Abstract
MicroRNAs (miRNAs) are small, noncoding RNA molecules that regulate gene expression post-transcriptionally through complementary base pairing with thousands of messenger RNAs. Although the precise biological functions of individual miRNAs are still unknown, miRNAs are speculated to play important roles in diverse biological processes through fine regulation of their target gene expression. A growing body of data indicates the deregulation of miRNAs during hepatocarcinogenesis. In this review, we summarize recent findings regarding deregulated miRNA expression and their possible target genes in hepatocarcinogenesis, with emphasis on inflammation-related hepatocarcinogenesis. Because miRNA-based strategies are being applied to clinical therapeutics, precise knowledge of miRNA functions is crucial both scientifically and clinically. We discuss the current open questions from these points of view, which must be clarified in the near future.
Similar content being viewed by others
Introduction
MicroRNAs (miRNAs) are short, single-stranded, non-coding RNAs, which are expressed in most organisms, from plants to vertebrates [1]. Since the discovery of the miRNA lin-4 in Caenorhabditis elegans [2, 3], 1,872 miRNA precursors and 2,578 mature miRNA sequences in humans have been deposited in miRBase, a public repository hosted by the Sanger Institute, as of November 2013 [4]. Bioinformatic predictions suggest that miRNAs regulate more than 30 % of human protein-coding genes [5–7]. Through the regulation of gene expression, miRNAs are involved in various physiological and pathological processes, including cell proliferation, apoptosis, differentiation, metabolism, oncogenesis and oncogenic suppression [8, 9]. Thus, it is not surprising that deregulation of miRNAs is linked closely to various human pathological conditions. In this review, we will describe the crucial role of miRNAs in liver carcinogenesis, especially inflammation-related hepatocarcinogenesis.
Biogenesis and functions of miRNAs
Transcription is the first step in miRNA expression (Fig. 1). Similar to most protein-coding genes, transcriptional factors, enhancers and silencers are involved in miRNA transcription [10–12]. Epigenetic mechanisms, such as promoter methylation or histone modification, also regulate miRNA transcription, and it was shown that histone deacetylase (HDAC) inhibition results in transcriptional changes in ~40 % of miRNAs [13].
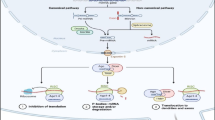
Biogenesis of miRNAs. The primary miRNA transcript (pri-miRNA) is transcribed from the genome by RNA polymerase II or III. The microprocessor complex Drosha–DGCR8 cleaves the pri-miRNA into the precursor hairpin, pre-miRNA in the nucleus. The pre-miRNA is exported from the nucleus by exportin-5–Ran-GTP. In the cytoplasm, the RNase Dicer in complex with the double-stranded RNA-binding protein, TRBP, cleaves the pre-miRNA hairpin to its mature length. The functional strand of the mature miRNA is loaded together with Argonaute (Ago2) proteins into the RNA-induced silencing complex (RISC), where it guides RISC to silence target mRNAs through mRNA cleavage or translational repression. The passenger strand (black) is degraded
Primary miRNAs, which possess stem-loop structures, are transcribed by RNA polymerase II [8]. These pri-miRNAs are processed by a microprocessor complex comprising Drosha (RNAase III) [14] and DGCR8/Pasha [15] in the nucleus [16]. The processed products are approximately 65-nucleotide hairpin-shaped precursors (pre-miRNAs) that are transported to the cytoplasm via exportin-5 [17, 18]. Pre-miRNAs are further cleaved into mature miRNAs by Drosha and Dicer RNA polymerase III. Mature miRNA duplexes are loaded onto an RNA-induced silencing complex (RISC) and are unwound into the single-stranded mature form [19–21]. The resulting co-complex directly targets the 3′-untranslated regions (3′-UTRs) of target mRNAs, depending on the sequence similarities, to negatively regulate their expression by enhancing mRNA cleavage or inhibiting translation (Fig. 1) [8, 22]. Because most miRNAs guide the recognition of imperfect matches of target mRNAs, individual miRNAs have multiple (probably hundreds) of mRNA targets. In addition, multiple miRNAs can cooperate to regulate the expression of the same transcript [6]. Thus, depending upon the identity of the target mRNAs, miRNAs play roles as “fine-tuners of gene expression” in the control of various biological functions.
Identifying functionally important miRNA target genes is crucial for understanding the impact of specific miRNAs on cellular function. However, this is challenging because miRNAs usually have imperfect complementarity with their targets [22]. In mammals, the most consistent requirement for miRNA-target interaction, although not always essential, is a contiguous and perfect pairing of the miRNA (nt 2–8), representing the “seed” sequence [22]. In many cases, the seed sequences determine this recognition, but in other cases, additional determinants are required, such as reasonable complementarity to the miRNA 3′ half to stabilize the interaction. In addition, target pairing to the center of some miRNAs has also been reported [23]. Although public miRNA target prediction algorithms, such as TargetScan [24] and PicTar [25], have facilitated the rapid identification of miRNA target genes [22], candidates should be validated experimentally.
miRNAs and cancer
The involvement of miRNAs in cancer pathogenesis is well established. miRNAs can affect six hallmarks of malignant cells, which are (1) self-sufficiency in growth signals, (2) insensitivity to anti-growth signals, (3) evasion of apoptosis, (4) limitless replicative potential, (5) angiogenesis, and (6) invasion and metastasis [26]. miRNAs are frequently up- or downregulated in malignant tissues and can be considered oncogenes or tumor suppressors, respectively. However, it is essential to test experimentally whether the deregulated miRNAs are actually causative to carcinogenesis, since miRNAs have a very restricted tissue-specific expression and the apparent miRNA modulation in cancer tissues may only reflect the different constituents of a cell population as compared to normal tissues. Extensive analyses have confirmed the causative roles of miRNAs in cancer by using either human cancer cells or genetically engineered animal models, such as transgenic expression of miR-155, miR-21 and miR-15-a/16-1, which are sufficient to initiate lymphomagenesis in mice [27–29]. These results suggest the potential role of miRNAs in the pathogenesis of carcinogenesis and as therapeutic targets.
miRNAs and hepatocarcinogenesis
Numerous reports regarding the deregulated expression of miRNAs in human hepatocellular carcinoma (HCC) are extant. Most studies compared the miRNA expression levels between cancer tissues and background non-tumorous tissues, selected candidate miRNA(s) and revealed their target genes, which may be involved in carcinogenesis. As shown in Tables 1 and 2, many miRNAs have been identified as downregulated or upregulated in recent studies (Tables 1, 2). However, these numerous results are not always superimposable due to the large variances in the results. These significant differences may be due to several reasons, such as the use of different techniques or different samples as controls, normal liver tissues versus peritumoral non-neoplastic tissues. In addition, one may need to take into consideration the fact that HCCs arise in background livers with different etiologies, such as hepatitis B, hepatitis C or steatohepatitis, and also the age or sex of the tissue-derived patients and background liver condition, such as fibrosis staging or inflammation activity, which may result in differences in the expression status of miRNAs. Despite these considerable limitations, the list suggests that diverse miRNAs play crucial roles in hepatocarcinogenesis. We will briefly describe some of them below.
The expression levels of miRNAs have restricted tissue specificities. In the liver, miR-122, miR-192 and miR-199a/b-3p are the three most expressed miRNAs, accounting for 52, 17 and 5 % of all mRNAs in the tissues, respectively [30]. The tumorigenic role of the loss of miR-122 was confirmed in gene-knockout mice [31, 32] and its expression is indeed decreased in half of the HCCs, especially non-viral HCCs [30]. We also reported that decreased expression of miR-122 is linked with poor prognosis of HCC [33]. While miR-192 does not appear to be deregulated in HCC samples in previous studies, miR-199a/b-3p is decreased with high frequency in HCC, which is closely linked to a poor prognosis of HCC [30]. In contrast, miR-21, whose expression is increased following rat hepatectomy [34], is upregulated as a known oncomiRNA and represses PTEN signaling, resulting in promotion of HCC development [35]. Although individual miRNAs may be involved in hepatocarcinogenesis, because miRNAs often function co-operatively, the extent of their involvement remains to be determined.
As described above, miRNAs usually have multiple mRNA targets. Thus, it is not practical to describe only a few genes as being responsible for the phenotypes by deregulation of specific miRNAs, while many studies identify specific genes as targets of specific miRNAs. Nonetheless, the identified targeted genes are generally related to at least one of the hallmarks of cancer, such as cell growth, apoptosis, invasion, and so on. These results suggest that the deregulation of miRNA expression might mediate hepatocarcinogenesis through deregulating the expression of their target genes.
The miRNAs identified as deregulated in hepatocarcinogenesis may be useful as diagnostic and prognostic markers [36], because miRNAs in the circulation are reported to be relatively stable [37]. Also, deregulated miRNAs may be candidate therapeutic and preventive targets against HCC. However, to include the obtained results in clinical interventional applications, it is necessary to confirm if the deregulated miRNAs are truly drivers or are simply passive in hepatocarcinogenesis. To this end, genetically modified mice may provide some information. In addition, to correctly interpret the data, a standard method of normalizing the microRNAome data between studies may also be crucial. Since there are multiple target genes of miRNAs and, conversely, one transcript can be targeted by multiple miRNAs, a more systematic comparison using miRNA data, transcriptome data and proteome data would increase our understanding of the consequences of the deregulation of miRNAs during hepatocarcinogenesis. From this point of view, systematic and comprehensive target gene analyses for in silico systems biology models may be one option to resolve these issues.
miRNAs linked to inflammation-mediated hepatocarcinogenesis
Inflammation is considered to be a major cause of cancer [38, 39]. In the liver, hepatocarcinogenesis frequently occurs in persistently inflamed liver tissues caused by chronic hepatitis viral infection or non-alcoholic steatohepatitis. However, the molecular linkage between chronic inflammation and carcinogenesis is not well characterized. miRNAs, as a new class of gene expression regulators, may be involved in chronic inflammation-induced carcinogenesis and, in fact, several studies have clarified one such linkage, in which miRNAs may serve as a bridge between continuous inflammation and carcinogenesis.
A flagship report addresses a positive feedback loop of an inflammatory response mediated by NF-κB that activates Lin28B transcription (Fig. 2) [40]. LIN28B, which is an inhibitor of miRNA processing, reduces let-7 levels. Let-7 inhibits IL-6 expression, resulting in higher levels of IL-6 than achieved by NF-κB activation. IL-6-mediated STAT3 activation is necessary for transformation and IL-6 activates NF-κB, completing a positive feedback loop. Although the experiments mainly used MCF10A cells (breast cancer cells), a similar feedback loop was observed in HCC tissues. The authors termed these mechanisms an “epigenetic switch” because the loop maintains the epigenetic transformed state even in the absence of induction by inflammation (Fig. 2).
A model bridging chronic inflammation and transformation by miRNA. Inflammation triggers activation of NF-κB, which leads to transcription of LIN28B. LIN28B inhibits the production of Let-7. Let-7 normally inhibits IL-6 expression, resulting in higher levels of IL-6 than are achieved by NF-κB activation. IL-6 mediated STAT3 activation is necessary for transformation and IL-6 activates NF-κB, completing a positive feedback loop
Another report addressed hepatocarcinogenesis induced by transient inhibition of HNF4α (Fig. 3) [41]. HNF4α was reported to be involved in liver oncogenesis, although discrepant reports have also been published [42–44]. In that report, transient HNF4α silencing was sufficient to maintain cell transformation. Through a miRNA library screen, miR-24 and miR-629 were identified to target HNF4α. Interestingly, both miRNAs were induced following HNF4α silencing, supporting their involvement in the HNF4α-dependent feedback loop. miR-24 and miR-629 contain the STAT3-binding motif in their promoter region. The authors showed that in response to IL-6, STAT3 binding to their promoters increased, resulting in miRNA expression. They also identified miR-124, whose promoter region contains HNF4α binding sites. miR-124 targets IL-6R and, thus, HNF4α silencing results in reduced expression of miR-124 and enhanced expression of IL-6R and activation of STAT3. The importance of these feedback loops was confirmed in vivo using a mouse HCC model induced by diethylnitrosamine. miR-124 delivery by cationic liposomes prevented tumor development. Thus, these microRNA feedback-inflammatory loops are important and can be a therapeutic target for liver cancer (Fig. 3) [41].
A model describing a positive feedback loop mediated by miRNAs from transient HNF4α inhibition to transformation. Transient silencing of HNF4α is mediated by miR-24 and miR-629, both of which are induced by STAT3 activation following IL-6 stimulation. miR-124, whose promoter region contains HNF4α-binding sites, targets IL-6R and, thus, HNF4α silencing results in reduced expression of miR-124 and enhanced expression of IL-6R and activation of STAT3, which induces miR-24 and miR-629. This microRNA feedback-inflammatory loop is thought to be crucial in IL-6-mediated liver cancer
A recent paper reported a similar but distinct observation (Fig. 4). The authors found that when using DEN-induced foci of altered hepatocytes (FAH), LIN28-expressing cells are present in FAH, in which let-7 is down-regulated, resulting in the enhanced expression of IL-6, mediating the progression of malignancies from progenitors. An important difference between the cells in FAH and those in early hepatocarcinogenesis is that IL-6 signaling is autocrine, being mediated by reduced let-7 due to upregulation of LIN28B in FAH cells. This mechanism may contribute to malignant progression from HCC progenitor cells (Fig. 4) [45].
A model bridging the malignant transformation of precursor cells and autocrine-mediated inflammation by microRNA. LIN28-expressing cells exist in the foci of altered hepatocytes, in which let-7 is downregulated, resulting in enhanced IL-6 expression, which mediates the progression of malignancies from progenitor cells
These three reports are from related research groups, and rely on the hypothesis that the IL-6-STAT3 pathway is crucial for hepatocarcinogenesis. Although IL-6 has been implicated as a growth factor in various epithelial cancers [46, 47], its relevance in hepatocarcinogenesis needs to be confirmed to determine the applicability and reproducibility of these findings to the clinical setting.
miRNAs as therapeutic targets in the liver
Recently, miravirsen, a LNA-modified DNA phosphorothioate antisense oligonucleotide against miR-122, became the first miRNA-targeting drug for clinical use [48]. It was developed to target HCV, as the stability and propagation of this virus is dependent on a functional interaction between the HCV genome and miR-122 [49, 50]. No harmful events were observed in Phase I studies in healthy volunteers, and Phase II studies proceeded to evaluate the safety and efficacy of miravirsen in 36 patients with chronic HCV genotype 1 infection. The patients were randomly assigned to receive 5 weeks of subcutaneous miravirsen injections at 3, 5 or 7 mg per kg body weight or a placebo over a 29-day period. Miravirsen resulted in a dose-dependent reduction in HCV levels, without major adverse events and with no escape mutations in the miR-122 binding sites of the HCV genome [48]. The success of miravirsen is promising, not only as a novel anti-HCV drug, but also as the first trial of miRNA-targeting therapy.
In addition to miravirsen, a clinical trial of MRX34 as a mimic of miR-34 is underway. MRX34 is a liposome-formulated mimic of the tumor suppressor miR-34 (Mirna Therapeutics, Austin, TX, USA). Further study of MRX34 is being conducted by Mirna Therapeutics, which initiated a Phase I study in May 2013 to examine the effects of MRX34 on unresectable primary liver cancer or advanced or metastatic cancer with liver involvement (ClinicalTrials.gov Identifier: NCT01829971). If these oligonucleotide therapies are successful, therapeutic options based on the numerous miRNAs deregulated during hepatocarcinogenesis appear promising [51].
Issues to be resolved in miRNA involvement in hepatocarcinogenesis
As described above, along with recent discoveries of the diverse effects of miRNAs in hepatocarcinogenesis, miRNA-mediated intervention is promising for the development of new diagnostic, preventive and therapeutic tools. However, the data obtained to date are far from complete. The following are some of the critical issues that we believe need to be resolved.
-
1.
The reason for the non-reproducible results among studies should be determined to utilize the available data more reasonably and efficiently.
-
2.
Identification of crucial driver miRNAs among the diverse deregulated miRNAs is critical to develop useful therapeutics in clinics, although even passive miRNAs may be utilized as markers for diagnosis or prediction of prognosis.
-
3.
Comprehensive target gene analyses using in silico systems biology models should be applied.
-
4.
For effective interventions using miRNA, the delivery method, improved oligonucleotide modification and safety must be further considered. Since miRNAs generally have diverse effects due to targeting multiple mRNAs, undesired outcomes, so called off-target effects, may be encountered, even when a specific miRNA is targeted.
Finding solutions to these issues should be considered as critically important for the near future in order to understand more fully the physiological function of miRNAs in hepatocarcinogenesis and utilize this knowledge in translational research.
Conclusions
The discovery of miRNA has, without doubt, opened up new possibilities for understanding the molecular mechanisms of gene regulation. As numerous findings regarding miRNA, from diverse perspectives, have been reported, the speed of discovery in this field is astonishing. In fact, novel therapeutics targeting miRNAs have already been successfully applied in clinical trials. Some miRNAs may be useful as novel biomarkers. Additionally, the discovery of novel concepts in the pathogenesis of hepatocarcinogenesis frequently involves miRNA. On the other hand, several important issues remain to be resolved in this field. Thus, continuous research in this field is still necessary to develop truly innovative concepts in our understanding of pathogenesis related to miRNA and to transform the obtained knowledge into real clinical applications.
References
Carrington J, Ambros V. Role of microRNAs in plant and animal development. Science. 2003;301:336–8.
Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75:843–54.
Wightman B, Ha I, Ruvkun G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell. 1993;75:855–62.
Kozomara A, Griffiths-Jones S. miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. 2011;39:D152–7.
John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS. Human MicroRNA targets. PLoS Biol. 2004;2:e363.
Krek A, Grün D, Poy MN, Wolf R, Rosenberg L, Epstein EJ, et al. Combinatorial microRNA target predictions. Nat Genet. 2005;37:495–500.
Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120:15–20.
Ambros V. The functions of animal microRNAs. Nature. 2004;431:350–5.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
Cai X, Hagedorn CH, Cullen BR. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA. 2004;10:1957–66.
Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek SH, et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 2004;23:4051–60.
Borchert GM, Lanier W, Davidson BL. RNA polymerase III transcribes human microRNAs. Nat Struct Mol Biol. 2006;13:1097–101.
Scott GK, Mattie MD, Berger CE, Benz SC, Benz CC. Rapid alteration of microRNA levels by histone deacetylase inhibition. Cancer Res. 2006;66:1277–81.
Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J, et al. The nuclear RNase III Drosha initiates microRNA processing. Nature. 2003;425:415–9.
Han J, Lee Y, Yeom KH, Kim YK, Jin H, Kim VN. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 2004;18:3016–27.
Denli AM, Tops BB, Plasterk RH, Ketting RF, Hannon GJ. Processing of primary microRNAs by the microprocessor complex. Nature. 2004;432:231–5.
Yi R, Qin Y, Macara IG, Cullen BR. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 2003;17:3011–6.
Lund E, Güttinger S, Calado A, Dahlberg JE, Kutay U. Nuclear export of microRNA precursors. Science. 2004;303:95–8.
Maniataki E, Mourelatos Z. A human, ATP-independent, RISC assembly machine fueled by pre-miRNA. Genes Dev. 2005;19:2979–90.
Mourelatos Z, Dostie J, Paushkin S, Sharma A, Charroux B, Abel L, et al. miRNPs: a novel class of ribonucleoproteins containing numerous microRNAs. Genes Dev. 2002;16:720–8.
Gregory RI, Chendrimada TP, Cooch N, Shiekhattar R. Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell. 2005;123:631–40.
Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136:215–33.
Shin C, Nam JW, Farh KK, Chiang HR, Shkumatava A, Bartel DP. Expanding the microRNA targeting code: functional sites with centered pairing. Mol Cell. 2010;38:789–802.
Friedman RC, Farh KK, Burge CB, Bartel DP. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009;19:92–105.
Lall S, Grün D, Krek A, Chen K, Wang YL, Dewey CN, et al. A genome-wide map of conserved microRNA targets in C. elegans. Curr Biol. 2006;16:460–71.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Costinean S, Sandhu SK, Pedersen IM, Tili E, Trotta R, Perrotti D, et al. Src homology 2 domain-containing inositol-5-phosphatase and CCAAT enhancer-binding protein beta are targeted by miR-155 in B cells of Emicro-MiR-155 transgenic mice. Blood. 2009;114:1374–82.
Medina PP, Nolde M, Slack FJ. OncomiR addiction in an in vivo model of microRNA-21-induced pre-B-cell lymphoma. Nature. 2010;467:86–90.
Klein U, Lia M, Crespo M, Siegel R, Shen Q, Mo T, et al. The DLEU2/miR-15a/16-1 cluster controls B cell proliferation and its deletion leads to chronic lymphocytic leukemia. Cancer Cell. 2010;17:28–40.
Hou J, Lin L, Zhou W, Wang Z, Ding G, Dong Q, et al. Identification of miRNomes in human liver and hepatocellular carcinoma reveals miR-199a/b-3p as therapeutic target for hepatocellular carcinoma. Cancer Cell. 2011;19:232–43.
Hsu SH, Wang B, Kota J, Yu J, Costinean S, Kutay H, et al. Essential metabolic, anti-inflammatory, and anti-tumorigenic functions of miR-122 in liver. J Clin Invest. 2012;122:2871–83.
Tsai WC, Hsu SD, Hsu CS, Lai TC, Chen SJ, Shen R, et al. MicroRNA-122 plays a critical role in liver homeostasis and hepatocarcinogenesis. J Clin Invest. 2012;122:2884–97.
Kojima K, Takata A, Vadnais C, Otsuka M, Yoshikawa T, Akanuma M, et al. MicroRNA122 is a key regulator of α-fetoprotein expression and influences the aggressiveness of hepatocellular carcinoma. Nat Commun. 2011;2:338.
Castro RE, Ferreira DM, Zhang X, Borralho PM, Sarver AL, Zeng Y, et al. Identification of microRNAs during rat liver regeneration after partial hepatectomy and modulation by ursodeoxycholic acid. Am J Physiol Gastrointest Liver Physiol. 2010;299:G887–97.
Meng F, Henson R, Wehbe-Janek H, Ghoshal K, Jacob ST, Patel T. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology. 2007;133:647–58.
Chiu LY, Kishnani PS, Chuang TP, Tang CY, Liu CY, Bali D, et al. Identification of differentially expressed microRNAs in human hepatocellular adenoma associated with type I glycogen storage disease: a potential utility as biomarkers. J Gastroenterol. 2013 (in press).
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA. 2008;105:10513–8.
Clevers H. At the crossroads of inflammation and cancer. Cell. 2004;118:671–4.
Grivennikov SI, Greten FR, Karin M. Immunity, inflammation, and cancer. Cell. 2010;140:883–99.
Iliopoulos D, Hirsch HA, Struhl K. An epigenetic switch involving NF-kappaB, Lin28, Let-7 MicroRNA, and IL6 links inflammation to cell transformation. Cell. 2009;139:693–706.
Hatziapostolou M, Polytarchou C, Aggelidou E, Drakaki A, Poultsides GA, Jaeger SA, et al. An HNF4α-miRNA inflammatory feedback circuit regulates hepatocellular oncogenesis. Cell. 2011;147:1233–47.
Horiguchi N, Takayama H, Toyoda M, Otsuka T, Fukusato T, Merlino G, et al. Hepatocyte growth factor promotes hepatocarcinogenesis through c-Met autocrine activation and enhanced angiogenesis in transgenic mice treated with diethylnitrosamine. Oncogene. 2002;21:1791–9.
Ning BF, Ding J, Yin C, Zhong W, Wu K, Zeng X, et al. Hepatocyte nuclear factor 4 alpha suppresses the development of hepatocellular carcinoma. Cancer Res. 2010;70:7640–51.
Walesky C, Edwards G, Borude P, Gunewardena S, O’Neil M, Yoo B, et al. Hepatocyte nuclear factor 4 alpha deletion promotes diethylnitrosamine-induced hepatocellular carcinoma in rodents. Hepatology. 2013;57:2480–90.
He G, Dhar D, Nakagawa H, Font-Burgada J, Ogata H, Jiang Y, et al. Identification of liver cancer progenitors whose malignant progression depends on autocrine IL-6 signaling. Cell. 2013;155:384–96.
Akira S, Kishimoto T. The evidence for interleukin-6 as an autocrine growth factor in malignancy. Semin Cancer Biol. 1992;3:17–26.
He G, Karin M. NF-κB and STAT3—key players in liver inflammation and cancer. Cell Res. 2011;21:159–68.
Janssen HL, Reesink HW, Lawitz EJ, Zeuzem S, Rodriguez-Torres M, Patel K, et al. Treatment of HCV infection by targeting microRNA. N Engl J Med. 2013;368:1685–94.
Jopling CL, Yi M, Lancaster AM, Lemon SM, Sarnow P. Modulation of hepatitis C virus RNA abundance by a liver-specific MicroRNA. Science. 2005;309:1577–81.
Fukuhara T, Matsuura Y. Role of miR-122 and lipid metabolism in HCV infection. J Gastroenterol. 2013;48:169–76.
Bouchie A. First microRNA mimic enters clinic. Nat Biotechnol. 2013;31:577.
Yang F, Yin Y, Wang F, Wang Y, Zhang L, Tang Y, et al. miR-17-5p Promotes migration of human hepatocellular carcinoma cells through the p38 mitogen-activated protein kinase-heat shock protein 27 pathway. Hepatology. 2010;51:1614–23.
Liu WH, Yeh SH, Lu CC, Yu SL, Chen HY, Lin CY, et al. MicroRNA-18a prevents estrogen receptor-alpha expression, promoting proliferation of hepatocellular carcinoma cells. Gastroenterology. 2009;136:683–93.
Wang B, Majumder S, Nuovo G, Kutay H, Volinia S, Patel T, et al. Role of microRNA-155 at early stages of hepatocarcinogenesis induced by choline-deficient and amino acid-defined diet in C57BL/6 mice. Hepatology. 2009;50:1152–61.
Jiang R, Deng L, Zhao L, Li X, Zhang F, Xia Y, et al. miR-22 promotes HBV-related hepatocellular carcinoma development in males. Clin Cancer Res. 2011;17:5593–603.
Wang B, Hsu SH, Frankel W, Ghoshal K, Jacob ST. Stat3-mediated activation of microRNA-23a suppresses gluconeogenesis in hepatocellular carcinoma by down-regulating glucose-6-phosphatase and peroxisome proliferator-activated receptor gamma, coactivator 1 alpha. Hepatology. 2012;56:186–97.
Fu X, Meng Z, Liang W, Tian Y, Wang X, Han W, et al. miR-26a enhances miRNA biogenesis by targeting Lin28B and Zcchc11 to suppress tumor growth and metastasis. Oncogene. 2013 (in press).
Ji J, Shi J, Budhu A, Yu Z, Forgues M, Roessler S, et al. MicroRNA expression, survival, and response to interferon in liver cancer. N Engl J Med. 2009;361:1437–47.
Yao J, Liang L, Huang S, Ding J, Tan N, Zhao Y, et al. MicroRNA-30d promotes tumor invasion and metastasis by targeting Galphai2 in hepatocellular carcinoma. Hepatology. 2010;51:846–56.
Varnholt H, Drebber U, Schulze F, Wedemeyer I, Schirmacher P, Dienes HP, et al. MicroRNA gene expression profile of hepatitis C virus-associated hepatocellular carcinoma. Hepatology. 2008;47:1223–32.
Shen G, Jia H, Tai Q, Li Y, Chen D. miR-106b downregulates adenomatous polyposis coli and promotes cell proliferation in human hepatocellular carcinoma. Carcinogenesis. 2013;34:211–9.
Ma S, Tang KH, Chan YP, Lee TK, Kwan PS, Castilho A, et al. miR-130b Promotes CD133(+) liver tumor-initiating cell growth and self-renewal via tumor protein 53-induced nuclear protein 1. Cell Stem Cell. 2010;7:694–707.
Liu S, Guo W, Shi J, Li N, Yu X, Xue J, et al. MicroRNA-135a contributes to the development of portal vein tumor thrombus by promoting metastasis in hepatocellular carcinoma. J Hepatol. 2012;56:389–96.
Zhang X, Liu S, Hu T, He Y, Sun S. Up-regulated microRNA-143 transcribed by nuclear factor kappa B enhances hepatocarcinoma metastasis by repressing fibronectin expression. Hepatology. 2009;50:490–9.
Zhu K, Pan Q, Zhang X, Kong LQ, Fan J, Dai Z, et al. MiR-146a enhances angiogenic activity of endothelial cells in hepatocellular carcinoma by promoting PDGFRA expression. Carcinogenesis. 2013;34:2071–9.
Luedde T. MicroRNA-151 and its hosting gene FAK (focal adhesion kinase) regulate tumor cell migration and spreading of hepatocellular carcinoma. Hepatology. 2010;52:1164–6.
Ding J, Huang S, Wu S, Zhao Y, Liang L, Yan M, et al. Gain of miR-151 on chromosome 8q24.3 facilitates tumour cell migration and spreading through downregulating RhoGDIA. Nat Cell Biol. 2010;12:390–9.
Yan XL, Jia YL, Chen L, Zeng Q, Zhou JN, Fu CJ, et al. Hepatocellular carcinoma-associated mesenchymal stem cells promote hepatocarcinoma progression: role of the S100A4-miR155-SOCS1-MMP9 axis. Hepatology. 2013;57:2274–86.
Zhang S, Shan C, Kong G, Du Y, Ye L, Zhang X. MicroRNA-520e suppresses growth of hepatoma cells by targeting the NF-κB-inducing kinase (NIK). Oncogene. 2012;31:3607–20.
Wang B, Hsu SH, Majumder S, Kutay H, Huang W, Jacob ST, et al. TGFbeta-mediated upregulation of hepatic miR-181b promotes hepatocarcinogenesis by targeting TIMP3. Oncogene. 2010;29:1787–97.
Ji J, Yamashita T, Budhu A, Forgues M, Jia HL, Li C, et al. Identification of microRNA-181 by genome-wide screening as a critical player in EpCAM-positive hepatic cancer stem cells. Hepatology. 2009;50:472–80.
Goeppert B, Schmezer P, Dutruel C, Oakes C, Renner M, Breinig M, et al. Down-regulation of tumor suppressor A kinase anchor protein 12 in human hepatocarcinogenesis by epigenetic mechanisms. Hepatology. 2010;52:2023–33.
Petrelli A, Perra A, Cora D, Sulas P, Menegon S, Manca C, et al. MiRNA/gene profiling unveils early molecular changes and NRF2 activation in a rat model recapitulating human HCC. Hepatology. 2013 (in press).
Ying Q, Liang L, Guo W, Zha R, Tian Q, Huang S, et al. Hypoxia-inducible microRNA-210 augments the metastatic potential of tumor cells by targeting vacuole membrane protein 1 in hepatocellular carcinoma. Hepatology. 2011;54:2064–75.
Chen PJ, Yeh SH, Liu WH, Lin CC, Huang HC, Chen CL, et al. Androgen pathway stimulates microRNA-216a transcription to suppress the tumor suppressor in lung cancer-1 gene in early hepatocarcinogenesis. Hepatology. 2012;56:632–43.
Xia H, Ooi LL, Hui KM. MicroRNA-216a/217-induced epithelial–mesenchymal transition targets PTEN and SMAD7 to promote drug resistance and recurrence of liver cancer. Hepatology. 2013;58:629–41.
Callegari E, Elamin BK, Giannone F, Milazzo M, Altavilla G, Fornari F, et al. Liver tumorigenicity promoted by microRNA-221 in a mouse transgenic model. Hepatology. 2012;56:1025–33.
Yuan Q, Loya K, Rani B, Möbus S, Balakrishnan A, Lamle J, et al. MicroRNA-221 overexpression accelerates hepatocyte proliferation during liver regeneration. Hepatology. 2013;57:299–310.
Gramantieri L, Fornari F, Ferracin M, Veronese A, Sabbioni S, Calin GA, et al. MicroRNA-221 targets Bmf in hepatocellular carcinoma and correlates with tumor multifocality. Clin Cancer Res. 2009;15:5073–81.
Fornari F, Gramantieri L, Ferracin M, Veronese A, Sabbioni S, Calin GA, et al. MiR-221 controls CDKN1C/p57 and CDKN1B/p27 expression in human hepatocellular carcinoma. Oncogene. 2008;27:5651–61.
Pineau P, Volinia S, McJunkin K, Marchio A, Battiston C, Terris B, et al. miR-221 overexpression contributes to liver tumorigenesis. Proc Natl Acad Sci USA. 2010;107:264–9.
Ladeiro Y, Couchy G, Balabaud C, Bioulac-Sage P, Pelletier L, Rebouissou S, et al. MicroRNA profiling in hepatocellular tumors is associated with clinical features and oncogene/tumor suppressor gene mutations. Hepatology. 2008;47:1955–63.
Lan SH, Wu SY, Zuchini R, Lin XZ, Su IJ, Tsai TF, et al. Autophagy suppresses tumorigenesis of hepatitis B virus-associated hepatocellular carcinoma through degradation of miR-224. Hepatology. 2013 (in press).
Scisciani C, Vossio S, Guerrieri F, Schinzari V, De Iaco R, D’Onorio de Meo P, et al. Transcriptional regulation of miR-224 upregulated in human HCCs by NFκB inflammatory pathways. J Hepatol. 2012;56:855–61.
Gao P, Wong CC, Tung EK, Lee JM, Wong CM, Ng IO. Deregulation of microRNA expression occurs early and accumulates in early stages of HBV-associated multistep hepatocarcinogenesis. J Hepatol. 2011;54:1177–84.
Wang Y, Lee AT, Ma JZ, Wang J, Ren J, Yang Y, et al. Profiling microRNA expression in hepatocellular carcinoma reveals microRNA-224 up-regulation and apoptosis inhibitor-5 as a microRNA-224-specific target. J Biol Chem. 2008;283:13205–15.
Lin J, Huang S, Wu S, Ding J, Zhao Y, Liang L, et al. MicroRNA-423 promotes cell growth and regulates G(1)/S transition by targeting p21Cip1/Waf1 in hepatocellular carcinoma. Carcinogenesis. 2011;32:1641–7.
Yang H, Cho ME, Li TW, Peng H, Ko KS, Mato JM, et al. MicroRNAs regulate methionine adenosyltransferase 1A expression in hepatocellular carcinoma. J Clin Invest. 2013;123:285–98.
Zhang LY, Liu M, Li X, Tang H. miR-490-3p modulates cell growth and epithelial to mesenchymal transition of hepatocellular carcinoma cells by targeting endoplasmic reticulum-Golgi intermediate compartment protein 3 (ERGIC3). J Biol Chem. 2013;288:4035–47.
Lim L, Balakrishnan A, Huskey N, Jones KD, Jodari M, Ng R, et al. MiR-494 within an oncogenic MicroRNA megacluster regulates G1/S transition in liver tumorigenesis through suppression of MCC. Hepatology. 2013 (in press).
Toffanin S, Hoshida Y, Lachenmayer A, Villanueva A, Cabellos L, Minguez B, et al. MicroRNA-based classification of hepatocellular carcinoma and oncogenic role of miR-517a. Gastroenterology. 2011;140(1618–1628):e1616.
Zhang L, Yang L, Liu X, Chen W, Chang L, Chen L, et al. MicroRNA-657 promotes tumorigenesis in hepatocellular carcinoma by targeting transducin-like enhancer protein 1 through nuclear factor kappa B pathways. Hepatology. 2013;57:1919–30.
Law PT, Qin H, Ching AK, Lai KP, Co NN, He M, et al. Deep sequencing of small RNA transcriptome reveals novel non-coding RNAs in hepatocellular carcinoma. J Hepatol. 2013;58:1165–73.
Wang Y, Lu Y, Toh ST, Sung WK, Tan P, Chow P, et al. Lethal-7 is down-regulated by the hepatitis B virus x protein and targets signal transducer and activator of transcription 3. J Hepatol. 2010;53:57–66.
Au SL, Wong CC, Lee JM, Fan DN, Tsang FH, Ng IO, et al. Enhancer of zeste homolog 2 epigenetically silences multiple tumor suppressor microRNAs to promote liver cancer metastasis. Hepatology. 2012;56:622–31.
Ji J, Zhao L, Budhu A, Forgues M, Jia HL, Qin LX, et al. Let-7g targets collagen type I alpha2 and inhibits cell migration in hepatocellular carcinoma. J Hepatol. 2010;52:690–7.
Fang Y, Xue JL, Shen Q, Chen J, Tian L. MicroRNA-7 inhibits tumor growth and metastasis by targeting the phosphoinositide 3-kinase/Akt pathway in hepatocellular carcinoma. Hepatology. 2012;55:1852–62.
Yan Y, Luo YC, Wan HY, Wang J, Zhang PP, Liu M, et al. MicroRNA-10a is involved in the metastatic process by regulating Eph tyrosine kinase receptor A4-mediated epithelial–mesenchymal transition and adhesion in hepatoma cells. Hepatology. 2013;57:667–77.
Maurel M, Jalvy S, Ladeiro Y, Combe C, Vachet L, Sagliocco F, et al. A functional screening identifies five microRNAs controlling glypican-3: role of miR-1271 down-regulation in hepatocellular carcinoma. Hepatology. 2013;57:195–204.
Wang Y, Jiang L, Ji X, Yang B, Zhang Y, Fu XD. Hepatitis B viral RNA directly mediates down-regulation of the tumor suppressor microRNA miR-15a/miR-16-1 in hepatocytes. J Biol Chem. 2013;288:18484–93.
Yang X, Liang L, Zhang XF, Jia HL, Qin Y, Zhu XC, et al. MicroRNA-26a suppresses tumor growth and metastasis of human hepatocellular carcinoma by targeting interleukin-6-Stat3 pathway. Hepatology. 2013;58:158–70.
Kota J, Chivukula RR, O’Donnell KA, Wentzel EA, Montgomery CL, Hwang HW, et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell. 2009;137:1005–17.
Xiong Y, Fang JH, Yun JP, Yang J, Zhang Y, Jia WH, et al. Effects of microRNA-29 on apoptosis, tumorigenicity, and prognosis of hepatocellular carcinoma. Hepatology. 2010;51:836–45.
Fang JH, Zhou HC, Zeng C, Yang J, Liu Y, Huang X, et al. MicroRNA-29b suppresses tumor angiogenesis, invasion, and metastasis by regulating matrix metalloproteinase 2 expression. Hepatology. 2011;54:1729–40.
Bae HJ, Noh JH, Kim JK, Eun JW, Jung KH, Kim MG, et al. MicroRNA-29c functions as a tumor suppressor by direct targeting oncogenic SIRT1 in hepatocellular carcinoma. Oncogene. 2013 (in press).
Yang P, Li QJ, Feng Y, Zhang Y, Markowitz GJ, Ning S, et al. TGF-β-miR-34a-CCL22 signaling-induced Treg cell recruitment promotes venous metastases of HBV-positive hepatocellular carcinoma. Cancer Cell. 2012;22:291–303.
Petrelli A, Perra A, Schernhuber K, Cargnelutti M, Salvi A, Migliore C, et al. Sequential analysis of multistage hepatocarcinogenesis reveals that miR-100 and PLK1 dysregulation is an early event maintained along tumor progression. Oncogene. 2012;31:4517–26.
Li D, Liu X, Lin L, Hou J, Li N, Wang C, et al. MicroRNA-99a inhibits hepatocellular carcinoma growth and correlates with prognosis of patients with hepatocellular carcinoma. J Biol Chem. 2011;286:36677–85.
Wang L, Zhang X, Jia LT, Hu SJ, Zhao J, Yang JD, et al. c-Myc-mediated epigenetic silencing of microRNA-101 contributes to dysregulation of multiple pathways in hepatocellular carcinoma. Hepatology. 2013 (in press).
Su H, Yang JR, Xu T, Huang J, Xu L, Yuan Y, et al. MicroRNA-101, down-regulated in hepatocellular carcinoma, promotes apoptosis and suppresses tumorigenicity. Cancer Res. 2009;69:1135–42.
Li S, Fu H, Wang Y, Tie Y, Xing R, Zhu J, et al. MicroRNA-101 regulates expression of the v-fos FBJ murine osteosarcoma viral oncogene homolog (FOS) oncogene in human hepatocellular carcinoma. Hepatology. 2009;49:1194–202.
Wang B, Hsu SH, Wang X, Kutay H, Bid HK, Yu J, et al. Reciprocal regulation of miR-122 and c-Myc in hepatocellular cancer: role of E2F1 and TFDP2. Hepatology. 2013 (in press).
Song K, Han C, Zhang J, Lu D, Dash S, Feitelson M, et al. Epigenetic regulation of MicroRNA-122 by peroxisome proliferator activated receptor-gamma and hepatitis b virus X protein in hepatocellular carcinoma cells. Hepatology. 2013 (in press).
Zeng C, Wang R, Li D, Lin XJ, Wei QK, Yuan Y, et al. A novel GSK-3 beta-C/EBP alpha-miR-122-insulin-like growth factor 1 receptor regulatory circuitry in human hepatocellular carcinoma. Hepatology. 2010;52:1702–12.
Gramantieri L, Ferracin M, Fornari F, Veronese A, Sabbioni S, Liu CG, et al. Cyclin G1 is a target of miR-122a, a microRNA frequently down-regulated in human hepatocellular carcinoma. Cancer Res. 2007;67:6092–9.
Zheng F, Liao YJ, Cai MY, Liu YH, Liu TH, Chen SP, et al. The putative tumour suppressor microRNA-124 modulates hepatocellular carcinoma cell aggressiveness by repressing ROCK2 and EZH2. Gut. 2012;61:278–89.
Furuta M, Kozaki KI, Tanaka S, Arii S, Imoto I, Inazawa J. miR-124 and miR-203 are epigenetically silenced tumor-suppressive microRNAs in hepatocellular carcinoma. Carcinogenesis. 2010;31:766–76.
Kim JK, Noh JH, Jung KH, Eun JW, Bae HJ, Kim MG, et al. Sirtuin7 oncogenic potential in human hepatocellular carcinoma and its regulation by the tumor suppressors MiR-125a-5p and MiR-125b. Hepatology. 2013;57:1055–67.
Fan DN, Tsang FH, Tam AH, Au SL, Wong CC, Wei L, et al. Histone lysine methyltransferase, suppressor of variegation 3–9 homolog 1, promotes hepatocellular carcinoma progression and is negatively regulated by microRNA-125b. Hepatology. 2013;57:637–47.
Gong J, Zhang JP, Li B, Zeng C, You K, Chen MX, et al. MicroRNA-125b promotes apoptosis by regulating the expression of Mcl-1, Bcl-w and IL-6R. Oncogene. 2013;32:3071–9.
Alpini G, Glaser SS, Zhang JP, Francis H, Han Y, Gong J, et al. Regulation of placenta growth factor by microRNA-125b in hepatocellular cancer. J Hepatol. 2011;55:1339–45.
Liang L, Wong CM, Ying Q, Fan DN, Huang S, Ding J, et al. MicroRNA-125b suppressesed human liver cancer cell proliferation and metastasis by directly targeting oncogene LIN28B2. Hepatology. 2010;52:1731–40.
Wong CC, Wong CM, Tung EK, Au SL, Lee JM, Poon RT, et al. The microRNA miR-139 suppresses metastasis and progression of hepatocellular carcinoma by down-regulating Rho-kinase 2. Gastroenterology. 2011;140:322–31.
Yang H, Fang F, Chang R, Yang L. MicroRNA-140-5p suppresses tumor growth and metastasis by targeting transforming growth factor β receptor 1 and fibroblast growth factor 9 in hepatocellular carcinoma. Hepatology. 2013;58:205–17.
Takata A, Otsuka M, Yoshikawa T, Kishikawa T, Hikiba Y, Obi S, et al. MicroRNA-140 acts as a liver tumor suppressor by controlling NF-κB activity by directly targeting DNA methyltransferase 1 (Dnmt1) expression. Hepatology. 2013;57:162–70.
Banaudha K, Kaliszewski M, Korolnek T, Florea L, Yeung ML, Jeang KT, et al. MicroRNA silencing of tumor suppressor DLC-1 promotes efficient hepatitis C virus replication in primary human hepatocytes. Hepatology. 2011;53:53–61.
Law PT, Ching AK, Chan AW, Wong QW, Wong CK, To KF, et al. MiR-145 modulates multiple components of the insulin-like growth factor pathway in hepatocellular carcinoma. Carcinogenesis. 2012;33:1134–41.
Gailhouste L, Gomez-Santos L, Hagiwara K, Hatada I, Kitagawa N, Kawaharada K, et al. miR-148a plays a pivotal role in the liver by promoting the hepatospecific phenotype and suppressing the invasiveness of transformed cells. Hepatology. 2013;58:1153–65.
Xu X, Fan Z, Kang L, Han J, Jiang C, Zheng X, et al. Hepatitis B virus X protein represses miRNA-148a to enhance tumorigenesis. J Clin Invest. 2013;123:630–45.
Zhang JP, Zeng C, Xu L, Gong J, Fang JH, Zhuang SM. MicroRNA-148a suppresses the epithelial–mesenchymal transition and metastasis of hepatoma cells by targeting Met/Snail signaling. Oncogene. 2013 (in press).
Han H, Sun D, Li W, Shen H, Zhu Y, Li C, et al. A c-Myc-MicroRNA functional feedback loop affects hepatocarcinogenesis. Hepatology. 2013;57:2378–89.
Huang J, Wang Y, Guo Y, Sun S. Down-regulated microRNA-152 induces aberrant DNA methylation in hepatitis B virus-related hepatocellular carcinoma by targeting DNA methyltransferase 1. Hepatology. 2010;52:60–70.
Ding J, Huang S, Wang Y, Tian Q, Zha R, Shi H, et al. Genome-wide screening reveals that miR-195 targets the TNF-α/NF-κB pathway by down-regulating IκB kinase alpha and TAB 3 in hepatocellular carcinoma. Hepatology. 2013;58:654–66.
Wang R, Zhao N, Li S, Fang JH, Chen MX, Yang J, et al. MicroRNA-195 suppresses angiogenesis and metastasis of hepatocellular carcinoma by inhibiting the expression of VEGF, VAV2, and CDC42. Hepatology. 2013;58:642–53.
Xu T, Zhu Y, Xiong Y, Ge YY, Yun JP, Zhuang SM. MicroRNA-195 suppresses tumorigenicity and regulates G1/S transition of human hepatocellular carcinoma cells. Hepatology. 2009;50:113–21.
Yuan JH, Yang F, Chen BF, Lu Z, Huo XS, Zhou WP, et al. The histone deacetylase 4/SP1/microrna-200a regulatory network contributes to aberrant histone acetylation in hepatocellular carcinoma. Hepatology. 2011;54:2025–35.
Shih TC, Tien YJ, Wen CJ, Yeh TS, Yu MC, Huang CH, et al. MicroRNA-214 downregulation contributes to tumor angiogenesis by inducing secretion of the hepatoma-derived growth factor in human hepatoma. J Hepatol. 2012;57:584–91.
Wong QW, Lung RW, Law PT, Lai PB, Chan KY, To KF, et al. MicroRNA-223 is commonly repressed in hepatocellular carcinoma and potentiates expression of Stathmin1. Gastroenterology. 2008;135:257–69.
Guichard C, Amaddeo G, Imbeaud S, Ladeiro Y, Pelletier L, Maad IB, et al. Integrated analysis of somatic mutations and focal copy-number changes identifies key genes and pathways in hepatocellular carcinoma. Nat Genet. 2012;44:694–8.
Chang Y, Yan W, He X, Zhang L, Li C, Huang H, et al. miR-375 inhibits autophagy and reduces viability of hepatocellular carcinoma cells under hypoxic conditions. Gastroenterology. 2012;143(177–187):e178.
He XX, Chang Y, Meng FY, Wang MY, Xie QH, Tang F, et al. MicroRNA-375 targets AEG-1 in hepatocellular carcinoma and suppresses liver cancer cell growth in vitro and in vivo. Oncogene. 2012;31:3357–69.
You X, Liu F, Zhang T, Li Y, Ye L, Zhang X. Hepatitis B virus X protein upregulates oncogene Rab18 to result in the dysregulation of lipogenesis and proliferation of hepatoma cells. Carcinogenesis. 2013;34:1644–52.
Buurman R, Gürlevik E, Schäffer V, Eilers M, Sandbothe M, Kreipe H, et al. Histone deacetylases activate hepatocyte growth factor signaling by repressing microRNA-449 in hepatocellular carcinoma cells. Gastroenterology. 2012;143:811–20. e811–15.
Tao ZH, Wan JL, Zeng LY, Xie L, Sun HC, Qin LX, et al. miR-612 suppresses the invasive-metastatic cascade in hepatocellular carcinoma. J Exp Med. 2013;210:789–803.
Zhang JF, He ML, Fu WM, Wang H, Chen LZ, Zhu X, et al. Primate-specific microRNA-637 inhibits tumorigenesis in hepatocellular carcinoma by disrupting signal transducer and activator of transcription 3 signaling. Hepatology. 2011;54:2137–48.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Otsuka, M., Kishikawa, T., Yoshikawa, T. et al. The role of microRNAs in hepatocarcinogenesis: current knowledge and future prospects. J Gastroenterol 49, 173–184 (2014). https://doi.org/10.1007/s00535-013-0909-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-013-0909-8