Abstract
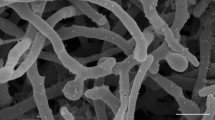
A novel actinomycete, designated strain NEAU-GRX11T, was isolated from muddy soil collected from a stream of Jinlong Mountain in Harbin, north China. The organism was found to have morphological and chemotaxonomic characteristics typical of the genus Micromonospora. The 16S rRNA gene sequence of strain NEAU-GRX11T showed highest similarity to Micromonospora zamorensis CR38T (99.2 %), Micromonospora saelicesensis Lupac 09T (99.0 %), Micromonospora chokoriensis 2-19/6T (98.7 %), Micromonospora coxensis 2-30-b/28T (98.5 %), Micromonospora aurantiaca ATCC 27029T (98.4 %) and Micromonospora lupini lupac 14NT (98.3 %). Phylogenetic analysis based on the 16S rRNA gene and gyrB gene demonstrated that strain NEAU-GRX11T was a member of the genus Micromonospora and supported the closest phylogenetic relationship to M. zamorensis CR38T, M. saelicesensis Lupac 09T, M. chokoriensis 2-19/6T and M. lupini lupac 14NT. A combination of DNA–DNA hybridization and some phenotypic characteristics indicated that the novel strain could be readily distinguished from these closest phylogenetic relatives. Therefore, it is proposed that NEAU-GRX11T represents a novel species of the genus Micromonospora, for which the name Micromonospora jinlongensis sp. nov. is proposed. The type strain is NEAU-GRX11T (=CGMCC 4.7103T=DSM 45876T).


Similar content being viewed by others
References
Ara I, Kudo T (2007) Two new species of the genus Micromonospora: micromonospora chokoriensis sp. nov. and Micromonospora coxensis sp. nov., isolated from sandy soil. J Gen Appl Microbiol 53:29–37
Carro L, Pukall R, Spröer C, Kroppenstedt RM, Trujillo ME (2012) Micromonospora cremea sp. nov. and Micromonospora zamorensis sp. nov., isolated from the rhizosphere of Pisum sativum. Int J Syst Evol Microbiol 62:2971–2977
Carro L, Pukall R, Spröer C, Kroppenstedt RM, Trujillo ME (2013) Micromonospora halotolerans sp. nov., isolated from the rhizosphere of a Pisum sativum plant. Antonie Leeuwenhoek Int J Gen 103:1245–1254
Collins MD (1985) Isoprenoid quinone analyses in bacterial classification and identification. In: Goodfellow M, Minnikin DE (eds) Chemical methods in bacterial systematics. Academic Press, London, pp 267–284
De Ley J, Cattoi H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Everest GJ, Meyers PR (2013) Micromonospora equina sp. nov., isolated from soil from a racecourse. Int J Syst Evol Microbiol 63:879–885
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Garcia LC, Martínez-Molina E, Trujillo ME (2010) Micromonospora pisi sp. nov., isolated from root nodules of Pisum sativum. Int J Syst Evol Microbiol 60:331–337
Gordon RE, Barnett DA, Handerhan JE, Pang C (1974) Nocardia coeliaca, Nocardia autotrophica, and the nocardin strain. Int J Syst Bacteriol 24:54–63
Gurovic MS, Müller S, Domin N, Seccareccia I, Nietzsche S, Martin K, Nett M (2013) Micromonospora schwarzwaldensis sp. nov., a producer of telomycin, isolated from soil in the Black Forest. Int J Syst Evol Microbiol. doi:10.1099/ijs.0.051623-0
Hayakawa M, Nonomura H (1987) Humic acid-vitamin agar, a new medium for the selective isolation of soil actinomycetes. J Ferment Technol 65:501–509
Huss VAR, Festl H, Schleifer KH (1983) Studies on the spectrometric determination of DNA hybridisation from renaturation rates. Syst Appl Microbiol 4:184–192
Jia FY, Liu CX, Wang XJ, Zhao JW, Liu QF, Zhang J, Gao RX, Xiang WS (2013) Wangella harbinensis gen. nov., sp. nov., a new member of the family Micromonosporaceae. Antonie Leeuwenhoek Int J Gen 103:399–408
Kasai H, Tamura T, Harayama S (2000) Intrageneric relationships among Micromonospora species deduced from gyrB-based phylogeny and DNA relatedness. Int J Syst Evol Microbiol 50:127–134
Kawamoto I (1989) Genus micromonospora Orskov 1923 147AL. In: Williams ST, Sharpe ME, Holt JG (eds) Ber-gey’s manual of systematic bacteriology, vol 4. Williams and Wilkins, Baltimore, pp 2442–2450
Kelly KL (1964) Inter-Society Color Council–National Bureau of Standards color name Charts illustrated with centroid colors. US Government Printing Office, Washington, DC
Kim SB, Brown R, Oldfield C, Gilbert SC, Iliarionov S, Goodfellow M (2000) Gordonia amicalis sp. nov., a novel dibenzothiophene-desulphurizing actinomycete. Int J Syst Evol Microbiol 50:2031–2036
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Koch C, Kroppenstedt RM, Stackebrandt E (1996) Intrageneric relationships of the actinomycete genus Micromonospora. Int J Syst Bacteriol 46:383–387
Kroppenstedt RM (1985) Fatty acid and menaquinone analysis of actinomycetes and related organisms. In: Goodfellow M, Minnikin DE (eds) Chemical methods in bacterial systematics. Academic press, London, pp 173–199
Lechevalier MP, Lechevalier HA (1980) The chemotaxonomy of actinomycetes. In: Dietz A, Thayer DW (eds) Actinomycete taxonomy special publication, vol 6. Society of Industrial Microbiology, Arlington, pp 227–291
Lechevalier MP, De Bièvre C, Lechevalier HA (1977) Chemotaxonomy of aerobic actinomycetes: phospholipid composition. Biochem Syst Ecol 5:249–260
Li L, Mao YJ, Xie QY, Deng Z, Hong K (2013a) Micromonospora avicenniae sp. nov., isolated from a root of Avicennia marina. Antonie Leeuwenhoek Int J Gen 103:1089–1096
Li L, Tang YL, Wei B, Xie QY, Deng Z, Hong K (2013b) Micromonospora sonneratiae sp. nov., isolated from a root of Sonneratia apetala. Int J Syst Evol Microbiol 63:2383–2388
Lǖdemann GM, Brodsky BC (1963) Taxonomy of gentamicin-producing Micromonospora. Antimicrob Agents Chemother 161:116–124
Maldonado LA, Fragoso-Yáñez D, Pérez-García A, Rosellón-Druker J, Quintana ET (2009) Actinobacterial diversity from marine sediments collected in Mexico. Antonie Leeuwenhoek Int J Gen 95:111–120
Mandel M, Marmur J (1968) Use of ultraviolet absorbance temperature profile for determining the guanine plus cytosine content of DNA. Methods Enzymol 12B:195–206
McKerrow J, Vagg S, McKinney T, Seviour EM, Maszenan AM, Brooks P, Seviour RJ (2000) A simple HPLC method for analysing diaminopimelic acid diastereomers in cell walls of Gram-positive bacteria. Lett Appl Microbiol 30:178–182
Mincer TJ, Jensen PR, Kauffman CA, Fenical W (2002) Widespread and persistent populations of a major actinomycete taxon in ocean sediments. Appl Environ Microbiol 68:5005–5011
Minnikin DE, Hutchinson IG, Caldicott AB, Goodfellow M (1980) Thin-layer chromatography of methanolysates of mycolic acid-containing bacteria. J Chromatogr 188:221–233
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JK (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Nimaichand S, Zhang YG, Cheng J, Li L, Zhang DF, Zhou EM, Dong L, Ningthoujam DS, Li WJ (2013) Micromonospora kangleipakensis sp. nov., isolated from a sample of Limestone quarry. Int J Syst Evol Microbiol. doi:10.1099/ijs.0.052746-0
Ørskov J (1923) Investigations into the morphology of the ray fungi. Levin and Munksgaard, Enhagen
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington DC, pp 607–654
Songsumanus A, Tanasupawat S, Igarashi Y, Kudo T (2013) Micromonospora maritima sp. nov., isolated from mangrove soil. Int J Syst Evol Microbiol 63:554–559
Supong K, Suriyachadkun C, Pittayakhajonwut P, Suwanborirux K, Thawai C (2013a) Micromonospora spongicola sp. nov., an actinomycete isolated from a marine sponge in the Gulf of Thailand. J Antibiot (Tokyo). doi:10.1038/ja.2013.35
Supong K, Suriyachadkun C, Tanasupawat S, Suwanborirux K, Pittayakhajonwut P, Kudo T, Thawai C (2013b) Micromonospora sediminicola sp. nov., isolated from marine sediment. Int J Syst Evol Microbiol 63:570–575
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: Molecular Evolutionary Genetics Analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Trujillo ME, Kroppenstedt RM, Fernández-Molinero C, Schumann P, Martínez-Molina E (2007) Micromonospora lupini sp. nov. and Micromonospora saelicesensis sp. nov., isolated from root nodules of Lupinus angustifolius. Int J Syst Evol Microbiol 57:2799–2804
Trujillo ME, Alonso-Vega P, Rodríguez R, Carro L, Cerda E, Alonso P, Martínez-Molina E (2010) The genus Micromonospora is widespread in legume root nodules: the example of Lupinus angustifolius. ISME J 4:1265–1281
Uchida K, Kudo T, Suzuki K, Nakase T (1999) A new rapid method of glycolate test by diethyl ether extraction, which is applicable to a small amount of bacterial cells of less than one milligram. J Gen Appl Microbiol 45:49–56
Waksman SA (1961) The Actinomycetes, vol. 2, Classification, identification and descriptions of genera and species. Williams and Wilkins, Baltimore
Waksman SA (1967) The Actinomycetes. A summary of current knowledge, Ronald
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE et al (1987) International Committee on Systematic Bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Syst Bacteriol 37:463–464
Wu C, Lu X, Qin M, Wang Y, Ruan J (1989) Analysis of menaquinone compound in microbial cells by HPLC. Microbiology [English translation of Microbiology (Beijing)] 16:176–178
Xiang WS, Liu CX, Wang XJ, Du J, Xi LJ, Huang Y (2011) Actinoalloteichus nanshanensis sp. nov., isolated from the rhizosphere of a fig tree (Ficus religiosa). Int J Syst Evol Microbiol 61:1165–1169
Xie QY, Lin HP, Li L, Brown R, Goodfellow M, Deng ZX, Hong K (2012) Verrucosispora wenchangensis sp. nov., isolated from mangrove soil. Antonie Leeuwenhoek Int J Gen 102:1–7
Yokota A, Tamura T, Hasegawa T, Huang LH (1993) Catenuloplanes japonicas gen. nov., sp. nov., nom. rev., a new genus of the order Actinomycetales. Int J Syst Bacteriol 43:805–812
Acknowledgments
This work was supported in part by grants from the Opening Fund of Key Laboratory of Soybean Biology in Chinese Ministry of Education (No. SB12B04), the National Key Project for Basic Research (No. 2010CB126102), the National Outstanding Youth Foundation (No. 31225024), the Special Foundation for Scientific and Technological Innovation Research of Harbin (No. 2011RFXXN038) and the Natural Science Foundation of Heilongjiang Province (No. C201029).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Ruixia Gao and Chongxi Liu contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gao, R., Liu, C., Zhao, J. et al. Micromonospora jinlongensis sp. nov., isolated from muddy soil in China and emended description of the genus Micromonospora . Antonie van Leeuwenhoek 105, 307–315 (2014). https://doi.org/10.1007/s10482-013-0074-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-013-0074-3