Abstract
Monoclonal antibodies are important tools for various applications in hantavirus diagnostics. Recently, we generated Puumala virus (PUUV)-reactive monoclonal antibodies (mAbs) by immunisation of mice with chimeric polyomavirus-derived virus-like particles (VLPs) harbouring the 120-amino-acid-long amino-terminal region of the PUUV nucleocapsid (N) protein. Here, we describe the generation of two mAbs by co-immunisation of mice with hexahistidine-tagged full-length N proteins of Sin Nombre virus (SNV) and Andes virus (ANDV), their characterization by different immunoassays and comparison with the previously generated mAbs raised against a segment of PUUV N protein inserted into VLPs. All of the mAbs reacted strongly in ELISA and western blot tests with the antigens used for immunization and cross-reacted to varying extents with N proteins of other hantaviruses. All mAbs raised against a segment of the PUUV N protein presented on chimeric VLPs and both mAbs raised against the full-length AND/SNV N protein reacted with Vero cells infected with different hantaviruses. The reactivity of mAbs with native viral nucleocapsids was also confirmed by their reactivity in immunohistochemistry assays with kidney tissue specimens from experimentally SNV-infected rodents and human heart tissue specimens from hantavirus cardiopulmonary syndrome patients. Therefore, the described mAbs represent useful tools for the immunodetection of hantavirus infection.
Similar content being viewed by others
Introduction
Hantaviruses, genus Hantavirus, family Bunyaviridae, are enveloped viruses with a segmented RNA genome of negative polarity [55]. The small (S) genome segment encodes the nucleocapsid (N) protein, which is associated with the RNA genome. The medium (M) genome segment codes for a glycoprotein precursor that is cleaved into the proteins G1 and G2, which are incorporated into the viral envelope. The large (L) genome segment contains the coding information for the RNA-dependent RNA polymerase responsible for virus transcription and replication [67].
In general, wild living rodents of the families Muridae and Cricetidae are the hantavirus reservoirs and thereby define the geographical distribution of the different hantavirus species. To this end, each hantavirus species is closely associated with a single rodent species or at least a closely related species of the same genus. According to this host specificity and other criteria of the International Committee on Taxonomy of Viruses, 22 different hantavirus species have been defined so far. Hantaan virus, the prototype virus of the genus, has been isolated from a striped field mouse (Apodemus agrarius, family Muridae, subfamily Murinae) trapped in Korea [32]. This hantavirus and Seoul virus, another Asian hantavirus transmitted by rats (Rattus rattus, R. norvegicus, R. losea; subfamily Murinae), can cause hemorrhagic fever with renal syndrome (HFRS) in humans, with case fatality ratios of 5-10% and 1-2%, respectively [47]. Milder courses of HFRS in Europe with a case fatality ratio of <1% are caused by Puumala virus (PUUV) carried by bank voles (Myodes glareolus, formerly Clethrionomys glareolus, subfamily Arvicolinae) [6]. Tula virus (TULV) represents another species of Arvicolinae-associated hantavirus that so far have only rarely been detected by serological assays as causative agents of human infections [25, 71, 79]. In contrast, human infections by Dobrava-Belgrade virus strains are well documented for several European countries, causing up to 12% case fatalities [2, 5, 43, 44, 48, 59, 65, 72]. This virus species includes three distinct genetic lineages associated with the striped field mouse (Apodemus agrarius), yellow-necked mouse (A. flavicollis) and the Caucasian wood mouse (A. ponticus), which most likely differ in their pathogenicity to humans [5, 27, 53, 72]. In 1993, a new disease, later designated as “hantavirus cardiopulmonary syndrome” (HCPS), was discovered in the Four Corners region, located in the southwestern United States [54]. This disease, characterized by pulmonary dysfunction, has a high case fatality ratio of about 36% [8]. The most important causative agents of HCPS in North and South America are Sin Nombre virus (SNV), carried by the deer mouse (Peromyscus maniculatus, subfamily Sigmodontinae), and Andes virus (ANDV), carried by the long-tailed pygmy rice rat Oligoryzomys longicaudatus (subfamily Sigmodontinae), respectively [66]. In addition to the rodent-borne hantaviruses, a large number of novel shrew- and mole-associated hantaviruses have been discovered recently [7, 23, 24, 26].
The diagnostics of human hantavirus infections is mainly based on serological assays. Hantavirus infections have been detected by indirect immunofluorescence assays (IFA), hemagglutination inhibition test, immune adherence hemagglutination test and high-density particle agglutination and different ELISA formats based on viral antigens [56]. More recently ELISA, immunoblot (western blot and strip blot) and rapid immunochromatographic assays have been developed using recombinant N protein antigens [20, 21, 31, 50]. Due to the strong cross-reactivity between N proteins of members of closely related hantavirus species, the use of N proteins of only selected hantaviruses of the Murinae, Arvicolinae and Sigmodontinae/Neotominae groups have been suggested to be sufficient for serological diagnostics [31]. However, in certain cases, a “homologous” antigen that is closely related to that of the infecting hantavirus might be required for a highly sensitive diagnosis in the acute phase of infection [12, 58, 68, 70, 74].
Monoclonal antibodies (mAbs) have widely been used as tools for diagnostic purposes, e.g., for detection of virus-specific antibodies in humans by different capture ELISA formats [37, 49, 84] or of viral antigen in infected cell culture or rodent tissues [39]. A comparative study using different recombinant N antigens and IgG ELISA formats has suggested the superior performance of the capture ELISA format using native viral antigens or baculovirus-expressed whole N antigens, in terms of specificity and sensitivity [73].
Recently, we developed a platform based on hamster polyomavirus major capsid protein (VP1)-derived chimeric virus-like particles (VLPs) for the generation of mAbs with the desired specificity. Immunisation of mice with VLPs harbouring the amino-terminal 120 amino acids (aa) of PUUV N protein at certain surface-exposed sites of VP1 resulted in the induction of a high-titered N-protein-specific antibody response [14]. Using these VLPs and a similar immunisation scheme, hantavirus N-protein-specific mAbs were generated [85].
The objective of the present study was to raise SNV- and ANDV-specific mAbs, to characterize these mAbs and compare them to those raised previously against a PUUV N protein segment presented on chimeric VLPs in terms of their cross-reactivity and reactivity with native viral antigens.
Materials and methods
Recombinant antigens
For immunisation of mice, yeast-expressed His-tagged N proteins of SNV strain 3H226 and ANDV strain AH-1 were used [68, 70]. In addition to these two recombinant proteins, different recombinant N proteins were used for cross-reactivity studies. For ELISA and western blot tests, yeast-expressed amino-terminally His-tagged N proteins of PUUV strains Vranica/Hällnäs (PUUV-Vra), Kazan (PUUV-Kaz) and Sotkamo (PUUV-Sot), HTNV strain Fojnica (HTNV), DOBV strain Slovenia (DOBV-Slo) [10, 62], SEOV [69], and TULV strain Moravia (Mertens et al., unpublished data) were used. The N proteins of PUUV strains Sotkamo and Kazan contain the entire sequence, whereas all others are lacking the amino-terminal methionine. For cross-reactivity testing of mouse antisera, yeast-expressed amino-terminally His-tagged N proteins of rabies virus (RABV) and human parainfluenza virus type 3 (hPIV3) were used (Petraityte et al., unpublished data). All of these recombinant proteins were expressed in yeast Saccharomyces cerevisiae strain gcn2 and purified by nickel chelation chromatography under denaturing conditions according to a protocol described previously [62]. For strip immunoassay (SIA), E. coli-expressed carboxy-terminally His-tagged N proteins were generated using the pET21 system (T7 system/E. coli; EMD Chemicals-Novagen, Gibbstown, NJ, USA). In each case, the N gene of ANDV, SNV, Muleshoe virus (MULEV), Bayou virus (BAYV), Rio Mamore virus (RMV), PUUV strain P360, Prospect Hill virus (PHV) and SEOV was amplified from animal tissues, virus-infected cells or from a pre-existing plasmid clone and inserted into the pET21b backbone using, usually, the HindIII and XhoI cloning sites. Two N proteins, those from ANDV and from SEOV, were slightly truncated, by 4 and 5 amino acids, respectively, at the carboxy terminus. The remaining N proteins were full-length. In all cases, the N proteins were expressed from the pET21b backbone in BL21 (DE3) cells using IPTG for induction. The proteins were affinity-purified using nickel chelation columns (Qiagen, Valencia, CA, USA).
Generation of monoclonal antibodies
The generation of mAbs against non-His-tagged chimeric polyomavirus-derived VLPs harbouring the 120-aa-long segment of PUUV-Vra N protein (clones 7A5, 5C5, 5E11 and 2C6; see Table 1) has been described recently [85]. The mAbs against SNV/ANDV N proteins were generated by immunization of BALB/c mice (n=3) subcutaneously with 50 μg of either ANDV or SNV His-tagged N proteins according to the following scheme: the first and the third immunizations were done with ANDV N protein; the second and the final boost were done with SNV N protein. These immunizations were administered with 28-day intervals between each. For the primary immunization, the antigen was emulsified in complete Freund’s adjuvant (Sigma Aldrich, St. Louis, MO, USA), and the subsequent immunizations were performed without any adjuvant. All procedures involving experimental mice were performed at the breeding colony of the Institute of Immunology (Vilnius University, Lithuania) under controlled laboratory conditions in strict accordance with Lithuanian and European legislation. The approval to use BALB/c mice for immunization experiments and mAb production was obtained from the Lithuanian State Food and Veterinary Agency (No. 0127, valid through 31.12.2010). Hybridomas were generated essentially as described by Kohler and Milstein (1975) [29]. Four days after the last immunization (corresponding to day 88 from the initial immunization), spleen cells of the best responder mouse were fused with mouse myeloma Sp2/0 cells using PEG 1500 as a fusion agent (PEG/DMSO solution, HybriMax, Sigma). Hybrid cells were selected in growth medium supplemented with hypoxanthine, aminopterin and thymidine (50x HAT media supplement, Sigma). Viable clones were screened by indirect ELISA using 96-well microtiter plates coated with the recombinant SNV and ANDV N proteins. Positive clones were stabilized by limiting dilution cloning on a macrophage feeder layer. Hybridoma cells were maintained in complete Dulbecco’s modified Eagle’s medium (DMEM) containing 15% fetal calf serum (FCS, Biochrom) and antibiotics. Heavy chain types of mAbs were determined by ELISA using a Monoclonal Antibody Isotyping Kit (Sigma, ISO-2). Specificity of mAbs was tested by ELISA and western blotting as described below.
Indirect ELISA
The specificity of mAbs was tested by an indirect ELISA. Microtiter plates (Nunc MaxiSorp, Roskilde, Denmark) were coated with the indicated antigens by adding 100 μl of the antigen solution (2 μg/ml) in coating buffer (50 mM Na-carbonate, pH 9.5) and incubation overnight at +4°C. The plates were blocked for 30 min at RT with 1% BSA in PBS and then incubated with undiluted hybridoma supernatants for 1 h at RT. After washing, the plates were incubated with horseradish peroxidase (HRP)-labeled anti-mouse IgG (Bio-Rad, 1:5000 in PBS-T) for 1 h at RT. Supernatants containing antigen-specific antibodies were detected with TMB substrate (Sigma), and the optical density (OD) was measured at 450 nm in a microtiter plate reader (Tecan, Groedig, Austria).
SDS-polyacrylamide gel electrophoresis and western blot analysis
Purified proteins (100 ng/lane) or crude cell lysates (1 μg of total protein per lane) were fractionated by sodium dodecylsulfate polyacrylamide gel electrophoresis (SDS-PAGE) in 12% minigels and transferred to polyvinyldifluoride (PVDF) membranes (Millipore, Bedford, USA) under semidry conditions. Membranes were blocked with 1% gelatin in PBS for 1 h at RT, washed with PBS containing 0.1% Tween-20 (PBS-T) and incubated with hybridoma supernatants for 1 h at RT. After washing, membranes were incubated with HRP-labelled anti-mouse IgG (Bio-Rad, 1:5000) for 1 h at RT. Blots were visualized using TMB-blotting (3’,5,5’-tetramethylbenzidine) ready-to-use substrate (Sigma).
Strip immunoblot assay
Affinity-purified recombinant N proteins were loaded at 2.5 ng/lane onto nitrocellulose membranes and applied to the membrane by suction. Control lanes included a Coomassie blue lane, which specified the orientation of the strip, and one or more lanes devoted to human or murine IgG or IgM to help the user determine that the anti-immunoglobulin conjugate has been correctly deployed in the assay. The intensity of mAb reactivity after probing the membrane was determined visually as positive or negative as compared to the anti-immunoglobulin controls. Hybridoma supernatants were used at dilutions of 1:500 and 1:2000. The results were scored visually as positive or negative in the context of the corresponding appropriate known positive or negative serum controls.
Indirect immunofluorescence assay
Indirect immunofluorescence assay (IFA) was done using commercial IFA slides with PUUV (strain Hällnäs B1 83-223L)-, HTNV (strain 76-118)- and SEOV (strain R22)-infected Vero E6 cells (Progen Biotechnik, Heidelberg, Germany) and PUUV (strain Sotkamo)-, DOBV (strain Slovenia)-, Saaremaa virus-, SEOV-, HTNV- and SNV-infected Vero E6 cells (EuroImmun, Lübeck, Germany) as recommended by the manufacturers. Briefly, cell culture supernatants of hybridomas were added to the slides in a dilution of 1:50 in PBS or undiluted. The negative control mAbs directed against Neospora caninum surface proteins were used at the same dilutions. Detection was done by incubation with a fluorescein isothiocyanate (FITC)-conjugated anti-mouse immunoglobulin (H+L) goat IgG (DakoCytomation, Hamburg, Germany) diluted 1:20 in PBS. Cells were counterstained with Evans blue and IFA slides were analysed using a Olympus VANOX AHBT 3 fluorescence microscope (Olympus Optical Co. Europa GmbH, Hamburg) with a 40x objective at an excitation of 495 nm and analySIS 3.2 software.
In addition to the commercial IFA, an in-house IFA using ANDV-infected Vero E6 cells cultivated in LabTek 16-well chamber slides (Nunc) was used. Cells were infected with ANDV stock, and after 7 days, they were dispersed by trypsinization, washed with PBS, fixed with acetone and incubated with the mAbs. After washing three times, FITC-labelled polyclonal rabbit anti-mouse F(ab′)2 (DakoCytomation) was added. Lung and heart tissues from seropositive rodents were also employed in the IFA technique to evaluate the reactivity of mAbs. Tissues were rinsed twice with PBS, and cells were attached to cover slips and stained as above. Preliminary experiments were done to select the optimal mAb concentration. The mAbs and the conjugate were diluted in PBS, and Evans blue (1:30000) was added to the conjugate dilution.
Immunohistochemistry of human and deer mouse tissue
Immunohistochemistry (IHC) was performed as described previously [19]. Tissue samples were fixed in 10% neutral-buffered formalin for 24-48 h before they were embedded in paraffin and subsequently sectioned at 4-6 μm thickness. The sections were placed onto (+)-charged poly-L-lysine microscope slides, subjected to antigen retrieval, deparaffinized and exposed to a 1:10,000 dilution in PBS of hyperimmune rabbit antiserum raised against recombinant SNV N protein or the indicated mAb diluted to 1:50 on an automated platform (Ventana Medical Systems Inc., Tucson, AZ, USA) for 1 h as described previously [19]. As a negative control, mAb against human influenza hemagglutinin (HA) at a 1:100 dilution was used (clone HA.C5, Abcam, Cambridge, UK). After antibody binding and several washes in PBS, the antibody-protein complexes were detected initially with a biotinylated goat anti-rabbit or goat anti-mouse secondary antibody applied for 1 h, followed by an HRP-avidin conjugate (Jackson ImmunoResearch Laboratories, Inc., West Grove, PA, USA). After further washes, the immune complexes were visualized with Fast Red substrate (Sigma, St. Louis, MO, USA), coverslipped, and analyzed by light microscopy. Specific staining appeared as red, punctate, cytoplasmic granules. Images were taken with the 60x magnification objective.
Results
Generation of mAbs against recombinant N proteins of ANDV and SNV
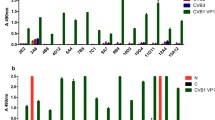
Subsequent immunizations with recombinant N proteins of ANDV and SNV elicited a strong N protein-specific antibody response in immunized mice. After the final immunization, the titers of IgG antibodies against SNV and ANDV N proteins in the sera of immunized mice were in the range of 1:70,000 - 1:100,000 (Fig. 1A). The specificity of the sera was confirmed by the fact that they did not show any reactivity with irrelevant yeast-expressed amino-terminally His-tagged N proteins of RABV and hPIV3 used as a negative control (Fig. 1A). Antisera were cross-reactive to varying extents with recombinant N proteins of other hantaviruses: they showed moderate to high reactivity to PUUV and HTNV N proteins (Fig. 1A). When compared to the cross-reactivity of antisera elicited by immunization with a PUUV N protein segment presented on chimeric VLPs [85], the immunization with full-length SNV/ANDV N proteins resulted in a higher level of cross-reactivity of mouse antisera. Nevertheless, immunization with chimeric VLPs harbouring a fragment of the PUUV N protein elicited a considerable level of cross-reactive antibodies against N proteins of SNV, ANDV and the different PUUV strains (Fig. 1B). The pattern of cross-reactivity of the antisera allowed us to conclude that immunization with recombinant antigens elicited the production of broadly cross-reactive antibodies.
Immunoreactivity of antisera collected from mice immunized either with full-length His-tagged SNV/ANDV N proteins (A) or non-His-tagged chimeric VLPs harbouring a 120-aa-long PUUV N protein segment (B) compared to recombinant N proteins of Sin Nombre virus (SNV), Andes virus (ANDV), Puumala virus strains Kazan (PUUV-Kaz), Sotkamo (PUUV-Sot) and Vranica-Hällnäs (PUUV-Vra), Hantaan virus (HTNV) and Dobrava-Belgrade virus, strain Slovenia (DOBV-Slo). As a negative control, yeast-expressed His-tagged N proteins of rabies virus (RABV) and human parainfluenza virus type 3 (hPIV3) were used
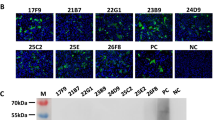
Fusion of spleen cells of the SNV/ANDV N-protein-immunized mouse with mouse myeloma cells resulted in two stable hybridoma clones (4H3 and 7G2; see Table 1). Typing by ELISA revealed a IgG1 subclass for both mAbs produced. The specificity of their immunoreactivities was confirmed by western blot analysis using the homologous recombinant SNV and ANDV N proteins used for immunization (see Fig. 2 and Table 3).
Western blot analysis of the cross-reactivity of the mAbs 4H3 (B) and 7G2 (C) with N proteins of Sin Nombre virus (lane 1), Andes virus (lane 2), Puumala virus (PUUV), strain Kazan (lane 3), PUUV, strain Sotkamo (lane 4), PUUV, strain Vranica/Hällnäs (lane 5), Tula virus (lane 6), Hantaan virus (lane 7), Dobrava-Belgrade virus, strain Slovenia (lane 8), and Seoul virus (lane 9). Lane 10, crude lysate of non-transformed yeast cells. Lane M, PageRulerTM Prestained Protein Ladder (UAB Fermentas, Vilnius, Lithuania, #SM0671). A, as a control, the same yeast cell crude lysates were run in a SDS polyacrylamide gel and stained with Coomassie blue
Cross-reactivities of mAbs raised against SNV/ANDV N protein with N proteins of different hantaviruses
The cross-reactivities of the novel mAbs 4H3 and 7G2 were analyzed in parallel by ELISA and western blot analysis using N proteins of SNV, ANDV, different PUUV strains, TULV, HTNV, DOBV-Slo and SEOV (Fig. 2, Tables 2 and 3) and by strip immunoblot assay (SIA) using N proteins of SNV, ANDV, BAYV, MULEV, RMV, PUUV and PHV and SEOV (Table 3). The mAb 4H3 reacted in ELISA and western blot exclusively with the SNV and ANDV N proteins used for immunization (Fig. 2B, Tables 2 and 3). In contrast, the mAb 7G2 demonstrated in the ELISA a broad cross-reactivity with the SNV and ANDV N proteins used for immunization as well as the N proteins of Arvicolinae-associated PUUV and TULV and Murinae-associated HTNV and SEOV (Table 2). This reactivity pattern, as well as the non-reactivity of this mAb with DOBV N antigen, was confirmed by western blot analysis (Fig. 2C, Table 3). The SIA analysis of mAb 7G2 confirmed the western blot reactivity pattern and demonstrated additional reactivities of this mAb with the N proteins of BAYV, MULEV, RMV and PHV (Table 3). Similarly, the mAb 4H3, which was reactive in the Western blot exclusively with SNV and ANDV N antigens, was found to react in SIA with N proteins of New World hantaviruses (BAYV, MULEV, RMV; Tables 2 and 3).
Cross-reactivities of mAbs raised against a PUUV N protein segment with N proteins of different hantaviruses
In addition to the previously reported cross-reactivities of mAbs 7A5, 5C5, 5E11 and 2C6 in western blot with N antigens of three different PUUV strains (see Table 3), we further evaluated the cross-reactivity characteristics of these mAbs with other antigens by ELISA, western blot and SIA analysis (Tables 2 and 3). In line with previous findings, the mAb 2C6 was found to react in the ELISA and western blot test exclusively with the N protein of PUUV strain Vranica/Hällnäs (Tables 2 and 3). Interestingly, this mAb reacted in the SIA with the N protein of PUUV strain P360, suggesting its antigenic relationship to the Vranica-Hällnäs strain (Table 3). The mAbs 5C5 and 5E11 reacted with N proteins of almost all Arvicolinae and New World rodent-associated hantaviruses (except MULEV), but only with one or two of the three analysed N antigens of Murinae-associated viruses (HTNV, DOBV and SEOV; Tables 2 and 3). The mAb 7A5 showed a similar reactivity pattern, but with a much lower intensity in SIA and with no reactivity in ELISA and western blot tests to the N proteins of HTNV, DOBV and SEOV that were tested (Tables 2 and 3).
Reactivities of mAbs with Vero cells infected with different hantaviruses in the immunofluorescence assay
The reactivities of all six mAbs with hantavirus-infected cells were tested by different commercial and an in-house immunofluorescence assays (IFA). All mAbs reacted with at least one antigen in the IFA, demonstrating that the mAbs are able to detect viral antigen in a native conformation (Table 4). In general, the results obtained correlate well with data from ELISA, western blot and SIA analysis. In addition, the reactivity patterns of the mAbs using two different commercially available slides (from Progen and Euroimmun) were similar. The mAb 4H3 was reactive with SNV and ANDV antigens, but not with those from PUUV, HTNV, SEOV, DOBV and Saaremaa virus (Table 4). The mAb 7G2 demonstrated a broad cross-reactivity with different hantaviruses, whereas mAb 2C6 was found to react with PUUV exclusively. The mAb 2C6 showed a slightly different immunoreactivity pattern from that observed in western blot: it was reactive with PUUV strains other than Vranica/Hällnäs and P360 (Fig. 3, Table 4). The other three mAbs raised against the PUUV N protein segment (clones 5C5, 5E11 and 7A5) showed a broader cross-reactivity with different hantaviruses (Fig. 3, Table 4).
Immunofluorescence assay with commercial slides based on Puumala virus, strain Sotkamo, with mAbs raised against SNV/ANDV N protein (clones 4H3, 7G2) and PUUV N protein segment (clones 2C6, 5C5, 5E11, 7A5). As a negative control, the same slides were incubated with the mAb specific for Neospora caninum surface proteins (NC)
In addition, the mAbs 5E11 and 4H3 were investigated for their reactivity with lung and heart tissues from 11 naturally hantavirus-infected rodents from Argentina that were confirmed by positive reverse transcriptase polymerase chain reaction (RT-PCR) and detection of anti-ANDV-IgG to contain ANDV of the South lineage. These included tissue specimens from 8 Oligoryzomys longicaudatus, 2 Abrothrix olivaceus and 1 A. longipilis captured in southern Argentina. One O. nigripes rodent captured in northern Argentina was confirmed to be infected with Juquitiba virus. Lung and heart tissue specimens from all 12 animals reacted with mAb 4H3. The reactivity of the mAb 5E11 was slightly lower. This antibody was not reactive with one O. longicaudatus tissue specimen (Table 5).
Reactivities of selected mAbs with tissue samples from SNV-infected mice and HCPS patients
To analyse the immunohistochemistry reactivity with tissue samples from hantavirus-infected mice and humans, we selected two mAbs that demonstrated strong reactivities by ELISA, western blot test, SIA and IFA. These mAbs were (i) mAb 5E11, raised against a PUUV N protein segment; and (ii) mAb 4H3, raised against full-length N protein of SNV/ANDV. To evaluate the immunodetection of viral antigen in situ, a known positive human heart tissue obtained from a subject who had succumbed to HCPS due to SNV infection, as well as deer mouse kidney tissue at day 14 after experimental infection with SNV were used. We found that both mAbs reacted strongly against SNV-positive tissue (Fig. 4A, B, E, F) and did not show any cross-reactivity on the non-infected tissue specimens used as a negative control (data not shown). The detection levels of SNV in human and mouse tissues were similar to those of a control hyperimmune rabbit serum raised against the SNV N protein (Fig. 4C, G). The irrelevant mAb used as a negative control did not show any reactivity with SNV-infected tissue specimens (Fig. 4D, H). This confirms the specificity of the mAb-based IHC test.
Immunohistochemistry detection of N protein in SNV-infected human and rodent tissue samples using mAbs 4H3 and 5E11. Photomicrographs are representative IHC stains for SNV N antigen of positive human heart tissues obtained post-mortem from an individual who succumbed to HCPS (A-D) and deer mouse kidney tissues at day 14 after experimental infection (E-H). Reactivities of the mAbs 4H3 (A, E) and 5E11 (B, F) are shown at 1:50 dilution of hybridoma supernatant. Control positive staining shown for both tissue types were done using SNV-N hyperimmune rabbit serum at 1:10,000 dilution (C, G). Control negative staining of infected tissue specimens were done using an irrelevant mAb against HA at a 1:100 dilution (D, H). Positive N-antigen expression in cells is seen as reddish, punctate staining. Arrows highlight representative positive stains as seen at 60x magnification
Discussion
The generation of hantavirus-specific mAbs followed different approaches. In the first attempts, HTNV-infected lung tissue from A. agrarius coreae or non-inactivated HTNV and SEOV were used to inoculate BALB/c mice, which resulted in the generation of mAbs with different properties [13, 75, 82]. Alternatively, complete but inactivated HTNV, SEOV and PUUV were used for immunization and subsequent generation of mAbs [4, 63]. A third approach was based on the infection of bank voles, the natural rodent host, with infectious PUUV, resulting in the generation of N-protein- and glycoprotein-specific mAbs [35, 36]. A fourth approach to generate hantavirus-specific antibodies explored materials from HFRS patients [38] in combination with the use of phage display techniques [28, 33, 34, 64]. Finally, recombinant N proteins were exploited for the generation of mAbs against HTNV, PUUV and TULV in laboratory mice [42, 46, 83, 84]. In a previous paper, we demonstrated that yeast-expressed polyomavirus-derived VLPs harbouring a PUUV N segment allowed the generation of specific mAbs in mice [85]. As we have previously demonstrated a strong immunogenicity of yeast-expressed His-tagged whole hantavirus N proteins in rabbits and mice [15, 62], we continued this approach by proving the potential of His-tagged yeast-expressed full-length N proteins of SNV/ANDV to induce a strong antibody response in mice and subsequently generate SNV/ANDV-reactive mAbs.
The broad analysis of mAbs raised against the amino-terminal segment (aa 1-120) of the PUUV N protein and those raised against the entire SNV/ANDV N proteins with N proteins of 14 different hantavirus strains, including four different PUUV strains, resulted in the identification of different cross-reactivity patterns. For both groups of mAbs, highly cross-reactive ones were identified reacting with almost all of the N proteins. The mAb 7G2, raised against full-length SNV/ANDV N protein, and the mAb 5E11, raised against a PUUV N protein segment, reacted with N proteins of SNV/ANDV, Arvicolinae- and Murinae-associated hantaviruses. This broad cross-reactivity pattern reflects the fact that the N protein is the most highly conserved hantaviral antigen [1]. Other mAbs demonstrated a lower level of cross-reactivity. The mAb 4H3, raised against full-length SNV/ANDV N protein, detected only the ANDV/SNV antigens used for immunisation and N antigens of other closely related Sigmodontinae/Neotominae-associated hantaviruses. Similarly, the mAbs 5C5 and 7A5, raised against a PUUV N protein segment, reacted almost exclusively with the N proteins of Sigmodontinae-, Neotominae- and Arvicolinae-associated hantaviruses. This pattern of cross-reactivity reflects the close phylogenetic and resulting antigenic similarity of these viruses [9]. The different cross-reactivity pattern is represented by mAb 2C6, which reacted in the ELISA, western blot and strip blot assay exclusively with the N proteins of PUUV strains Vranica/Hällnäs and P360, but not with the N proteins of other PUUV strains or with the N proteins of other hantaviruses. This is highly surprising, as the aa sequence divergence between N proteins of different PUUV strains reached only 5% at maximum [1, 9].
Previous investigations of the immune response in natural infection as well as experimental infection or immunisation demonstrated that the the amino-terminal region of the hantavirus N protein is highly immunogenic and cross-reactive [12, 16–18, 22, 41, 81]. In addition, the mAbs raised against the N proteins of PUUV, HTNV and TULV were found to bind to the amino-terminal region of the corresponding N proteins [41, 42, 83]. In line with these data, our mAbs 5C5, 5E11 and 7A5, which mapped to aa 1-45 or aa 1-120 of the N protein, were found to be cross-reactive to N proteins of related hantaviruses. However, the differences of the cross-reactivity patterns of mAbs 5C5 and 5E11 suggested a slightly different epitope localisation of these two mAbs. In line with these data, a previous investigation also identified different epitopes in the amino-terminal region (aa 1-45) of the PUUV N protein [45]. The exclusive reactivity of mAb 2C6 with the N proteins of PUUV strains Vranica/Hällnäs and P360 can be attributed to aa position 35, where only these two strains showed a D35Y aa exchange (Fig. 5). Interestingly, an aa exchange exactly at this position was shown previously to abrogate binding activity of the mAb 5A3 to the amino-terminal region of N protein of PUUV strain Vranica/Hällnäs while maintaining its reactivity to the corresponding region of the CG18-20 N protein [45]. In addition to the amino-terminal region, further B-cell epitopes have been identified along the N protein when human or animal sera or murine mAbs were studied [40, 41, 57, 76, 78, 83]. Previously, a region with a high degree of sequence variation has been identified in the hantavirus N protein between residues 241 and 300 [1, 60, 61], where a binding site for TULV-specific mAbs has been mapped between aa residues 226-293 [42]. Similarly, studies on the cross-reactivity of SNV- and ANDV-specific mAbs with different hantavirus N proteins have indicated that epitopes located within aa 244-286 are related to serotype specificity [77]. The lack of reactivity observed here for the PUUV-cross-reactive mAb 7G2 with the amino-terminal region (aa 1-120) of the PUUV N protein suggests that the epitope may lie downstream from this sequence.
Multiple amino acid (aa) sequence alignment of the amino-terminal region (aa 1-120) of the N proteins of Puumala virus strains Kazan (PUUV-Kaz, GenBank accession No. Z84204), P-360 (PUUV-P360, L11347), Sotkamo (PUUV-Sot, X61035), Vranica/Hällnäs (PUUV-Vra, U14137), Tula virus, strain Moravia (TULV, Z69991), Prospect Hill virus (PHV, M34011), Bayou virus (BAYV, L36929), Muleshoe virus (MULEV, U54575), Sin Nombre virus (SNV, NC_005216), Andes virus (ANDV, AF004660), Rio Mamore virus (RMV, U52136), Hantaan virus, strain Fojnica (HTNV, M14626), Seoul virus (SEOV, AY273791) and Dobrava-Belgrade virus, strain Slovenia (DOBV-Slo, L41916). The alignment was generated using ClustalW software. “*” indicates identical aa residues in all sequences in the alignment, “:” indicates conserved substitutions, and “.” indicates semi-conserved substitutions in the alignment
The observed cross-reactivity pattern of different mAbs and their epitope localization confirms the presence of multiple cross-reactive epitopes and few serotype-specific epitopes in the N protein of different hantaviruses. Therefore, the type-specific diagnosis of closely related hantavirus infections is complicated by the cross-reactivity of antibodies generated against the immunodominant amino-terminal N protein segment. This assumption was the basis for previous approaches to generate amino-terminally truncated N protein derivatives and their use for serotyping of hantavirus infections [3, 30, 51, 52, 80].
The mAbs characterized here were demonstrated to react with native viral antigen in infected Vero E6 cells in the indirect IFA as well as IHC with native viral antigen in tissue samples of naturally infected rodent reservoirs and human patients. This observation indicates that the recombinant antigens, both the N protein derivatives presented on polyomavirus-derived VLPs and the His-tagged N proteins, share linear and accessible epitopes with the native viral antigen in hantavirus-infected cells. The different reactivity patterns of the mAbs that were observed in the IFA may also allow their use for strain typing or antigenic characterization of different hantavirus strains, as already demonstrated previously for other mAbs [11, 35, 42, 75]. Interestingly, the strain-specific mAb 2C6 demonstrated a slightly increased cross-reactivity in the IFA with N proteins of PUUV strains other than Vranica/Hällnäs and P360 than in the western blot, suggesting an influence of the protein folding or a discontinuous structure of the antibody binding site in the N protein. In line with this assumption, a previous investigation also suggested a discontinuous nature of the epitopes of PUUV N-specific antibodies [45].
In conclusion, these novel mAbs represent useful tools for future immunohistochemistry-based pathogenicity studies in rodent reservoirs as well as in human samples, allowing a comparative analysis of the antigen expression in different tissue and cell types. In addition, the antibodies can be used for establishment of novel serological assays in hantavirus diagnostics, including antigenic classification of hantavirus strains.
References
Antic D, Kang CY, Spik K, Schmaljohn C, Vapalahti O, Vaheri A (1992) Comparison of the deduced gene products of the L, M and S genome segments of hantaviruses. Virus Res 24(1):35–46
Antoniadis A, Stylianakis A, Papa A, Alexiou-Daniel S, Lampropoulos A, Nichol ST, Peters CJ, Spiropoulou CF (1996) Direct genetic detection of Dobrava virus in Greek and Albanian patients with hemorrhagic fever with renal syndrome. J Infect Dis 174:407–410
Araki K, Yoshimatsu K, Ogino M, Ebihara H, Lundkvist A, Kariwa H, Takashima I, Arikawa J (2001) Truncated hantavirus nucleocapsid proteins for serotyping Hantaan, Seoul, and Dobrava hantavirus infections. J Clin Microbiol 39:2397–2404
Arikawa J, Schmaljohn AL, Dalrymple JM, Schmaljohn CS (1989) Characterization of Hantaan virus envelope glycoprotein antigenic determinants defined by monoclonal antibodies. J Gen Virol l70(Pt 3):615–624
Avsic-Zupanc TS, Xiao SY, Stojanovic R, Gligic A, van der Groen G, LeDuc JW (1992) Characterization of Dobrava Virus: a hantavirus from Slovenia, Yugoslavia. J Med Virol 38:132–137
Brummer-Korvenkontio M, Henttonen H, Vaheri A (1982) Hemorrhagic fever with renal syndrome in Finland: ecology and virology of nephropathia epidemica. Scand J Infect Dis Suppl 36:88–91
Carey DE, Reuben R, Panicker KN, Shope RE, Myers RM (1971) Thottapalayam virus: a presumptive arbovirus isolated from a shrew in India. Indian J Med Res 59:1758–1760
CDC hantaviruses (2010). http://www.cdc.gov/ncidod/diseases/hanta/hps/noframes/caseinfo.htm
Chu YK, Rossi C, Leduc JW, Lee HW, Schmaljohn CS, Dalrymple JM (1994) Serological relationships among viruses in the Hantavirus genus, family Bunyaviridae. Virology 198(1):196–204
Dargeviciute A, Brus Sjölander K, Sasnauskas K, Krüger DH, Meisel H, Ulrich R, Lundkvist A (2002) Yeast-expressed Puumala hantavirus nucleocapsid protein induces protection in a bank vole model. Vaccine 20:3523–3531
Dzagurova T, Tkachenko E, Slonova R, Ivanov L, Ivanidze E, Markeshin S, Dekonenko A, Niklasson B, Lundkvist A (1995) Antigenic relationships of hantavirus strains analysed by monoclonal antibodies. Arch Virol 140:1763–1773
Elgh F, Linderholm M, Wadell G, Tarnvik A, Juto P (1998) Development of humoral cross-reactivity to the nucleocapsid protein of heterologous hantaviruses in nephropathia epidemica. FEMS Immunol Med Microbiol 22:309–315
Franko MC, Gibbs CJ Jr, Lee PW, Gajdusek DC (1983) Monoclonal antibodies specific for Hantaan virus. Proc Natl Acad Sci USA 80(13):4149–4153
Gedvilaite A, Zvirbliene A, Staniulis J, Sasnauskas K, Krüger DH, Ulrich R (2004) Segments of Puumala hantavirus nucleocapsid protein inserted into chimeric polyomavirus-derived virus-like particles induce a strong immune response in mice. Viral Immunol 17:51–68
Geldmacher A, Schmaler M, Krüger DH, Ulrich R (2004) Yeast-expressed hantavirus Dobrava nucleocapsid protein induces a strong, long-lasting and highly cross-reactive immune response in mice. Viral Immunol 17:115–122
Geldmacher A, Skrastina D, Petrovskis I, Borisova G, Berriman JA, Roseman AM, Crowther RA, Fischer J, Musema S, Gelderblom HR, Lundkvist A, Renhofa R, Ose V, Krüger DH, Pumpens P, Ulrich R (2004) An amino-terminal segment of hantavirus nucleocapsid protein presented on hepatitis B virus core particles induces a strong and highly cross-reactive antibody response in mice. Virology 323:108–119
Geldmacher A, Skrastina D, Borisova G, Petrovskis I, Krüger DH, Pumpens P, Ulrich R (2005) A hantavirus nucleocapsid protein segment exposed on hepatitis B virus core particles is highly immunogenic in mice when applied without adjuvants or in the presence of pre-existing anti-core antibodies. Vaccine 23:3973–3983
Gött P, Zöller L, Darai G, Bautz EK (1997) A major antigenic domain of hantaviruses is located on the aminoproximal site of the viral nucleocapsid protein. Virus Genes 14(1):31–40
Green W, Feddersen R, Yousef O, Behr M, Smith K, Nestler J, Jenison S, Yamada T, Hjelle B (1998) Tissue distribution of hantavirus antigen in naturally infected humans and deer mice. J Infect Dis 177(6):1696–1700
Hjelle B, Jenison S, Torrez-Martinez N, Herring B, Quan S, Polito A, Pichuantes S, Yamada T, Morris C, Elgh F, Lee HW, Artsob H, Dinello R (1997) Rapid and specific detection of Sin Nombre virus antibodies in patients with hantavirus pulmonary syndrome by a strip immunoblot assay suitable for field diagnosis. J Clin Microbiol 35:600–608
Hujakka H, Koistinen V, Kuronen I, Eerikainen P, Parviainen M, Lundkvist A, Vaheri A, Vapalahti O, Narvanen A (2003) Diagnostic rapid tests for acute hantavirus infections: specific tests for Hantaan, Dobrava and Puumala viruses versus a hantavirus combination test. J Virol Methods 108:117–122
Jenison S, Yamada T, Morris C, Anderson B, Torrez-Martinez N, Keller N, Hjelle B (1994) Characterization of human antibody responses to four corners hantavirus infections among patients with hantavirus pulmonary syndrome. J Virol 68:3000–3006
Kang HJ, Bennett SN, Sumibcay L, Arai S, Hope AG, Mocz G, Song JW, Cook JA, Yanagihara R (2009) Evolutionary insights from a genetically divergent hantavirus harbored by the European common mole (Talpa europaea). PLoS One 4:e6149
Kang HJ, Arai S, Hope AG, Cook JA, Yanagihara R (2010) Novel Hantavirus in the Flat-Skulled Shrew (Sorex roboratus). Vector Borne Zoonotic Dis. doi:10.1089/vbz.2009.0159
Klempa B, Meisel H, Rath S, Bartel J, Ulrich R, Krüger DH (2003) Occurrence of renal and pulmonary syndrome in a region of northeast Germany where Tula virus circulates. J Clin Microbiol 41:4894–4897
Klempa B, Fichet-Calvet E, Lecompte E, Auste B, Aniskin V, Meisel H, Barriere P, Koivogui L, Meulen J, Krüger DH (2007) Novel hantavirus sequences in shrew, Guinea. Emerg Infect Dis 13:520–522
Klempa B, Tkachenko EA, Dzagurova TK, Yunicheva YV, Morozov VG, Okulova NM, Slyusareva GP, Smirnov A, Krüger DH (2008) Hemorrhagic fever with renal syndrome caused by 2 lineages of Dobrava hantavirus, Russia. Emerg Infect Dis 14(4):617–625
Koch J, Liang M, Queitsch I, Kraus AA, Bautz EK (2003) Human recombinant neutralizing antibodies against hantaan virus G2 protein. Virology 308(1):64–73
Kohler G, Milstein C (1975) Continuous culture of fused cells secreting antibody of predefined specificity. Nature 7:495–497
Koma T, Yoshimatsu K, Pini N, Safronetz D, Taruishi M, Levis S, Endo R, Shimizu K, Yasuda SP, Ebihara H, Feldmann H, Enria D, Arikawa J (2010) Truncated hantavirus nucleocapsid proteins for serotyping Sin Nombre, Andes, and Laguna Negra hantavirus infections in humans and rodents. J Clin Microbiol 48(5):1635–1642
Krüger DH, Ulrich R, Lundkvist A (2001) Hantavirus infections and their prevention. Microbes Infect 3:1129–1144
Lee HW, Lee PW, Johnson KM (1978) Isolation of the etiologic agent of Korean hemorrhagic fever. J Infect Dis 137:298–308
Liang M, Guttieri M, Lundkvist A, Schmaljohn C (1997) Baculovirus expression of a human G2-specific, neutralizing IgG monoclonal antibody to Puumala virus. Virology 235(2):252–260
Liang M, Mahler M, Koch J, Ji Y, Li D, Schmaljohn C, Bautz EK (2003) Generation of an HFRS patient-derived neutralizing recombinant antibody to Hantaan virus G1 protein and definition of the neutralizing domain. J Med Virol 69(1):99–107
Lundkvist A, Fatouros A, Niklasson B (1991) Antigenic variation of European haemorrhagic fever with renal syndrome virus strains characterized using bank vole monoclonal antibodies. J Gen Virol 72:2097–2103
Lundkvist A, Niklasson B (1992) Bank vole monoclonal antibodies against Puumala virus envelope glycoproteins: identification of epitopes involved in neutralization. Arch Virol 126:93–105
Lundkvist A, Horling J, Niklasson B (1993) The humoral response to Puumala virus infection (nephropathia epidemica) investigated by viral protein specific immunoassays. Arch Virol 130(1–2):121–130
Lundkvist A, Horling J, Athlin L, Rosen A, Niklasson B (1993) Neutralizing human monoclonal antibodies against Puumala virus, causative agent of nephropathia epidemica: a novel method using antigen-coated magnetic beads for specific B cell isolation. J Gen Virol 74(Pt 7):1303–1310
Lundkvist A, Horling J, Björsten S, Niklasson B (1995) Sensitive detection of hantaviruses by biotin-streptavidin enhanced immunoassays based on bank vole monoclonal antibodies. J Virol Methods 52(1–2):75–86
Lundkvist A, Björsten S, Niklasson B, Ahlborg N (1995) Mapping of B-cell determinants in the nucleocapsid protein of Puumala virus: definition of epitopes specific for acute immunoglobulin G recognition in humans. Clin Diagn Lab Immunol 2(1):82–86
Lundkvist A, Kallio-Kokko H, Sjölander KB, Lankinen H, Niklasson B, Vaheri A, Vapalahti O (1996) Characterization of Puumala virus nucleocapsid protein: identification of B-cell epitopes and domains involved in protective immunity. Virology 216(2):397–406
Lundkvist A, Vapalahti O, Plyusnin A, Sjölander KB, Niklasson B, Vaheri A (1996) Characterization of Tula virus antigenic determinants defined by monoclonal antibodies raised against baculovirus-expressed nucleocapsid protein. Virus Res 45(1):29–44
Lundkvist A, Apekina N, Myasnikov Y, Vapalahti O, Vaheri A, Plyusnin A (1997) Dobrava hantavirus outbreak in Russia. Lancet 350:781–782
Lundkvist A, Vasilenko V, Golovljova I, Plyusnin A, Vaheri A (1998) Human Dobrava hantavirus infections in Estonia. Lancet 352:369
Lundkvist A, Meisel H, Koletzki D, Lankinen H, Cifire F, Geldmacher A, Sibold C, Gött P, Vaheri A, Krüger DH, Ulrich R (2002) Mapping of B-cell epitopes in the nucleocapsid protein of Puumala hantavirus. Viral Immunol 15:177–192
Mazzarotto GA, Raboni SM, Stella V, Carstensen S, de Noronha L, Levis S, Zanluca C, Zanetti CR, Bordignon J, Duarte dos Santos CN (2009) Production and characterization of monoclonal antibodies against the recombinant nucleoprotein of Araucaria hantavirus. J Virol Methods 162(1–2):96–100
McCaughey C, Hart CA (2000) McCoughey and Hart 2000 Hantaviruses. J Med Microbiol 49(7):587–599
Meisel H, Lundkvist A, Gantzer K, Bar W, Sibold C, Krüger DH (1998) First case of infection with hantavirus Dobrava in Germany. Eur J Clin Microbiol Infect Dis 17:884–885
Meisel H, Wolbert A, Razanskiene A, Marg A, Kazaks A, Sasnauskas K, Pauli G, Ulrich R, Krüger DH (2006) Development of novel immunoglobulin G (IgG), IgA, and IgM enzyme immunoassays based on recombinant Puumala and Dobrava hantavirus nucleocapsid proteins. Clin Vaccine Immunol 13:1349–1357
Mertens M, Wolfel R, Ullrich K, Yoshimatsu K, Blumhardt J, Romer I, Esser J, Schmidt-Chanasit J, Groschup MH, Dobler G, Essbauer SS, Ulrich RG (2009) Seroepidemiological study in a Puumala virus outbreak area in South-East Germany. Med Mircobiol Immunol 198:83–91
Morii M, Yoshimatsu K, Arikawa J, Zhou G, Kariwa H, Takashima I (1998) Antigenic characterization of Hantaan and Seoul virus nucleocapsid proteins expressed by recombinant baculovirus: application of a truncated protein, lacking an antigenic region common to the two viruses, as a serotyping antigen. J Clin Microbiol 36(9):2514–2521
Nakamura I, Yoshimatsu K, Lee BH, Okumura M, Taruishi M, Araki K, Kariwa H, Takashima I, Arikawa J (2008) Development of a serotyping ELISA system for Thailand virus infection. Arch Virol 153(8):1537–1542
Nemirov K, Vapalahti O, Lundkvist A, Vasilenko V, Golovljova I, Plyusnina A, Niemimaa J, Laakkonen J, Henttonen H, Vaheri A, Plyusnin A (1999) Isolation and characterization of Dobrava hantavirus carried by the striped field mouse (Apodemus agrarius) in Estonia. J Gen Virol 80(Pt 2):371–379
Nichol ST, Spiropoulou CF, Morzunov S, Rollin PE, Ksiazek TG, Feldmann H, Sanchez A, Childs J, Zaki S, Peters CJ (1993) Genetic identification of a hantavirus associated with an outbreak of acute respiratory illness. Science 262(5135):914–917
Nichol ST, Beaty BJ, Elliott RM, Goldbach R, Plyusnin A, Schmaljohn CS, Tesh RB (2005) Family Bunyaviridae. In: Fauquet CM, Mayo MA, Maniloff J, Desselberger U, Ball LA (eds) Virus taxonomy. Eight report of the international committee on taxonomy of viruses. Elsevier Academic Press, San Diego, CA, pp 695–716
Niklasson B, Lundkvist A, Rossi CA, Ksiazek T, Summers PL, McClain DJ, Hooper J, Lee PW, Tsai TF, Kitamura T, Tomiyama T, Lee HW, Ivanov AP, Rezapkin GV, Tkachenko EA, Hjelle B, Torrez-Martinez N, Bharadwaj M (1998) Virus detection and identification with serological tests. In: Lee HW, Calisher C, Schmaljohn CS (eds) Manual of hemorrhagic fever with renal syndrome and hantavirus pulmonary syndrome. WHO Collaborating Center for Virus Reference and Research, Seoul, pp 80–130
Okumura M, Yoshimatsu K, Araki K, Lee BH, Asano A, Agui T, Arikawa J (2004) Epitope analysis of monoclonal antibody E5/G6, which binds to a linear epitope in the nucleocapsid protein of hantaviruses. Arch Virol 149(12):2427–2434
Padula PJ, Rossi CM, Della Valle MO, Martinez PV, Colavecchia SB, Edelstein A, Miguel SDL, Rabinovich RD, Segura EL (2000) Development and evaluation of a solid-phase enzyme immunoassay based on Andes hantavirus recombinant nucleoprotein. J Med Microbiol 49:149–155
Papa A, Johnson AM, Stockton PC, Bowen MD, Spiropoulou CF, Alexiou-Daniel S, Ksiazek TG, Nichol ST, Antoniadis A (1998) Retrospective serological and genetic study of the distribution of hantaviruses in Greece. J Med Virol 55:321–327
Plyusnin A, Vapalahti O, Lankinen H, Lehväslaiho H, Apekina N, Myasnikov Y, Kallio-Kokko H, Henttonen H, Lundkvist A, Brummer-Korvenkontio M et al (1994) Tula virus: a newly detected hantavirus carried by European common voles. J Virol 68(12):7833–7839
Plyusnin A, Cheng Y, Vapalahti O, Pejcoch M, Unar J, Jelinkova Z, Lehväslaiho H, Lundkvist A, Vaheri A (1995) Genetic variation in Tula hantaviruses: sequence analysis of the S and M segments of strains from Central Europe. Virus Res 39(2–3):237–250
Razanskiene A, Schmidt J, Geldmacher A, Ritzi A, Niedrig M, Lundkvist A, Krüger DH, Meisel H, Sasnauskas K, Ulrich R (2004) High yields of stable and highly pure nucleocapsid proteins of different hantaviruses can be generated in the yeast Saccharomyces cerevisiae. J Biotechnol 111:319–333
Ruo SL, Sanchez A, Elliott LH, Brammer LS, McCormick JB, Fisher-Hoch SP (1991) Monoclonal antibodies to three strains of hantaviruses: Hantaan, R22, and Puumala. Arch Virol 119(1–2):1–11
Salonen EM, Parren PW, Graus YF, Lundkvist A, Fisicaro P, Vapalahti O, Kallio-Kokko H, Vaheri A, Burton DR (1998) Human recombinant Puumala virus antibodies: cross-reaction with other hantaviruses and use in diagnostics. J Gen Virol 79(Pt4):659–665
Sandmann S, Meisel H, Razanskiene A, Wolbert A, Pohl B, Krüger DH, Sasnauskas K, Ulrich R (2005) Detection of human hantavirus infections in Lithuania. Infection 33:66–72
Schmaljohn C, Hjelle B (2007) Hantaviruses: a global disease problem. Emerg Infect Dis 3(2):95–104
Schmaljohn CS, Nichol ST (2001) Hantaviruses. In: Current Topics in microbiology and immunology, vol 256. Springer, Berlin
Schmidt J, Meisel H, Hjelle B, Krüger DH, Ulrich R (2005) Development and evaluation of serological assays for detection of human hantavirus infections caused by Sin Nombre virus. J Clin Virol 33:247–253
Schmidt J, Jandrig B, Klempa B, Yoshimatsu K, Arikawa J, Meisel H, Niedrig M, Pitra C, Krüger DH, Ulrich R (2005) Nucleocapsid protein of cell culture-adapted Seoul virus strain 80–39: Analysis of its encoding sequence, expression in yeast and immuno-reactivity. Virus Genes 30:37–48
Schmidt J, Meisel H, Capria SG, Petraityte R, Lundkvist A, Hjelle B, Vial PA, Padula P, Krüger DH, Ulrich R (2006) Serological assays for the detection of human Andes hantavirus infections based on its yeast-expressed nucleocapsid protein. Intervirology 49(3):173–184
Schultze D, Lundkvist A, Blauenstein U, Heyman P (2002) Tula virus infection associated with fever and exanthema after a wild rodent bite. Eur J Clin Microbiol Infect Dis 21:304–306
Sibold C, Ulrich R, Labuda M, Lundkvist A, Martens H, Schütt M, Gerke P, Leitmeyer K, Meisel H, Krüger DH (2001) Dobrava hantavirus causes hemorrhagic fever with renal syndrome in central Europe and is carried by two different Apodemus mice species. J Med Virol 63:158–167
Sjölander KB, Elgh F, Kallio-Kokko H, Vapalahti O, Hagglund M, Palmcrantz V, Juto P, Vaheri A, Niklasson B, Lundkvist A (1997) Evaluation of serological methods for diagnosis of Puumala hantavirus infection (nephropathia epidemica). J Clin Microbiol 35:3264–3268
Sjölander KB, Lundkvist A (1999) Dobrava virus infection: serological diagnosis and cross-reactions to other hantaviruses. J Virol Methods 80:137–143
Sugiyama K, Morikawa S, Matsuura Y, Tkachenko EA, Morita C, Komatsu T, Akao Y, Kitamura T (1987) Four serotypes of haemorrhagic fever with renal syndrome viruses identified by polyclonal and monoclonal antibodies. J Gen Virol 68(Pt4):979–987
Tischler ND, Galeno H, Rosemblatt M, Valenzuela PD (2005) Human and rodent humoral immune responses to Andes virus structural proteins. Virology 334(2):319–326
Tischler ND, Rosemblatt M, Valenzuela PD (2008) Characterization of cross-reactive and serotype-specific epitopes on the nucleocapsid proteins of hantaviruses. Virus Res 135(1):1–9
Vapalahti O, Kallio-Kokko H, Narvanen A, Julkunen I, Lundkvist A, Plyusnin A, Lehvaslaiho H, Brummer-Korvenkontio M, Vaheri A, Lankinen H (1995) Human B-cell epitopes of Puumala virus nucleocapsid protein, the major antigen in early serological response. J Med Virol 46(4):293–303
Vapalahti O, Lundkvist A, Kukkonen SKJ, Cheng Y, Gilljam M, Kanerva M, Manni T, Pejcoch M, Niemimaa J, Kaikusalo A, Henttonen H, Vaheri A, Plyusnin A (1996) Isolation and characterization of Tula virus, a distinct serotype in the genus Hantavirus, family Bunyaviridae. J Gen Virol 77:3063–3067
Wang M, Rossi C, Schmaljohn CS (1993) Expression of non-conserved regions of the S genome segments of three hantaviruses: evaluation of the expressed polypeptides for diagnosis of haemorrhagic fever with renal syndrome. J Gen Virol 74(Pt6):1115–1124
Yamada T, Hjelle B, Lanzi R, Morris C, Anderson B, Jenison S (1995) Antibody responses to Four Corners hantavirus infections in the deer mouse (Peromyscus maniculatus): identification of an immunodominant region of the viral nucleocapsid protein. J Virol 69:1939–1943
Yamanishi K, Dantas JR, Takahashi M, Yamanouchi T, Domae K, Takahashi Y, Tanishita O (1984) Antigenic differences between two viruses, isolated in Japan and Korea, that cause hemorrhagic fever with renal syndrome. J Virol 52(1):231–237
Yoshimatsu K, Arikawa J, Tamura M, Yoshida R, Lundkvist A, Niklasson B, Kariwa H, Azuma I (1996) Characterization of the nucleocapsid protein of Hantaan virus strain 76–118 using monoclonal antibodies. J Gen Virol 77:695–704
Zöller LG, Yang S, Gött P, Bautz EKF, Darai G (1993) A novel μ-capture enzyme-linked immunosorbent assay based on recombinant proteins for sensitive and specific diagnosis of hemorrhagic fever with renal syndrome. J Clin Microbiol 31:1194–1199
Zvirbliene A, Samonskyte L, Gedvilaite A, Voronkova T, Ulrich R, Sasnauskas K (2006) Generation of monoclonal antibodies of desired specificity using chimeric polyomavirus-derived virus-like particles. J Immunol Methods 311:57–70
Acknowledgments
The authors thank Detlev H. Krüger and Helga Meisel (Berlin, Germany) for support at the initial stage of work, Connie S. Schmaljohn (Fort Detrick, MD, USA) for provision of the PUUV-P360 N protein-encoding plasmid, Gereon Schares (Wusterhausen, Germany) for provision of negative control mAbs raised against Neospora caninum surface proteins and Lieselotte Minke (Wusterhausen) for excellent technical assistance. The experiments were supported by grants from DLR-Internationales Büro des BMBF (grant LTU 02/001), from the Lithuanian State and Study Foundation (project B-16/2006) and United States Public Health Services grant U19 AI45452.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kucinskaite-Kodze, I., Petraityte-Burneikiene, R., Zvirbliene, A. et al. Characterization of monoclonal antibodies against hantavirus nucleocapsid protein and their use for immunohistochemistry on rodent and human samples. Arch Virol 156, 443–456 (2011). https://doi.org/10.1007/s00705-010-0879-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-010-0879-6