Abstract
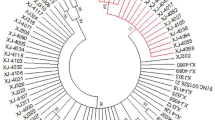
The E1 ORF is one of the most conserved regions in the human papillomavirus (HPV) genome. The complete E1 gene of the HPV16 genome was amplified with four overlapping primer sets in 16 high-grade (CIN II, III) and 13 low-grade cervical (CIN I) intraepithelial neoplasias as well as in one cervical cancer case. Sequence analysis of the E6 and E7 genes was also carried out in the same cervical samples in order to confirm the association between nucleotide sequence variations in the HPV16 E1 ORF and HPV16 variant lineages. Analysis of the E1 ORF revealed 27 nucleotide changes, and these changes were correlated with those found in HPV16 Asian American and African type II variants. Of these nucleotide variations, A1668G, G2073A, T2169C, T2189C, A2453T, C2454T, A2587T and G2650A were identified only in high-grade dysplasia cases. A phylogenetic tree of the E1 ORF and nucleotide sequence analysis of the E1, E6 and E7 genes revealed that intratypic nucleotide sequence polymorphisms located in the E1 ORF can be used to identify the major phylogenetic branch to which a HPV16 genome belongs. Moreover, amplification of the E1 ORF revealed a disruption between nucleotides 878 and 1523 in five high- and two low-grade cervical cases, indicating that integration of HPV DNA occurs at an early stage of viral infection.

Similar content being viewed by others
References
Bernard HU, Burk RD, Chen Z, van Doorslaer K, Hausen Hz, de Villiers EM (2010) Classification of papillomaviruses (PVs) based on 189 PV types and proposal of taxonomic amendments. Virology 401:70–79
de Villiers EM, Fauquet C, Broker TR, Bernard HU, zur Hausen H (2004) Classification of papillomaviruses. Virology 324:17–27
de Sanjose S, Quint WG, Alemany L, Geraets DT, Klaustermeier JE, Lloveras B, Tous S, Felix A, Bravo LE, Shin HR, Vallejos CS, de Ruiz PA, Lima MA, Guimera N, Clavero O, Alejo M, Llombart-Bosch A et al (2010) Retrospective International Survey and HPV Time Trends Study Group. Human papillomavirus genotype attribution in invasive cervical cancer: a retrospective cross-sectional worldwide study. Lancet Oncol 11:1048–1056
Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM (2010) Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer 127:2893–2917
Li N, Franceschi S, Howell-Jones R, Snijders PJF, Clifford GM (2011) Human papillomavirus type distribution in 30,848 invasive cervical cancers worldwide: Variation by geographical region, histological type and year of publication. Int J Cancer 128:927–935
Bernard HU (2005) The clinical importance of the nomenclature, evolution and taxonomy of human papillomaviruses. J Clin Virol 32:S1–S6
Gottschling M, Göker M, Stamatakis A, Bininda-Emonds OR, Nindl I, Bravo IG (2011) Quantifying the phylodynamic forces driving papillomavirus evolution. Mol Biol Evol 28:2101–2113
Chan SY, Bernard HU, Ong CK, Chan SP, Hofmann B, Delius H (1992) Phylogenetic analysis of 48 papillomavirus types and 28 subtypes and variants: a showcase for the molecular evolution of DNA viruses. J Virol 66:5714–5725
Yamada T, Wheeler CM, Halpern AL, Stewart AC, Hildesheim A, Jenison SA (1995) Human papillomavirus type 16 variant lineages in United States populationscharacterized by nucleotide sequence analysis of the E6, L2, and L1 coding segments. J Virol 69:7743–7753
Chen Z, Terai M, Fu L, Herrero R, DeSalle R, Burk RD (2005) Diversifying selection in human papillomavirus type 16 lineages based on complete genome analyses. J Virol 79:7014–7023
Ho L, Chan SY, Burk RD, Das BC, Fujinaga K, Icenogle JP, Kahn T, Kiviat N, Lancaster W, Mavromara-Nazos P (1993) The genetic drift of human papillomavirus type 16 is a means of reconstructing prehistoric viral spread and the movement of ancient human populations. J Virol 67:6413–6423
Ho L, Chan SY, Chow V, Chong T, Tay SK, Villa LL, Bernard HU (1991) Sequence variants of human papillomavirus type 16 in clinical samples permit verification and extension of epidemiological studies and construction of a phylogenetic tree. J Clin Microbiol 29:1765–1772
Yamada T, Manos MM, Peto J, Greer CE, Munoz N, Bosch FX, Wheeler CM (1997) Human papillomavirus type 16 sequence variation in cervical cancers: a worldwide perspective. J Virol 71:2463–2472
Carvajal-Rodríguez A (2008) Detecting recombination and diversifying selection in human alpha-papillomavirus. Infect Genet Evol 8:689–692
Tsakogiannis D, Ruether IGA, Kyriakopoulou Z, Pliaka V, Skordas V, Gartzonika C, Levidiotou-Stefanou S, Markoulatos P (2012) Molecular and phylogenetic analysis of the HPV 16 E4 gene in cervical lesions from women in Greece. Arch Virol 157:1729–1739
Cornet I, Gheit T, Franceschi S, Vignat J, Burk RD, Sylla BS, Tommasino M, Clifford GM, IARC HPV Variant Study Group (2012) Human papillomavirus type 16 genetic variants: phylogeny and classification based on E6 and LCR. J Virol 86:6855–6861
Smith B, Chen Z, Reimers L, van Doorslaer K, Schiffman M, Desalle R, Herrero R, Yu K, Wacholder S, Wang T, Burk RD (2011) Sequence imputation of HPV16 genomes for genetic association studies. PLoS One 6:e21375
Chen Z, Schiffman M, Herrero R, Desalle R, Anastos K, Segondy M, Sahasrabuddhe VV, Gravitt PE, Hsing AW, Burk RD (2011) Evolution and taxonomic classification of human papillomavirus 16 (HPV16)-related variant genomes: HPV31, HPV33, HPV35, HPV52, HPV58 and HPV67. PLoS One 6:e20183
Huertas-Salgado A, Martín-Gámez DC, Moreno P, Murillo R, Bravo MM, Villa L, Molano M (2011) E6 molecular variants of human papillomavirus (HPV) type 16: an updated and unified criterion for clustering and nomenclature. Virology 410:201–215
García-Vallvé S, Alonso A, Bravo IG (2005) Papillomaviruses: different genes have different histories. Trends Microbiol 13:514–521
Titolo S, Brault K, Majewski J, White PW, Archambault J (2003) Characterization of the minimal DNA binding domain of the human papillomavirus e1 helicase: fluorescence anisotropy studies and characterization of a dimerization-defective mutant protein. J Virol 77:5178–5191
Titolo S, Pelletier A, Pulichino AM, Brault K, Wardrop E, White PW, Cordingley MG, Archambault J (2000) Identification of domains of the human papillomavirus type 11 E1 helicase involved in oligomerization and binding to the viral origin. J Virol 74:7349–7361
Amin AA, Titolo S, Pelletier A, Fink D, Cordingley MG, Archambault J (2000) Identification of domains of the HPV11 E1 protein required for DNA replication in vitro. Virology 272:137–150
White PW, Pelletier A, Brault K, Titolo S, Welchner E, Thauvette L, Fazekas M, Cordingley MG, Archambault J (2001) Characterization of recombinant HPV6 and 11 E1 helicases: effect of ATP on the interaction of E1 with E2 and mapping of a minimal helicase domain. J Biol Chem 276:22426–22438
Stenlund A (2003) E1 initiator DNA binding specificity is unmasked by selective inhibition of non-specific DNA binding. EMBO J 22:954–963
Clower RV, Hu Y, Melendy T (2006) Papillomavirus E2 protein interacts with and stimulates human topoisomerase I. Virology 348:13–18
Loo YM, Melendy T (2004) Recruitment of replication protein A by the papillomavirus E1 protein and modulation by single-stranded DNA. J Virol 78:1605–1615
Parker LM, Harris S, Gossen M, Botchan MR (2000) The bovine papillomavirus E2 transactivator is stimulated by the E1 initiator through the E2 activation domain. Virology 270:430–443
Conger KL, Liu JS, Kuo SR, Chow LT, Wang TS (1999) Human papillomavirus DNA replication. Interactions between the viral E1 protein and two subunits of human dna polymerase alpha/primase. J Biol Chem 274:2696–2705
Masterson PJ, Stanley MA, Lewis AP, Romanos MA (1998) A C-terminal helicase domain of the human papillomavirus E1 protein binds E2 and the DNA polymerase alpha-primase p68 subunit. J Virol 72:7407–7419
Cricca M, Venturoli S, Leo E, Costa S, Musiani M, Zerbini M (2009) Disruption of HPV16 E1 and E2 genes in precancerous cervical lesions. J Virol Methods 158:180–183
Arias-Pulido H, Peyton CL, Joste NE, Vargas H, Wheeler CM (2006) Human papillomavirus type 16 integration in cervical carcinoma in situ and in invasive cervical cancer. J Clin Microbiol 44:1755–1762
Huang LW, Choa SL, Lee BH (2008) Integration of human papillomavirus type-16 and type-18 is a very early event in cervical carcinogenesis. J Clin Pathol 61:627–631
Kulmala SM, Syrjänen SM, Gyllensten UB, Shabalova IP, Petrovichev N, Tosi P, Syrjänen KJ, Johansson BC (2006) Early integration of high copy HPV16 detectable in women with normal and low grade cervical cytology and histology. J Clin Pathol 59:513–517
Casas I, Powell L, Klapper PE, Cleator GM (1995) New method for the extraction of viral RNA and DNA from cerebrospinal fluid for use in the polymerase chain reaction assay. J Virol Methods 53:25–36
Tsakogiannis D, Ruether IG, Kyriakopoulou Z, Pliaka V, Theoharopoulou A, Skordas V, Panotopoulou E, Nepka C, Markoulatos P (2012) Sequence variation analysis of the E2 gene of human papilloma virus type 16 in cervical lesions from women in Greece. Arch Virol 157:825–832
Puranen M, Saarikoski S, Syrjänen K, Syrjänen S (1996) Polymerase chain reaction amplification of human papillomavirus DNA from archival, Papanicolaou-stained cervical smears. Acta Cytol 40:391–395
Li W, Wang W, Si M, Han L, Gao Q, Luo A, Li Y, Lu Y, Wang S, Ma D (2008) The physical state of HPV16 infection and its clinical significance in cancer precursor lesion and cervical carcinoma. J Cancer Res Clin Oncol 134:1355–1361
Sotlar K, Diemer D, Dethleffs A, Hack Y, Stubner A, Vollmer N, Menton S, Menton M, Dietz K, Wallwiener D, Kandolf R, Bültmann B (2004) Detection and typing of human papillomavirus by e6 nested multiplex PCR. J Clin Microbiol 42:3176–3184
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Kalantari M, Karlsen F, Kristensen G, Holm R, Hagmar B, Johansson B (1998) Disruption of the E1 and E2 reading frames of HPV 16 in cervical carcinoma is associated with poor prognosis. Int J Gynecol Pathol 17:146–153
Briolat J, Dalstein V, Saunier M, Joseph K, Caudroy S, Prétet JL, Birembaut P, Clavel C (2007) HPV prevalence, viral load and physical state of HPV-16 in cervical smears of patients with different grades of CIN. Int J Cancer 121:2198–2204
Schiffman M, Rodriguez AC, Chen Z, Wacholder S, Herrero R, Hildesheim A, Desalle R, Befano B, Yu K, Safaeian M, Sherman ME, Morales J, Guillen D, Alfaro M, Hutchinson M, Solomon D, Castle PE, Burk RD (2010) A population-based prospective study of carcinogenic human papillomavirus variant lineages, viral persistence, and cervical neoplasia. Cancer Res 70:3159–3169
Sichero L, Ferreira S, Trottier H, Duarte-Franco E, Ferenczy A, Franco EL, Villa LL (2007) High grade cervical lesions are caused preferentially by non-European variants of HPVs 16 and 18. Int J Cancer 120:1763–1768
Quint KD, de Koning MN, van Doorn LJ, Quint WG, Pirog EC (2010) HPV genotyping and HPV16 variant analysis in glandular and squamous neoplastic lesions of the uterine cervix. Gynecol Oncol 117:297–301
Tornesello ML, Losito S, Benincasa G, Fulciniti F, Botti G, Greggi S, Buonaguro L, Buonaguro FM (2011) Human papillomavirus (HPV) genotypes and HPV16 variants and risk of adenocarcinoma and squamous cell carcinoma of the cervix. Gynecol Oncol 121:32–42
Acknowledgments
This work was supported by research grants from the Postgraduate Programmes ‘Applications of Molecular Biology-Genetics. Diagnostic Biomarkers’, code 3817, and “Biotechnology”, code 3439, of the University of Thessaly, School of Health Sciences, Department of Biochemistry & Biotechnology.
Conflict of interest
All authors declare that they have no conflicting or dual interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tsakogiannis, D., Darmis, F., Gortsilas, P. et al. Nucleotide polymorphisms of the human papillomavirus 16 E1 gene. Arch Virol 159, 51–63 (2014). https://doi.org/10.1007/s00705-013-1790-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-013-1790-8