Abstract
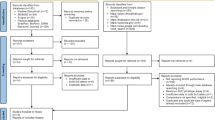
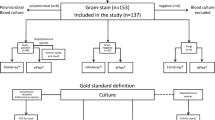
The Verigene Gram-Positive Blood Culture (BC-GP) Assay (Nanosphere Inc., Northbrook, IL) is a microarray-based test designed to rapidly identify directly from positive blood cultures multiple bacterial species and their antimicrobial resistance markers. Nonduplicate blood cultures from 118 patients admitted to Erasme Hospital were prospectively enrolled. All but six organisms were members of the panel (95.6 %). For the identification of pathogens and detection of the mecA gene, the agreement with routine methods was 87.6 % and 97.7 %, respectively. The performance of the BC-GP assay was lower with polymicrobial than with monomicrobial blood cultures. Another concern of the BC-GP assay was the misidentification of Streptococcus mitis as S. pneumoniae (3/8). The BC-GP assay is a rapid and accurate tool for the simultaneous detection of multiple sepsis-causing bacteria and resistant genes from blood cultures, which could have an impact on patient management and healthcare cost.

Similar content being viewed by others
References
Clinical and Laboratory Standards Institute (CLSI) (2013) Performance standards for antimicrobial susceptibility testing; 23rd informational supplement. CLSI document M100-S23. CLSI, Wayne, PA
Maes N, Magdalena J, Rottiers S, De Gheldre Y, Struelens MJ (2002) Evaluation of a triplex PCR assay to discriminate Staphylococcus aureus from coagulase-negative staphylococci and determine methicillin resistance from blood cultures. J Clin Microbiol 40:1514–1517
Mestas J, Polanco CM, Felsenstein S, Dien Bard J (2014) Performance of the Verigene Gram-positive blood culture assay for direct detection of Gram-positive organisms and resistance markers in a pediatric hospital. J Clin Microbiol 52:283–287
Beal SG, Ciurca J, Smith G, John J, Lee F, Doern CD, Gander RM (2013) Evaluation of the Nanosphere Verigene Gram-positive blood culture assay with the VersaTREK blood culture system and assessment of possible impact on selected patients. J Clin Microbiol 51:3988–3992
Wojewoda CM, Sercia L, Navas M, Tuohy M, Wilson D, Hall GS, Procop GW, Richter SS (2013) Evaluation of the Verigene Gram-positive blood culture nucleic acid test for rapid detection of bacteria and resistance determinants. J Clin Microbiol 51:2072–2076
Samuel LP, Tibbetts RJ, Agotesku A, Fey M, Hensley R, Meier FA (2013) Evaluation of a microarray-based assay for rapid identification of Gram-positive organisms and resistance markers in positive blood cultures. J Clin Microbiol 51:1188–1192
Scott LJ (2013) Verigene® Gram-positive blood culture nucleic acid test. Mol Diagn Ther 17:117–122
Sullivan KV, Turner NN, Roundtree SS, Young S, Brock-Haag CA, Lacey D, Abuzaid S, Blecker-Shelly DL, Doern CD (2013) Rapid detection of Gram-positive organisms by use of the Verigene Gram-positive blood culture nucleic acid test and the BacT/Alert Pediatric FAN system in a multicenter pediatric evaluation. J Clin Microbiol 51:3579–3584
Buchan BW, Ginocchio CC, Manii R, Cavagnolo R, Pancholi P, Swyers L, Thomson RB Jr, Anderson C, Kaul K, Ledeboer NA (2013) Multiplex identification of Gram-positive bacteria and resistance determinants directly from positive blood culture broths: evaluation of an automated microarray-based nucleic acid test. PLoS Med 10:e1001478
Alby K, Daniels LM, Weber DJ, Miller MB (2013) Development of a treatment algorithm for streptococci and enterococci from positive blood cultures identified with the Verigene Gram-positive blood culture assay. J Clin Microbiol 51:3869–3871
Ikryannikova LN, Filimonova AV, Malakhova MV, Savinova T, Filimonova O, Ilina EN, Dubovickaya VA, Sidorenko SV, Govorun VM (2013) Discrimination between Streptococcus pneumoniae and Streptococcus mitis based on sorting of their MALDI mass spectra. Clin Microbiol Infect 19:1066–1071
Dubois D, Segonds C, Prere MF, Marty N, Oswald E (2013) Identification of clinical Streptococcus pneumoniae isolates among other alpha and nonhemolytic streptococci by use of the Vitek MS matrix-assisted laser desorption ionization-time of flight mass spectrometry system. J Clin Microbiol 51:1861–1867
La Scola B, Raoult D (2009) Direct identification of bacteria in positive blood culture bottles by matrix-assisted laser desorption ionisation time-of-flight mass spectrometry. PLoS One 4:e8041
Kok J, Thomas LC, Olma T, Chen SC, Iredell JR (2011) Identification of bacteria in blood culture broths using matrix-assisted laser desorption-ionization Sepsityper™ and time of flight mass spectrometry. PLoS One 6:e23285
Tissari P, Zumla A, Tarkka E, Mero S, Savolainen L, Vaara M, Aittakorpi A, Laakso S, Lindfors M, Piiparinen H, Mäki M, Carder C, Huggett J, Gant V (2010) Accurate and rapid identification of bacterial species from positive blood cultures with a DNA-based microarray platform: an observational study. Lancet 375:224–230
Martin GS, Mannino DM, Eaton S, Moss M (2003) The epidemiology of sepsis in the United States from 1979 through 2000. N Engl J Med 348:1546–1554
Acknowledgments
We thank Soultana Pavlov, Angélique Fourmeaux, Sandra Cuvellier, and Anne-Laurence Debyttere for their technical assistance with the blood cultures processing and Nanosphere, which kindly provided the kits used in this study.
Conflict of interest
The authors declare no conflict of interest in relation to this work.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 17 kb)
Rights and permissions
About this article
Cite this article
Dodémont, M., De Mendonça, R., Nonhoff, C. et al. Evaluation of Verigene Gram-positive blood culture assay performance for bacteremic patients. Eur J Clin Microbiol Infect Dis 34, 473–477 (2015). https://doi.org/10.1007/s10096-014-2250-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-014-2250-4