.2018.06.006
Welch HG, Black WC (2010) Overdiagnosis in cancer. J Natl Cancer Inst 102:605–613. https://doi.org/10.1093/jnci/djq099
Whitton H, Singh LN, Patrick MA et al (2018) Changes at the nuclear lamina alter binding of pioneer factor Foxa2 in aged liver. Aging Cell 17:e12742. https://doi.org/10.1111/acel.12742
Wiech T, Stein S, Lachenmaier V et al (2009) Spatial allelic imbalance of BCL2 genes and chromosome 18 territories in nonneoplastic and neoplastic cervical squamous epithelium. Eur Biophys J 38:793–806. https://doi.org/10.1007/s00249-009-0474-5
Wutz G, Varnai C, Nagasaka K et al (2017) Topologically associating domains and chromatin loops depend on cohesin and are regulated by CTCF, WAPL, and PDS5 proteins. EMBO J 36:3573–3599. https://doi.org/10.15252/embj.201798004
Yamanaka M, Smith NI, Fujita K (2014) Introduction to super-resolution microscopy. Microscopy 63:177–192. https://doi.org/10.1093/jmicro/dfu007
Yamazaki T, Souquere S, Chujo T et al (2018) Functional domains of NEAT1 architectural lncRNA induce paraspeckle assembly through phase separation. Mol Cell 70:1038–1053.e7. https://doi.org/10.1016/j.molcel.2018.05.019
Zink D, Amaral MD, Englmann A et al (2004a) Transcription-dependent spatial arrangements of CFTR and adjacent genes in human cell nuclei. J Cell Biol 166:815–825. https://doi.org/10.1083/jcb.200404107
Zink D, Fischer AH, Nickerson JA (2004b) Nuclear structure in cancer cells. Nat Rev Cancer 4:677–687. https://doi.org/10.1038/nrc1430
Zullo JM, Demarco IA, Piqué-Regi R et al (2012) DNA sequence-dependent compartmentalization and silencing of chromatin at the nuclear lamina. Cell 149:1474–1487. https://doi.org/10.1016/j.cell.2012.04.035
Acknowledgements
Work in the Misteli lab was supported by funding from the Intramural Research Program of the National Institutes of Health (NIH), National Cancer Institute, and Center for Cancer Research and by the NIH 4D Nucleome Common Fund Project (5U54DK107980-01). PT was supported by the Polish National Science Centre Grants (2014/15/N/NZ2/00379 and 2017/24/T/NZ2/00307), KM was supported by a Department of Defense Idea Award (W81XWH-15-1-0322), CA was supported by an IWT fellowship (no. 141372) from the Belgian Research Foundation—Flanders (FWO) and a Gustave Boël—Sofina travel grant from the Koning Boudewijn Stichting, Belgium, SV was supported by an Erwin Schroedinger Fellowship from the Austrian Science Fund (J3849-B28), and MF is supported by a Postdoctoral Research Associate Training (PRAT) fellowship from the National Institute of General Medical Sciences (NIGMS, 1Fi2GM128585-01).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Adriaens, C., Serebryannyy, L.A., Feric, M. et al. Blank spots on the map: some current questions on nuclear organization and genome architecture. Histochem Cell Biol 150, 579–592 (2018). https://doi.org/10.1007/s00418-018-1726-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-018-1726-1