Abstract
The Mycobacterium tuberculosis (M. tuberculosis)-specific culture filtrate protein-10 (CFP-10) is highly recognized by M. tuberculosis infected subjects. In the present study, the proliferative response and IFN-γ secretion was found for C-terminal peptides of the protein (Cfp651–70, Cfp761–80, Cfp871–90, and Cfp981–100). The alleles HLA DRB1 *04 and HLA DRB1 *10 recognized the C-terminal peptides Cfp7, Cfp8, and Cfp9 in HHC. Cfp6 was predominantly recognized by the alleles HLA DRB1 *03 and HLA DRB1 *15 by PTB. The minimal nonameric epitopes from the C-terminal region were CFP-1056–64 and CFP-1076–84. These two peptides deserve attention for inclusion in a vaccine against tuberculosis in this region.
Similar content being viewed by others
Introduction
The scourge of diseases has afflicted the world since time immemorial. Tuberculosis (TB) remains one among such diseases, which has caused widespread morbidity and mortality worldwide [1]. It proves to be a dangerous liaison in the context of opportunistic infection in human immunodeficiency virus (HIV) infected individuals. The disease becomes an even more challenging one when it becomes resistant to TB drugs—multidrug resistant tuberculosis (MDR) and extremely drug resistant tuberculosis (XDR) [2].
Two secreted proteins of Mycobacterium tuberculosis (M. tuberculosis) which are of much interest because of their role in protection are region of deletion-1 (RD-1) locus encoded early secreted antigenic target-6 (ESAT-6) and culture filtrate protein-10 (CFP-10). CFP-10 is a low molecular weight culture filtrate protein, which is cotranscribed along with ESAT-6 gene, its heterodimeric counterpart. RD-1 region is present in pathogenic M. tuberculosis genome and in Mycobacterium bovis, but absent in non-pathogenic Bacille Calmette Guerin (BCG) strains [3–7]. Specific immunity against these proteins is known to enhance host resistance against M. tuberculosis infection [8]. For successful containment of tuberculosis infection, a strong Th1 response is required [9]. Both ESAT-6 and CFP-10 are known to stimulate T cells to produce interferon gamma (IFN-γ), a Th1 cytokine, and exhibit cytolytic T lymphocyte (CTL) activity in animals and in humans, making them excellent candidates for including in a tuberculosis vaccine [10–13].
In this study, we have identified peptides of CFP-10, which induce IFN-γ production and CD4, CD8 T cell proliferation. Of nine peptides, the C-terminal Cfp7 (Cfp-10 peptide 7), Cfp8, and Cfp9 peptides induced IFN-γ production and proliferation of CD4 and CD8 cells in HHC.
Materials and methods
Study participants
This study was approved by the Institutional Ethics Committee. Informed consent was obtained from the subjects before drawing blood.
Healthy household contacts of TB patients (n = 23)
Healthy household contacts (HHC) were recruited from families where there was at least one sputum positive PTB (index case) living in the same household, sharing the kitchen and bathroom, for at least 3 months immediately preceding the start of treatment of the index case, who were sure to be infected [14], whose age ranged from 27 to 54 years. The male/female ratio was 13:10. During recruitment, the subjects were clinically evaluated for symptoms of tuberculosis and chest radiographs were taken. None of the HHC presented with clinical symptoms or lesions in chest X-rays excluding the possibility of TB disease. Sputum smears and cultures were also negative in this group. Quantiferon-TB gold (in-tube) test was done to confirm the M. tuberculosis infection in the subjects. The subjects were followed for a period of 6 months and none broke down with tuberculosis disease.
Tuberculosis patients (n = 34)
Pulmonary tuberculosis patients before treatment (n = 21)
All pulmonary tuberculosis patients before treatment (PTB) were positive for sputum smears and culture. The subjects of this group were naïve for anti-tuberculous treatment. Their age ranged from 28 to 52 years and the male/female ratio was 17:4. Examination of smear and culture was done according to the methods already established in our centre [15]. Two spot and 1 overnight sputum specimens were collected from each patient. For smears, acid-fast bacilli (AFB) staining was done and they were examined under fluorescence microscope. For culture, sputum was concentrated and inoculated onto Lowenstein–Jensen (LJ) media and incubated for up to 8 weeks at 37°C and checked for positivity.
Treated tuberculosis patients (n = 13)
The subjects in this category had completed 6 months course of anti-tuberculous therapy (ATT), and there was no evidence for recent infection. The age range was 26–50 years, and the male/female ratio was 10:3. Chest X-ray and sputum smears were done to exclude active TB in this group. The treated tuberculosis patients (TR) were recruited after a period range of 7–36 months of start of their treatment. The drug regimen followed for these patients was 2EHRZ3/4RH3.
The patients and contacts recruited for the study were seronegative for HIV by two enzyme immunoassays—Tridot (J. Mitra & Co., New Delhi, India) and Retroquic (Qualprodiagnostics, Goa, India) in serum.
PPD (purified protein derivative) skin testing was not done in our study subjects, as it is neither sensitive nor specific. It has been observed that >70% of women and >80% of men turn positive for PPD by 25 years, due to exposure to M. tuberculosis as well as other environmental mycobacteria. Thus, PPD positivity in our population can mean infection not only with M. tuberculosis but also with other environmental mycobacteria. The more sensitive Quantiferon test uses antigens unique for M. tuberculosis; the test identifies infected subjects and BCG vaccination or exposure to environmental mycobacteria does not affect the test results. This test was done after subject recruitment and was not used for preselecting positive subjects. All the HHC recruited for the study were positive for Quantiferon-TB Gold test (≥0.35 IU/ml).
Quantiferon-TB Gold In-tube assay
Infection status of tuberculosis was assessed by an ELISA based Quantiferon-TB Gold kit (Cellestis Inc., Victoria, Australia) as per the manufacturer’s instructions. Blood was collected in tubes labeled nil control, mitogen, and TB antigen (ESAT-6 and CFP-10 overlapping peptides and a TB 10.4 peptide). The tubes were then incubated at 37°C for 16–24 h, and then plasma was collected from the tubes. The cut-off point was set as 0.35 IU/ml as per the manufacturer’s instructions.
Recombinant proteins and synthetic peptides
Recombinant protein CFP-10 was prepared in the laboratory of Dr. Pawan Sharma, New Delhi. The protein was prepared as already described [16]. CFP-10 peptides (a kind gift from Dr. Thomas B Nutman, NIAID, NIH, USA) covered the entire primary structure of the protein (20-mers with 10-amino acid overlap). Protein estimation was done for the peptides using BCA protein assay kit (Pierce Biotechnology, Rockford, IL, USA). Lyophilized peptides were reconstituted in dimethyl sulfoxide, aliquoted, and stored at −80°C until use.
The peptide sequences used in the present study were as follows:
CFP-10 (1–20) Cfp1 | MAEMKTDAATLAQEAGNFER |
CFP-10 (11–30) Cfp2 | LAQEAGNFERISGDLKTQID |
CFP-10 (21–40) Cfp3 | ISGDLKTQIDQVESTAGSLQ |
CFP-10 (31–50) Cfp4 | QVESTAGSLQGQWRGAAGTA |
CFP-10 (41–60) Cfp5 | GQWRGAAGTAAQAAVVRFQE |
CFP-10 (51–70) Cfp6 | AQAAVVRFQEAANKQKQELD |
CFP-10 (61–80) Cfp7 | AANKQKQELDEISTNIRQAG |
CFP-10 (71–90) Cfp8 | EISTNIRQAGVQYSRADEEQ |
CFP-10 (81–100) Cfp9 | VQYSRADEEQQQALSSQMGF |
The peptides used in in vitro studies were not identified previously by in silico analysis. All the overlapping peptides covering the entire sequence of the CFP-10 protein were tested in the in vitro experiments and peptides were not pooled for the experiments.
Concentrations of peptides
The overlapping peptides of CFP-10 were used at different concentrations. A standardization experiment with a range of concentrations (1, 2, 5, and 10 μg/ml) of peptides was done and based on lymphocyte proliferation and IFN-γ secretion (from culture supernatants); the concentration which gave the maximal response was fixed and used.
IFN γ ELISA
Five-day culture supernatants from BrdU culture peripheral blood mononuclear cells (PBMCs) were assayed for the presence of IFN γ by ELISA as per the manufacturer’s instructions (BD Biosciences, San Diego, CA). The detection limit of the assay ranged from 4.7 to 300 pg/ml. The lowest detection limit of the kit was 1 pg/ml.
Intracellular cytokine staining
Whole blood was diluted in the ratio of 1:2 with Roswell Park Memorial Institute medium (RPMI; Sigma–Aldrich corporation, St. Louis, MO, USA) to which costimulatory molecules (CD49d and CD28 1 μg/ml each/ml of culture), overlapping peptides and recombinant proteins of CFP-10 were added at optimal concentrations and incubated in 48-well plates for 18 h in the presence of brefeldin A (BD Biosciences, San Diego, CA, USA) during the last 16 h (Costar, Corning Inc., NY, USA) at 37°C in 5% CO2 atmosphere. After stimulation, cells were washed and labeled with fluorochrome-conjugated specific antibodies anti-CD4 APC, CD8 PE-Cy5, IFN-γ PE, and CD4 APC, CD8 PE-Cy5, IL-4 PE in separate tubes (BD Biosciences, San Diego, CA, USA). Fixing and permeabilising of cells were done using Cytofix/Cytoperm buffer (BD Biosciences, San Diego, CA, USA), washed with Perm/wash buffer (BD Biosciences, San Diego, CA, USA), and then stained for cytokines. After washing, cells were fixed using 4% paraformaldehyde and analyzed using a FACSCalibur Flow cytometer (BD Biosciences, San Diego, CA, USA). Percentage of cytokine secreting CD4 and CD8 cells were analyzed using FlowJo software (Tree Star Inc., San Carlos, CA, version 7.1.1).
BrdU (bromodeoxyuridine) incorporation assay
PBMCs were separated from venous blood using Histopaque (Sigma–Aldrich Corporation, St. Louis, MO, USA). To the separated cells [1 × 106/ml in RPMI + 10% heat inactivated human AB serum (Sigma–Aldrich corporation, St. Louis, MO, USA)], CFP-10 peptides, recombinant CFP-10 protein, and M. tuberculosis culture filtrate antigen were added. The plate (Costar, Corning Inc., NY, USA) was incubated at 37°C in a 5% CO2 atmosphere for 5 days. BrdU incorporation assay was carried out using a BrdU flow kit (BD Biosciences, San Diego, CA) as per the manufacturer’s instructions.
Briefly, BrdU (50 μM/ml) (BD Biosciences, San Diego, CA, USA) was added 16 h prior to termination of the experiment. Cells were removed from the plate using 2 mM EDTA, surface labels (CD4 APC, CD8 PE-Cy5; BD Biosciences, San Diego, CA, USA) were added and incubated for 30 min. The cells were then permeabilized using Cytofix/Cytoperm buffer (BD Biosciences, San Diego, CA, USA) for 30 min. Repermeabilization was carried out by incubation with cytoperm plus buffer for 15 min. To expose the incorporated BrdU, the cells were treated with 100 μg of DNAse/0.5 × 106 PBMCs per tube (BD Biosciences, San Diego, CA, USA) and incubated at 37°C for an hour. This was followed by addition of anti-BrdU FITC antibody (BD Biosciences, San Diego, CA, USA) and incubation for 30 min. Finally, the cells were fixed in 4% paraformaldehyde and then analyzed on a FACSCalibur Flow cytometer (BD Biosciences, San Diego, CA, USA). Data were collected and analyzed using FlowJo software (Tree Star Inc., San Carlos, CA, version 7.1.1).
MHC typing
DNA extraction was done from polymorphonuclear cells, as well as from PBMCs by a salting out method as described elsewhere [17]. Typing of DRB was done by PCR with Sequence-specific primers (SSP). Low resolution PCR-SSP was performed for DRB1 alleles [18].
In silico prediction of potential binding regions in CFP-10
MHC binding peptides (15-mers) were predicted from CFP-10 using MHC-II binding prediction server [19] available at http://tools.immuneepitope.org/main/jsp/menu.jsp. For prediction, consensus method was chosen since the method has been reported to be better than other methods [19]. The potential nonameric epitopes were predicted using ProPred available at http://www.imtech.res.in/raghava/propred/ [20]. Default threshold was set at which the sensitivity and specificity were found to be similar. HLA were typed in our study only by low resolution (two digits) while in the bioinformatics server we used, higher resolution for HLA were available. In both the methods, HLA were restricted by HLA subtypes observed in our study population. In vivo, the protein sequence can be processed and cleaved in any position to form the epitopes. Therefore, to predict the possible HLA binders present in the whole sequence, the full-length sequence of the CFP-10 was used. Therefore, nonamers predicted to be present in the overlapping sequence have been represented in two consequent 20-mers in the in silico analysis (Table 6).
Statistical analysis
GraphPad Prism (Graph PAD Software 4.0, San Diego, USA) was used for result analysis. The differences between IFN-γ secretion (ELISA) and intracellular IFN-γ levels among three groups and proliferation among the groups were assessed by one-way analysis of variance (ANOVA) with Bonferroni’s correction for comparing multiple columns. For calculating percent responders among various groups, mean + 3SD of unstimulated culture of subjects in all groups was taken which was ~100 pg/ml. This was considered the common cut-off for all the groups. Those values which were above this cut-off point were considered as positive [21, 22]. Overall IFN-γ response among various groups was calculated by subtracting the unstimulated mean values from stimulated culture mean values. For BrdU incorporation assays, the cut-off was set by considering mean + 3SD of unstimulated culture of each group. The values above this cut-off were considered positive. Intracellular IFN-γ and IL-4 response in PTB and HHC groups was analyzed by Student’s t-test.
Results
Recognition of CFP-10 peptides
The response to CFP-10 peptides was measured by an IFN-γ ELISA. To calculate positivity, mean + 3SD of unstimulated culture value for all subjects of a group was considered. This was ~100 pg/ml for all the groups. So, this was used as the common cut-off for all the groups. The values above this cut-off point were deemed positive [21, 22].
For M. tuberculosis culture filtrate antigen (CFA), responders were 17/21 (71%) among PTB group. CFP-10 protein response of PTB was 48% (13/21). Individual peptide responses were as follows: Out of 13 CFP-10 responders, 5 responded to Cfp6 giving 39% response. For Cfp5, Cfp7, and Cfp8, 23% (3/13) response was observed. Other peptide responders were 15% (2/13) for rest of the peptides (Table 1).
Among HHC, the CFA responders were 91% (21/23). CFP-10 responders among HHC were 65% (15/23). Individual peptides gave the following results: for Cfp6 and Cfp8 peptides 53% of HHC reacted (8/15). The peptide Cfp7 and Cfp9 gave 47% (7/15) response. The Cfp2 and Cfp3 peptides were recognized by 40% (6/15) of HHC. It was observed that responders to Cfp1 and Cfp5 were 33% (5/15). The responders for Cfp 4 were 27% (4/15) (Table 1).
A 100% (13/13) response was found for CFA among TR. Eighty-five percent of TR reacted to CFP-10 protein (11/13). Of 11 CFP-10 positive subjects, 7 TR were positive for Cfp6 accounting for 64% responders. The responders for peptides Cfp1 and Cfp8 were 55% (6/11). A response of 46% (5/11) was observed for peptides Cfp2 and Cfp5. For rest of the peptides, the responses were 36% (4/11) (Cfp3, Cfp4, Cfp7, and Cfp9; Table 1).
The peptides Cfp6, Cfp7, Cfp8, and Cfp5 were the peptides predominantly recognized by PTB. The HHC recognized the peptides Cfp6, Cfp8, Cfp7, Cfp9 and Cfp2, Cfp3. TR reacted to the peptides Cfp6, Cfp1, Cfp8, Cfp2, and Cfp5.
The levels of IFN-γ were studied in various groups. Cfp7 induced highest mean IFN-γ response (4,292 pg/ml) in HHC. The peptides Cfp8, Cfp9, Cfp6 induced 3,910, 2,672, 2,617 pg/ml of IFN-γ by T cells, respectively.
The peptide Cfp6 induced highest level of IFN-γ (279 pg/ml) in PTB. For Cfp8, the IFN-γ induced was 216 pg/ml. In case of TR, an increased response was observed for peptide Cfp8 (335 pg/ml). Other peptides for which high response was observed were Cfp3 (279 pg/ml), Cfp6 (278 pg/ml), Cfp7 (260 pg/ml), Cfp5 (247 pg/ml), and Cfp1 (221 pg/ml).
It was observed that Cfp6, Cfp7, Cfp8, and Cfp9 were the peptides for which predominantly higher IFN-γ was produced by T lymphocytes in HHC when compared to PTB. In comparison with PTB, the TR group responded to Cfp8 (Fig. 1). The differences in IFN-γ levels between the groups were not statistically significant.
IFN-γ response to overlapping peptides of CFP-10. The IFN-γ levels in each group were assessed for all CFP-10 overlapping peptides by considering mean values (pg/ml) for all the groups. For all the groups, the unstimulated mean values were subtracted from stimulated mean values. The difference between groups was studied by one-way ANOVA with Bonferroni’s posttest. P pulmonary tuberculosis patients, C healthy household contacts, T treated TB patients, CFP-10 culture filtrate protein-10
Proliferative responses to overlapping peptides of CFP-10
Representative flow cytometry plots for proliferative response to a peptide (Cfp8) are given in Fig. 2a, b, c, d. The proliferative response to overlapping CFP-10 peptides was studied by a flow cytometry based BrdU incorporation assay. The responses were studied in HHC, PTB, and TR in both CD4 as well as CD8 cells. The response of each individual to a CFP-10 peptide was studied by taking mean + 3SD of unstimulated culture. Those values which were above this cut-off point were considered positive. Of 23 HHC, 19 responded to CFP-10 protein (83%) for CD4+ T cells. It was observed that among HHC, the peptides for which response was observed were Cfp9 (11/23–48%), Cfp7 (10/23–44%), Cfp6 (10/23–44%), Cfp8 (9/23–39%; Table 2), for rest of the peptides the response was less (8–30%). In PTB group, the response was obtained for peptides Cfp6 (48%), Cfp5 (8/21–38%) and Cfp4 (38%). The remaining peptide response ranged from 5 to 29%. The response in TR for CFP-10 protein was 100%. The responding peptides in TR group were Cfp5 (92%), Cfp7 (85%), Cfp3 (77%), Cfp8 (62%), Cfp2 (62%), Cfp1 (62%). The response range for rest of the peptides was 23–40% (data not shown).
CD4 (a, b) and CD8 (c, d) cell proliferation response in HHC by BrdU incorporation assay. a Unstimulated culture—CD4. b Peptide Cfp8—CD4. c Unstimulated culture—CD8. d Peptide Cfp8—CD8. For flow cytometric analysis of proliferating cell population, lymphocyte population was gated in forward scatter/side scatter plot and within the gate; BrdU-positive CD4 and CD8 cells were calculated; HHC healthy household contacts
The peptides for which an increased percentage of proliferating CD4+ cells were found: Cfp7, Cfp9, Cfp6, and Cfp8 in HHC; Cfp4, Cfp5, and Cfp6 in PTB; and Cfp5, Cfp3, Cfp7, Cfp8, Cfp2, and Cfp1 in TR.
The proliferative responses in three groups for CD8 cells gave the following results: The increased percentage of CD8 cells positive for BrdU was observed for peptides Cfp6 (48%), Cfp8 (44%), Cfp1, Cfp5 and Cfp7 (39%), Cfp3 (30%) in HHC group in comparison with other peptides (Table 3). The peptides Cfp8 (38%), Cfp5, Cfp6, and Cfp9 (24%) responded in PTB group for which increased number of proliferating CD8 cells were observed. In TR group, the responding peptides were Cfp5 (85%), Cfp7 and Cfp9 (69%), Cfp6 and Cfp8 (62%; data not shown).
The peptides Cfp6, Cfp8, Cfp1, Cfp3, Cfp5, and Cfp7 were the ones to which CD8+ cell proliferation was observed in HHC. In PTB, Cfp8, Cfp5, Cfp6, and Cfp9 were the peptides for which response was observed, and the response was found for peptides Cfp5, Cfp7, Cfp9, Cfp6, and Cfp8 in TR.
Th1 and Th2 responses to peptides (IFN-γ and IL-4)
Since Th2 immunity also plays a role in mycobacterial immunity, the signature cytokine of Th2 cells (IL-4) was also studied. The ratio of IFN-γ to IL-4 was studied by intracellular cytokine staining of CD4 cells. For CD4 cells, the ratio ranged from 1.3 to 1.9 for the peptides Cfp6, Cfp7, Cfp8, and Cfp9 in HHC when compared to other peptides for which it was low (0.6–1.2). In the case of PTB, the ratio ranged from 0.4 to 1.5 for all the peptides. Peptide Cfp1 (1.1) and Cfp5 (1.5) showed an elevated ratio when compared to other peptides. Although there was an increased response among HHC when compared to PTB, the difference was not statistically significant (Table 4). The mean IFN-γ and IL-4 for both HHC and PTB studied within CD4+ populations have also been represented in Table 4. When IFN-γ/IL-4 ratio was studied for CD8 cells, no differences in ratio between peptides were observed (data not shown).
MHC class II allele (HLA-DRB1) typing
HLA DRB1 gene typing gave the following results:
High IFN-γ response was observed for the peptide Cfp7 (aa61–80), which was recognized by the alleles HLA DRB1 *04 and *10 among HHC. Other peptides for which these alleles gave an elevated response were Cfp8 (aa71–90) and Cfp9 (aa81–100). The potential minimal epitopic regions may be lying within these highly responding C-terminal peptides. The region Cfp6 (aa51–70) was recognized (30,364.06 pg/ml) by the allele HLA DRB1 *14 by an HHC (Table 5). Other alleles for which a positive IFN-γ response was not observed for CFP-10 protein, and its peptides were HLA DRB1 *07, *11, *12, *03.
The allele HLA DRB1 *03 and DRB1 *15 of PTB recognized the region Cfp6 (Table 5). For other alleles of PTB subjects (HLA DRB1 *07, *13, *14), no positive response was observed for CFP-10 and its peptides.
Among TR, the peptides Cfp3 (aa21–40) and Cfp5 (aa41–60) were the ones to be recognized by HLA DRB1 *14, *10, *04, *12 alleles. Although response was observed for other peptides, Cfp6 (HLA DRB1 *14, HLA DRB1 *15), Cfp7 (HLA DRB1 *10, HLA DRB1 *04), Cfp8 (HLA DRB1 *14, HLA DRB1 *10, HLA DRB1 *12), and Cfp9 (HLA DRB1 *10, HLA DRB1 *04), few of the four above-mentioned alleles were recognized. HLA DRB1 alleles of subjects for whom a positive IFN-γ response was not observed was HLA DRB1 *11 for CFP-10 protein.
In silico analysis of CFP-10
Thirteen 15-mers were predicted to be starting in three regions of CFP-10 sequence as follows: (a) at amino acid position 1, (b) between 37 and 39, at 42 and 43 and (c) at 67 and between 70 and 75. The IEDB server calculates the affinity to all possible 15-mers for the given protein sequence. Lower the Consensus Percentile Rank, better the affinity of the peptide with the MHC. Only the topmost ranked 15-mers alone were shown in the results which were potential HLA binders. On this basis, thirteen 15-mers were predicted to be potential HLA binders with nine different HLA subtypes (HLA_DRB1-0301, HLA_DRB1-0401, HLA_DRB1-0404, HLA_DRB1-0405, HLA_DRB1-0701, HLA_DRB1-0802, HLA_DRB1-1101, HLA_DRB1-1302, and HLA_DRB1-1501), which were also observed in the study population (Table 6).
Eight nonamers were found to be predicted as potential epitopes recognized by HLA-DRB1 subtypes. Among them, epitopes CFP-1056–64 and CFP-1076–84 were predicted as potential promiscuous epitopes as the former was predicted as potential binder with 32 different HLA-DRB1 alleles while the latter was predicted to be recognized by 41 alleles. Remaining epitopes were predicted to be binders to HLA ranging from one to eight different HLA-DRB1 subtypes (Table 6).
Discussion
It has been reported that CFP-10 protein is recognized by T cells from active tuberculosis patients and latently infected subjects [13, 23, 24].
Till date, there are no good immunological parameters that correlate with protective immunity against TB in humans. Experimental evidence shows the dependence of antigen-specific T lymphocytes and their ability to stimulate antimycobacterial activity of macrophages by release of IFN-γ. The role of IFN-γ in TB control has been demonstrated in IFN-γ gene-disrupted mice [25], and the mutation of this gene in humans leads to increased susceptibility to tuberculosis [26]. The ability to stimulate T cell release of IFN-γ is a criterion for identifying protective antigens in tuberculosis. Various studies have provided evidence that antigens recognized by “protected group”, but not active TB patients, can be considered for vaccine development strategies using IFN-γ as a protective correlate. This approach is an efficient one for protective antigens identification [27–29]. So, we have used IFN-γ as a correlate for protection in screening the CFP-10 peptide responses.
Our results from this study suggest that 20-mer peptides of the C-terminal region of CFP-10 protein are immunogenic eliciting IFN-γ production from HHC. The peptides recognized by HHC in the present study were Cfp651–70 (53%), Cfp761–80 (47%), Cfp871–90 (53%), and Cfp981–100 (47%). Previous studies on CFP-10 peptides in Indian and Zambian population have shown that the carboxy terminal region is immunogenic and the peptide Cfp871–90 elicits responses by 30–50% of healthy asymptomatic subjects [13, 23]. In concordance with this result, we have found about 53% of HHC recognizing this peptide in our present study. This peptide produced 3,910 pg/ml of IFN-γ (mean value) and ranked next to Cfp7.
A study in Indian population by Lalvani et al. [13] on CFP-10 overlapping peptides (15-mers overlapping by 10 amino acids) has identified two immunodominant regions CFP51–70 and CFP71–90. These regions have been shown to be recognized by healthy subjects. Our study shows Cfp7 (61–80) and Cfp8 (71–90) peptides to produce highest IFN-γ response in HHC. The response observed in our study is a 5-day response, in contrast to 14-h response observed in Lalvani et al. [13] study.
In a study by Shams et al. [30], in American population, a 15-mer peptide, CFP-1071–85 has been identified, which has been shown to elicit IFN-γ production by ELISPOT assay. In our study too, this region showed a high IFN-γ response, but Cfp6 (51–70) and Cfp8 (71–90) of our study exhibited the highest response. This may be due to genetic differences between the populations which led to differential recognition of peptides.
It has been shown by Lewinsohn et al. [31] that CD4+ T cells recognized certain peptidic regions like CFP101–23, CFP1069–100 more, and the peptide CFP1041–59 was recognized to a lesser extent. The study was carried out in American population and CD4 T cell reactivity to overlapping peptides of CFP-10 was ascertained by an IFN-γ ELISPOT assay. The subjects enrolled in the study were normal healthy subjects. In their study, CD4+ T cells were purified by magnetic separation and then they were coincubated with peptide pulsed dendritic cells. In our present study, PBMCs were stimulated with peptides and IFN-γ secretion was measured in the supernatants by an ELISA. The second highly recognized peptide (CFP1069–100) of Lewisohn et al. [31] study overlaps our study peptides Cfp7, Cfp8 and Cfp9.
In a study by Kamath et al. [32], a minimal epitope in CFP-10 (32–39) has been identified to elicit immune response in mice by activating CD4 as well as CD8 cells. A marked immune response was not observed for this peptide in all the three subject groups recruited in our study. This fact suggests that results from animal studies do not always extrapolate to human subjects. The other reason may be the result obtained in our study has explored a small HLA pool, and there may be other populations who might respond to this unique peptide.
Our present study has recruited HHC as a group, because these subjects live in the same household of PTB but do not develop the disease. Hence, this group is considered as “protected.” The immune responses are worth studying in this group.
The immune response was low (IFN-γ production and proliferation) for PTB when compared to HHC in this study. Although there are reports on high level of IFN-γ in PTB, like our study, few studies have shown the higher response in HHC than PTB [33–35]. Many factors are attributed to this decreased response viz., advanced disease stage and hence cell recruitment to site of infection, genetic predisposition, defects in antigen presenting cells, etc.
In the present study, treated subjects have been included as a group to investigate whether drugs alter the immune status or restore the immune system to normalcy. The response observed for TR was decreased, when compared to PTB, as well as HHC. This decrease might be due to the fact that the circulating IFN-γ secreting cells specific for CFP-10 peptides might have decreased due to clearance of the infection [36, 37]. This might be due to the clearance of bacilli due to ATT and hence decreased antigenic stimulus and lowered circulating IFN-γ secreting cells.
In the present study, 20-mers were used for which, a CD4+ mediated response is expected because of the peptide size. The 20-mer peptides we used elicited response (proliferation as well as intracellular IFN-γ and IL-4) in CD8+ T cells also. The reason for their CD8+ cell activation may due to the 20-mer peptides being processed and binding class I molecules in endocytic compartments or at the cell surface after regurgitation of processed peptides. It has been shown by Eberl et al. [38] that a 69-mer synthetic polypeptide (a malarial parasite circumsporozoite antigen) is presented to MHC class I restricted CD8 cells. That study involved an in vivo animal (mouse) model as well as in vitro (cell line) system and has suggested that serum components may allow the processing and loading of exogenous polypeptides (which do not enter cytosolic processing pathway) onto empty cell surface class I MHC molecules for presentation to CD8+ cells.
Human AB serum has been used in our study in the cell culture medium whose components might have played a role in enhancing the presentation of breakdown products of 20-mer peptides to CD8 cells and activating it.
Exogenous peptides are also presented to CD8 cells by other mechanisms:
-
1.
Macropinocytosis of phagocytosis of polypeptide by a particular set of macrophages followed by disruption of pinosome or phagosome.
-
2.
Pinocytosis or endocytosis of antigens bound to proteins for which receptors exist on cell surface followed by lysosomal processing, escape of the proteins into cytosol or regurgitation to the cell surface.
In immune response to antigens, an enhanced Th1 and a lowered Th2 response is considered efficient for protection. In mice as well as humans, a lowered IL-4 response has been observed to be associated with protection [39, 40]. In Balb/c mice with IL-4 gene knockout, absence of IL-4 led to restricted growth of M. tuberculosis indicating the absence of this cytokine and protection [39]. In humans, it has been observed that TB health care workers with pre-existing IL-4 responses had a higher rate of progression to tuberculosis [40]. So, this Th1/Th2 axis response was addressed in the present study.
In this study, IFN-γ/IL-4 ratio was studied for all 20-mer peptides used in the study. The Th2 axis of immune response is usually considered in very few epitope mapping studies or not considered at all. We studied the signature cytokine of Th2 arm of immunity and found that the peptides Cfp6, Cfp7, Cfp8 and Cfp9 all showed a high ratio of IFN-γ/IL-4 in HHC group, suggesting that the immune response elicited by these peptides is predominantly Th1 type. In proliferation and IFN-γ secretion, TR was included but not in later studies due to non-availability of samples.
The proliferative responses showed that the percentage of CD4+ cells increased for peptides Cfp6, Cfp7, Cfp8, and Cfp9 in HHC. Thus, it can be concluded that these peptides are able to elicit proliferation of CD4+ T cells and production of IFN-γ. In the case of PTB, the peptides Cfp4, Cfp5, and Cfp6 were the predominant proliferation inducing peptides. The peptides Cfp1, Cfp2, Cfp3, Cfp5, Cfp7, and Cfp8 were the ones which induced proliferation in the TR group. The increased number of peptide inducing proliferation in the TR may be due to prolonged exposure to antigen and increased number of antigen-specific cells for many immunodominant regions of CFP-10 protein.
The CD8+ cell proliferation responses differed from that of CD4+ responses. The peptides Cfp1, Cfp3, Cfp5, Cfp6, Cfp7, Cfp8 elicited proliferation in HHC. The PTB as well as TR group showed CD8+ response to peptides Cfp5, Cfp6, Cfp8, and Cfp9. TR responded to the peptide Cfp7 also. The difference between CD4 and CD8 responses may be due to the presence of unique CD8 epitopes which might have caused proliferation.
In the present study, the strong responders from HHC1, 2, 3, as well as TR6 have allele similarities but T cells respond to distinct epitopes. The probable reason for the lesser number of peptidic regions recognized by HHC1, 2, 3 (Cfp7, Cfp8, Cfp9) when compared to TR6 (Cfp3,Cfp5, Cfp7, Cfp8, Cfp9) might be due to restriction or influence by HLA-DQ, DP class II alleles or HLA class I alleles (A, B, and C) of these subjects. Apart from this, the inter individual variation in IFN-γ response to even those who share the same HLA might be due to polymorphisms in their IFN-γ gene. Another possible reason for differences in epitope recognition between subjects who share the same allele might be due to the epitope being recognized not being a dominant one. This may lead to differential binding of the epitopic regions to the same alleles. The bioinformatics data analysis supports this possibility, because the potential epitopes predicted were found to lie within the peptides Cfp7, Cfp8 and Cfp9 (CFP-1056–64 and CFP-1076–84). The regions Cfp3 and Cfp5 lack immunodominant CFP-10 epitopes. These reasons might be speculated to be the cause for the observed difference in responses between HHC and TR sharing the same alleles.
An association between HLA and tuberculosis has been reported [41]. Among the MHC class II molecules, HLA-DR molecules are reported to be the main presenters of mycobacterial antigens to T cells [42–45]. In vaccine studies, HLA profiles deserve importance because different HLA molecules present peptides to T cells with variable efficiencies [46]. Thus, a need always arises for designing HLA-based vaccines for different geographic areas and specific ethnic groups. The other class II alleles, HLA-DP and DQ alleles have also been studied in tuberculosis [47] and are known to present antigens. They also play a role in immune effector functions. HLA-DRB1, a more polymorphic allele, has been typed in this study but not the other alleles.
In this study, HLA DRB1 *04 and DRB1 *10 were the alleles which showed a predominant response among HHC presenting C-terminal peptides of CFP-10 protein. Cfp6 peptide was recognized by this group. Two peptides, Cfp3 and Cfp5, were recognized by multiple alleles in the TR group. In south India, the association of HLA DRB1 *10 with prevention and DRB1 *15 allele with predisposition to tuberculosis infection has already been studied [47]. In our study, the allele DRB1 *10 of the protected group recognized the peptide Cfp7, Cfp8 as well as Cfp9. These peptides were least recognized by the PTB group in our study hence are unique and may have implications for vaccine design. Response was not observed for certain HLA DRB1 alleles. Major conclusions cannot be drawn from these HLA studies; so a larger cohort of subjects is needed to confirm the observed results.
In vitro response was observed for 20-mers Cfp6, Cfp7, Cfp8, and Cfp9 in HHC group. Though HLA analysis reveals peptide recognition ranging from the length of 9–21 amino acids, 9-mers and 15-mers are more frequently recognized length as epitopes. In order to predict epitopes with minimal length, two bioinformatics servers were chosen in this study, IEDB server to predict possible 15-mers as epitope and to further narrow down the length (nonamers as minimal epitopes) of the epitope in the CFP-10 protein, ProPred was chosen. The minimal epitopes, CFP-1056–64 (VRFQEAANK) and CFP-1076–84 (IRQAGVQYS) were predicted as potential promiscuous nonamers by ProPred. The prediction of nonamer 76–84 is also supported by the IEDB result in which 15-mers starting from any of amino acids, 70–75 (15-mers 70–84, 71–85, 72–86, 73–87, 74–88, and 75–89) were predicted as potential binders with different HLA subtypes (HLA_DRB1-0301, HLA_DRB1-0404, HLA_DRB1-0405, and HLA_DRB1-0802). The epitope 76–84 was present in Cfp7, 8, 9 and all of those peptides were found to induce immune response in vitro. The nonamer 56–64 was predicted as a potential promiscuous epitope in ProPred analysis while the same nonamer was part of Cfp5, 6, and 7. The immune response has been observed in vitro for Cfp6 and 7 while no response was found for the peptide Cfp5. Therefore, amino acids in the carboxy terminal of the nonamer of 56–64 may be responsible for immune response. The structural biology study of these two nonamers (56–64 and 76–84) and their HLA association may help us to better understand the importance of specific amino acid in the immune response.
Two methods were chosen here for the in silico analysis in order to enhance the quality of the prediction. Nonamers predicted using ProPred start at position 4, 39, 43, 55, 56, and 76 were found to be part of or subsequence of the 15-mers predicted using the Consensus method (IEDB server). Prediction of 15-mers as HLA binders by IEDB which are already demonstrated to be immunogenic in vitro and further narrowing down to nonamers by prediction using ProPred server enhances the potential of these nonamers to be potential candidates for immunodominance. However, further characterization is required to confirm which of these nonamers predicted are specifically responsible for such immune response.
There are few drawbacks in this study. Instead of single cytokine-secreting cells, recently polyfunctional cytokine-secreting cells are being studied which has not been attempted in this study. Regulatory T cell response is another area of interest which can unravel many hidden facts about regulatory nature of epitopes. The members of the cohort of TR recruited were not the same as PTB after treatment and in the present study. If the same PTB were studied after treatment, the response would be more meaningful with regard to T cell recognition pattern. Only HLA typing has been done and confirmation of peptide binding to HLA by peptide binding assays has not been done in this study. Moreover, HLA typing has been done in a few subjects and in silico analysis has been done with few HLA alleles in the present study. The number of subjects for HLA typing has to be increased to get a better picture of the scenario.
The C-terminal peptides of CFP-10 viz., Cfp7, Cfp8, and Cfp9 are recognized by HHC and not by PTB. The in silico approaches have showed two minimal epitopes CFP-1056–64 and CFP-1076–84 to be immunogenic which have prospects for vaccine development.
References
WHO Report (2006) Global tuberculosis control—surveillance, planning and financing
Zager EM, McNerney R (2008) Multidrug-resistant tuberculosis. BMC Infect Dis 8:10
Brosch R, Gordon SV, Marmiesse M, Brodin P, Buchrieser C, Eiglmeier K et al (2002) A new evolutionary scenario for the Mycobacterium tuberculosis complex. Proc Natl Acad Sci USA 99(6):3684–3689
Pym AS, Brodin P, Brosch R, Huerre M, Cole ST (2002) Loss of RD1 contributed to the attenuation of the live tuberculosis vaccines Mycobacterium bovis BCG and Mycobacterium microti. Mol Microbiol 46(3):709–717
Cole ST, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D et al (1998) Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393(6685):537–544
Harboe M, Oettinger T, Wiker HG, Rosenkrands I, Andersen P (1996) Evidence for occurrence of the ESAT-6 protein in Mycobacterium tuberculosis and virulent Mycobacterium bovis and for its absence in Mycobacterium bovis BCG. Infect Immun 64(1):16–22
Sorensen AL, Nagai S, Houen G, Andersen P, Andersen AB (1995) Purification and characterization of a low-molecular-mass T-cell antigen secreted by Mycobacterium tuberculosis. Infect Immun 63(5):1710–1717
Pym AS, Brodin P, Majlessi L, Brosch R, Demangel C, Williams A et al (2003) Recombinant BCG exporting ESAT-6 confers enhanced protection against tuberculosis. Nat Med 9(5):533–539
Bulat-Kardum L, Etokebe GE, Knezevic J, Balen S, Matacovic-Mileusnic N, Zaputovic L et al (2006) Interferon-γ receptor-1 gene promoter polymorphisms (G-611A; T-56C) and susceptibility to tuberculosis. Scand J Immunol 63:142–150
Arend SM, Geluk A, van Meijgaarden KE, van Dissel JT, Theisen M, Andersen P et al (2000) Antigenic equivalence of human T-cell responses to Mycobacterium tuberculosis-specific RD1-encoded protein antigens ESAT-6 and culture filtrate protein 10 and to mixtures of synthetic peptides. Infect Immun 68(6):3314–3321
Lalvani A, Brookes R, Wilkinson RJ, Malin AS, Pathan AA, Andersen P et al (1998) Human cytolytic and interferon gamma-secreting CD8+ T lymphocytes specific for Mycobacterium tuberculosis. Proc Natl Acad Sci U S A 95(1):270–275
Brandt L, Oettinger T, Holm A, Andersen AB, Andersen P (1996) Key epitopes on the ESAT-6 antigen recognized in mice during the recall of protective immunity to Mycobacterium tuberculosis. J Immunol 157(8):3527–3533
Lalvani A, Nagvenkar P, Udwadia Z, Pathan AA, Wilkinson KA, Shastri JS et al (2001) Enumeration of T cells specific for RD1-encoded antigens suggests a high prevalence of latent Mycobacterium tuberculosis infection in healthy urban Indians. J Infect Dis 183(3):469–477
Behr A, Hopewell PC, Paz EA, Kamamura LM, Schecter GF, Small PM (1998) Predictive value of contact investigation for identifying recent transmission of Mycobacterium tuberculosis. Am J Respir Crit Care Med 158:465–469
Selvakumar N, Vanajakumar PG, Gopi PG, Venkataramu KV, Datta M, Paramasivan N et al (1995) Isolation of tubercle bacilli from sputum samples of patients in the field studies by the cetylpyridinium chloride-sodium chloride and sodium hydroxide methods. Indian J Med Res 102:149–151
Trajkovic V, Singh G, Singh B, Singh S, Sharma P (2002) Effect of Mycobacterium tuberculosis-specific 10-kilodalton antigen on macrophage release of tumor necrosis factor alpha and nitric oxide. Infect Immun 70(12):6558–6566
Miller SA, Dykes DD, Polesky HF (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 16(3):1215
Bunce M, O’Neill CM, Barnardo MC, Krausa P, Browning MJ, Morris PJ et al (1995) Phototyping: comprehensive DNA typing for HLA-A, B, C, DRB1, DRB3, DRB4, DRB5 & DQB1 by PCR with 144 primer mixes utilizing sequence-specific primers (PCR-SSP). Tissue Antigens 46(5):355–367
Wang P, Sidney J, Dow C, Mothe B, Sette A, Peters B (2008) A systematic assessment of MHC class II peptide binding predictions and evaluation of a consensus approach. PLoS Comput Biol 4(4):e1000048
Singh H, Raghava GP (2001) ProPred: prediction of HLA-DR binding sites. Bioinformatics 17(12):1236–1237
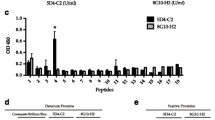
Tavares RCO, Salgado J, Moreira VB, Ferreira MA, Mello FC, Leung JW et al (2007) Interferon gamma response to combinations 38 kDa/CFP-10, 38 kDa/MPT-64, ESAT-6/MPT-64 and ESAT-6/CFP-10, each related to a single recombinant antigen of Mycobacterium tuberculosis in individuals from tuberculosis endemic areas. Microbiol Immunol 51(3):289–296
Doherty TM, Demissie A, Menzies D, Andersen P, Rook G, Zumla A (2005) Effect of sample handling on analysis of cytokine responses to Mycobacterium tuberculosis in clinical samples using ELISA, ELISPOT and quantitative RT-PCR. J Immunol Methods 298(1–2):129–141
Chapman AL, Munkanta M, Wilkinson KA, Pathan AA, Ewer K, Ayles H et al (2002) Rapid detection of active and latent tuberculosis infection in HIV-positive individuals by enumeration of Mycobacterium tuberculosis-specific T cells. AIDS 16(17):2285–2293
Skjot RL, Oettinger T, Rosenkrands I, Ravn P, Brock I, Jacobsen S et al (2000) Comparative evaluation of low-molecular-mass proteins from Mycobacterium tuberculosis identifies members of the ESAT-6 family as immunodominant T-cell antigens. Infect Immun 68(1):214–220
Flynn JL, Chan J, Triebold KJ, Dalton DK, Stewart TA, Bloom BR (1993) An essential role for interferon gamma in resistance to Mycobacterium tuberculosis infection. J Exp Med 178(6):2249–2254
Jouanguy E, Altare F, Lamhamedi S, Revy P, Emile JF, Newport M et al (1996) Interferon-gamma-receptor deficiency in an infant with fatal bacilli Calmette-Guérin infection. N Engl J Med 335(26):1956–1961
Havlir DV, Wallis RS, Boom WH, Daniel TM, Chervenak K, Ellner JJ (1991) Human immune response to Mycobacterium tuberculosis antigens. Infect Immun 9(2):665–670
Torres M, Herrera T, Villareal H, Rich EA, Sada E (1994) Cytokine profiles for peripheral blood lymphocytes from patients with active pulmonary tuberculosis and healthy household contacts in response to the 30-kilodalton antigen of Mycobacterium tuberculosis. Infect Immun 66(1):176–180
Grotzke JE, Lewinsohn DM (2005) Role of CD8+ T lymphocytes in control of Mycobacterium tuberculosis infection. Microbes Infect 7(4):776–788
Shams H, Klucar P, Weis SE, Lalvani A, Moonan PK, Safi H et al (2004) Characterization of a Mycobacterium tuberculosis peptide that is recognized by human CD4+ and CD8+ T cells in the context of multiple HLA alleles. J Immunol 173(3):1966–1977
Lewinsohn DM, Zhu L, Madison VJ, Dillon DC, Fling SP, Reed SG et al (2001) Classically restricted human CD8+ T lymphocytes derived from Mycobacterium tuberculosis-infected cells: definition of antigenic specificity. J Immunol 166(1):439–446
Kamath AB, Woodworth J, Xiong X, Taylor C, Weng Y, Behar SM (2004) Cytolytic CD8+ T cells recognizing CFP10 are recruited to the lung after Mycobacterium tuberculosis infection. J Exp Med 200(11):1479–1489
Brock I, Munk ME, Kok-Jensen A, Andersen P (2001) Performance of whole blood IFN-γ test for tuberculosis diagnosis based on PPD or the specific antigens ESAT-6 and CFP-10. Int J. Tuberc Lung Dis 5:462–467
Jo EK, Kim HJ, Lim JH, Min D, Song Y, Song C et al (2000) Dysregulated production of interferon-γ, interleukin-4 and interleukin-6 in early tuberculosis patients in response to antigen 85B of Mycobacterium tuberculosis. Scand J Immunol 51:209–217
Lee JS, Song CH, Kim CH, Kong SJ, Shon MH, Kim HJ et al (2002) Profiles of IFN-γ and its regulatory cytokines (IL-12, IL-18 and IL-10) in peripheral blood mononuclear cells from patients with multidrug-resistant tuberculosis. Clin Exp Immunol 128:16–524
Pathan AA, Wilkinson KA, Klenerman P, McShane H, Davidson RN, Pasvol G et al (2001) Direct ex vivo analysis of antigen-specific IFN-gamma-secreting CD4 T cells in Mycobacterium tuberculosis-infected individuals: associations with clinical disease state and effect of treatment. J Immunol 167(9):5217–5225
Carrara S, Vincenti D, Petrosillo N, Amicosante M, Girardi E, Goletti D (2004) Use of a T cell-based assay for monitoring efficacy of antituberculosis therapy. Clin Infect Dis 38(5):754–756
Eberl G, Renggli J, Men Y, Roggero MA, Lopez JA, Corradin G (1999) Extracellular processing and presentation of a 69-mer synthetic polypeptide to MHC class I-restricted T cells. Mol Immunol 36:103–112
Hernandez-Pando R, Aguilar D, Garcia Hernandez ML, Orozco H, Rook GAW (2004) Pulmonary tuberculosis in Balb/c mice with non-functional IL-4 genes; changes in the inflammatory effects of TNF-α in the regulation of fibrosis. Eur J Immunol 34:174–183
Ordway DJ, Costa L, Martins M, Silveira H, Amaral L, Arroz MJ et al (2004) Increased interleukin-4 production by CD8 and gamma delta T cells in health-care workers is associated with the subsequent development of active tuberculosis. J Infect Dis 190:756–766
Bothamley GH (2002) Treatment, tuberculosis, and human leukocyte antigen. Am J Respir Crit Care Med 166(7):907–908
Mustafa AS, Qvigstad E (1989) HLA-DR-restricted antigen-induced proliferation and cytotoxicity mediated by CD4+ T-cell clones from subjects vaccinated with killed M. leprae. Int J Lepr Other Mycobact Dis 57(1):1–11
Oftung F, Shinnick TM, Mustafa AS, Lundin KE, Godal T, Nerland AH (1990) Heterogeneity among human T cell clones recognizing an HLA-DR4, Dw4-restricted epitope from the 18-kDa antigen of Mycobacterium leprae defined by synthetic peptides. J Immunol 144(4):1478–1483
Mustafa AS, Lundin KE, Oftung F (1993) Human T cells recognize mycobacterial heat shock proteins in the context of multiple HLA-DR molecules: studies with healthy subjects vaccinated with Mycobacterium bovis BCG and Mycobacterium leprae. Infect Immun 61(12):5294–5301
Oftung F, Geluk A, Lundin KE, Meloen RH, Thole JE, Mustafa AS et al (1994) Mapping of multiple HLA class II-restricted T-cell epitopes of the mycobacterial 70-kilodalton heat shock protein. Infect Immun 62(12):5411–5418
De Groot AS, McMurry J, Marcon L, Franco J, Rivera D, Kutzler M et al (2005) Developing an epitope-driven tuberculosis (TB) vaccine. Vaccine 23(17–18):2121–2131
Ravikumar M, Dheenadhayalan V, Rajaram K, Lakshmi SS, Kumaran PP, Paramasivan CN et al (1999) Associations of HLA-DRB1, DQB1 and DPB1 alleles with pulmonary tuberculosis in south India. Tuber Lung Dis 79(5):309–317
Acknowledgments
Mr. Madhan Kumar is a recipient of Senior Research Fellowship from Council of Scientific and Industrial Research (CSIR), New Delhi, India. Overlapping peptides of CFP-10 were a kind gift from Dr. Thomas B Nutman, NIAID/NIH, USA (under ICER/TRC program). CFP-10 whole protein was a gift from Dr. Pawan Sharma, ICGEB (International Centre for Genetic Engineering and Biotechnology), New Delhi. We would like to thank Mr. Anbalagan and Mr. Murugesh for their help in acquiring samples in flow cytometry. We are grateful to patients, healthy household contacts, and treated patients who participated in this study and gave blood. The help rendered by the RNTCP staff Mrs. Kasthuri and Mrs. Thilagavathi in recruiting patients is kindly acknowledged. This project was not a funded project.
Conflict of interest statement
None.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kumar, M., Sundaramurthi, J.C., Mehra, N.K. et al. Cellular immune response to Mycobacterium tuberculosis-specific antigen culture filtrate protein-10 in south India. Med Microbiol Immunol 199, 11–25 (2010). https://doi.org/10.1007/s00430-009-0129-2
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00430-009-0129-2