Abstract
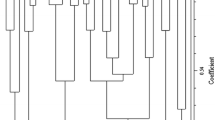
Olive (Olea europaea) is an ancient and important crop in both olive oil production and table use. It is important to identify the genetic diversity of olive genetic resources for cultivar development and evaluation of olive germplasm. In the study, 14 microsatellite markers (UDO4, UDO8, UDO9, UDO11, UDO12, UDO22, UDO24, UDO26, UDO28, DCA9, DCA11, DCA13, DCA15, and DCA18) were used to assess the genetic variation on 76 olive (Olea europaea L.) genotypes from Mardin province together with 6 well-known Turkish and 4 well-known foreign reference cultivars. All microsatellite markers showed polymorphism and the number of alleles varied between 9 and 22, with an average of 14.57. The most informative loci were DCA 11 (22 alleles) and DCA 9 (21 alleles). Dendrogram based on genetic distances was constructed for the 86 olive genotypes/cultivars, which revealed the existence of different clusters. The high genetic similarity was evident between Bakırkire2 and Zinnar5 (0.74) genotypes, while the most genetically divergent genotypes were Gürmeşe5 and Yedikardeşler2 (0.19). It was concluded that there was abundant SSR polymorphism in olive germplasm in southern Anatolia in Turkey and could be important for future breeding activities.

Similar content being viewed by others
References
Abdessemed S, Muzzalupo I, Benbouza H (2015) Assessment of genetic diversity among Algerian olive (Olea europaea L.) cultivars using SSR marker. Sci Hortic 192:10–20
Alba V, Montemurro C, Sabetta W, Pasqualone A, Blanco A (2009) SSR-based identification key of cultivars of Olea europaea L. diffused in Southern-Italy. Sci Hortic 123:11–16
Albertini R, Torricelli E, Bitocchi E, Raggi L, Marconi G, Pollastri L, Di Minco G, Battistini A, Papa R, Veronesi F (2011) Structure of genetic diversity in Olea europaea L. cultivars from central Italy. Mol Breed 27(4):533–547
Barranco D, Cimato A, Fiorino P, Rallo L, Touzani A, Castaneda C, Serafin F, Trujillo I (2000) World catalogue of olive varieties. International Olive Oil Council, Madrid 360 pp
Belaj A, Satovic Z, Cipriani G, Baldoni L, Testolin R, Rallo L, Trujillo I (2003) Comparative study of the discriminating capacity of RAPD, AFLP and SSR markers and their effectiveness in establishing genetic relationships in olive. Theor Appl Genet 107:736–744
Besnard G, Baradat P, Berville A (2001) Genetic relationships in the olive (Olea europaea L.) reflect multilocal selection of cultivars. Theor Appl Genet 102:251–258
Bowcock AM, Ruiz-Linares A, Tomfohrde T, Minch E, Kidd JR, Cavalli-Sforza LL (1994) High resolution of human evolutionary trees with polymorphic microsatellites. Nature 368:455–457
Cheng XM, Xu JS, Xia S, Gu JX, Yang Y, Fu J, Qian XJ, Zhang SC, Wu JS, Liu KD (2009) Development and genetic mapping of microsatellite markers froö genome survey sequences in Brassica napus. Theor Appl Genet 118:1121–1131
Cipriani G, Marrazzo MT, Marconi R, Cimato A, Testolin R (2002) Microsatellite markers isolated in olive (Olea europaea L.) are suitable for individual fingerprinting and reveal polymorphism within ancient cultivars. Theor Appl Genet 104:223–228
Contento A, Ceccarelli M, Gelati MT, Maggini F, Baldoni L, Cionini PG (2002) Diversity of olea genotypes and the origin of cultivated olives. Theor Appl Genet 104:1229–1238
Ercisli S, Ipek A, Barut E (2011) SSR marker-based DNA fingerprinting and cultivar identification of olives (Olea europaea L.). Biochem Genet 49(9–10):555–561
Ipek A, Barut E, Gulen H, Ipek M (2012) Assessment of inter and intra-cultivar variations in olive using SSR markers. Sci Agricola 69:327–335
Kaczmarska E, Gawronski J, Dyduch-Sieminska M, Najda A, Marecki W, Zebrowska J (2015) Genetic diversity and chemical characterization of selected Polish and Russian cultivars and clones of blue honeysuckle (Lonicera caerulea). Turk J Agric For 39:394402
Kaya HB, Cetin O, Kaya H, Sahin M, Sefer F, Kahraman A, Tanyolac B (2013) SNP discovery by Illumina-based transcriptome sequencing of the olive and the genetic characterization of Turkish olive genotypes revealed by AFLP, SSR and SNP markers. PloS One 8:e73674
Lopes MS, Mendonca D, Sefc KM, Sabino Gil F, da Camara Machado A (2004) Genetic evidence of intra-cultivar variability within Iberian olive cultivars. HortScience 39:1562–1565
Minch E, Ruiz-Linares A, Goldstein DB, Feldman M, Cavalli-Sforza LL (1995) Microsat (Version 1.4d): a computer program for calculating various statistics on microsatellite allele data. Stanford University Medical Center, Stanford
Mishra PK, Ram RB, Kumar N (2015) Genetic variability, heritability, and genetic advance in strawberry (Fragaria × ananassa Duch.). Turk J Agric For 39:451–458
Mlcek J, Valsikova M, Druzbikova H, Ryant P, Jurikova T, Sochor J, Borkovcova M (2015) The antioxidant capacity and macroelement content of several onion cultivars. Turk J Agric For 39:999–1004
Muzzalupo I, Vendramin GG, Chiappetta A (2014) Genetic biodiversity of Italian olives (Olea europaea) germplasm analyzed by SSR markers. Sci World J 116(3):352–359
Nemli S, Kianoosh T, Tanyolaç MD (2015) Genetic diversity and population structure of common bean (Phaseolus vulgaris L.) accessions through retrotransposon-based interprimer binding sites (iPBSs) markers. Turk J Agric For 39:940–948
Noormohammadi Z, Hosseini-Mazinani M, Trujillo I, Belaj A (2009) Study of intracultivar variation among main Iranian olive cultivars using SSR markers. Acta Biol Szeged 53(1):27–32
Paetkau D, Calvert W, Stirling I, Strobeck C (1995) Microsatellite analysis of population structure in Canadian polar bears. Mol Ecol 4:347–354
Poljuha D, Sladonja B, Setic E, Milotic A, Bandelj D, Jakse J, Javornik B (2008) DNA fingerprinting of olive varieties in Istria (Croatia) by microsatellite markers. Sci Hortic 115:223–230
Rohlf FJ (2004) NTSYS-pc numerical taxonomy and multivariate analysis system. Version 2.11V. Exeter software, Setauket
Roubos K, Moustakas M, Aravanopoulos FA (2010) Molecular identification of greek olive (Olea europaea) cultivars based on microsatellite loci. Genet Mol Res 9(3):1865–1876
Sakar E, Celik M, Ergul A (2013) Molecular analysis of olive germplasms grown in Adiyaman based on SSR markers. Zeytin Dostu 27:78–83
Sneath PHA, Sokal RR (1973) Numerical taxonomy: the principles and practice of numerical classification. Freeman, San Francisco
Zaher H, Boulouha B, Baaziz M, Sikaoui L, Gaboun F, Udupa SM (2011) Morphological and genetic diversity in olive (Olea europaea subsp. europaea L.) clones and varieties. Plant Omics J 4(7):370–376
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sakar, E., Unver, H. & Ercisli, S. Genetic Diversity Among Historical Olive (Olea europaea L.) Genotypes from Southern Anatolia Based on SSR Markers. Biochem Genet 54, 842–853 (2016). https://doi.org/10.1007/s10528-016-9761-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-016-9761-x