Abstract
Objective
To obtain novel glucoamylase from Daqu microbe.
Results
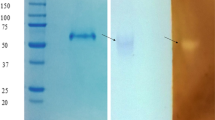
A dominant strain known as LZ2 with high activity of hydrolyzing starch was isolated from Luzhou Daqu, a Chinese traditional fermentation starter. The LZ2 was identified as Aspergillus oryzae by 18S rDNA sequence analysis. Glucoamylase from LZ2, named as GA-LZ2, was purified to homogeneity and showed a single band with expected molecular mass of 60 kD. The GA-LZ2 effectively degraded amylose, rice starch and wheat starch. Optimal temperature and pH value of enzyme were 60 °C and pH 4.0 respectively. The GA-LZ2 displayed significant thermal stability and pH stability at moderate temperature and low pH. Intriguingly, the thermostability was enhanced in the presence of starch. In addition, GA-LZ2 exhibited insensitivity to glucose, independence of metal ions and tolerance to organic solvents. The GA-LZ2 retained complete activity in the presence of 100 mM glucose and 5% ethanol and methanol.
Conclusion
Glucoamylase GA-LZ2 displayed broad substrate specificity, strong stability and tolerance, suggesting that GA-LZ2 carry potential for industrial application in bioethanol production.





Similar content being viewed by others
References
Aquino ACMM, Jorge JA, Terenzi HF, de Polizel MLTM (2011) Thermostable glucose-tolerant glucoamylase produced by the thermophilic fungus Scytalidium thermophilum. Folia Microbiol 46:11–16
Bagheri A, Khodarahmi R, Mostafaie A (2014) Purification and biochemical characterisation of glucoamylase from a newly isolated Aspergillus niger: Relation to starch processing. Food Chem 161:270–278
Benassi VM, Pasin TM, Facchini FDA, Jorge JA, de Lourdes Teixeira de Moraes Poliz M (2013) A novel glucoamylase activated by manganese and calcium produced in submerged fermentation by Aspergillus phoenicis. J Basic Microbiol 54: 333–339
Campos L, Felix R (1995) Purification and characterization of a glucoamylase from Humicola grisea. Appl Environ Microbiol 6:2436–2438
Coutinho PM, Reilly PJ (1994) Structure-function relationships in the catalytic and starch binding domains of glucoamylase. Protein Eng Des Sel 7:393–400
da Silva WB, Peralta RM (1998) Purification and characterization of a thermostable glucoamylase from Aspergillus fumigatus. Can J Microbiol 44:493–497
da Silva TM, Maller A, de Lima Damásio AR, Michelin M, Ward RJ, Hirata IY, Jorge JA, Terenzi HF, de Polizeli MLTM (2009) Properties of a purified thermostable glucoamylase from Aspergillus niveus. J Ind Microbiol Biotechnol 36:1439–1446
Ganghofner D, Kellermann J, Staudenbauer WL, Bronnenmeier K (1988) Purification and properties of an amylopullulanase, a glucoamylase, and an α-glucosidase in the amylolytic enzyme system of Thermoanaerobacterium thermosaccharolyticum. Biosci Biotechnol Biochem 62:302–308
Gill RK, Kaur J (2004) A thermostable glucoamylase from a thermophilic Bacillus sp.: characterization and thermostability. J Ind Microbiol Biotechnol 31:540–543
Hata Y, Kitamoto K, Gomi K, Kumagai C, Tamura G, Hara S (1991) The glucoamylase cDNA from Aspergillus oryzae: Its cloning, nucleotide sequence, and expression in Saccharomyces cerevisiae. Agric Biol Chem 55:941–949
Hua H, Luo H, Bai Y, Wang K, Niu C, Huang H, Shi P, Wang C, Yang P, Yao B (2014) A thermostable glucoamylase from Bispora sp. MEY-1 with stability over a broad pH range and significant starch hydrolysis capacity. PLoS ONE 9:e113581
Jang S-W, Kim J-S, Park J-B, Jung J-H, Park C-S, Shin WC, Ha S-J (2015) Characterization of the starch degradation activity from newly isolated Rhizopus oryzae WCS-1 and mixed cultures with Saccharomyces cerevisiae for efficient ethanol production from starch. Food Sci Biotechnol 24:1805–1810
Karim KMR, Husaini A, Sing NN, Tasnim T, Mohd Sinang F, Hussain H, Hossain MA, Roslan H (2019) Characterization and expression in Pichia pastoris of a raw starch degrading glucoamylase (GA2) derived from Aspergillus flavus NSH9. Protein Express Purif 164:1–10
Kim M-S, Park J-T, Kim Y-W, Lee H-S, Nyawira R, Shin H-S, Park C-S, Yoo S-H, Kim Y-R, Moon T-W, Park K-H (2004) Properties of a novel thermostable glucoamylase from the hyperthermophilic archaeon Sulfolobus solfataricus in relation to starch processing. Appl Environ Microbiol 70:3933–3940
Kumar S, Satyanarayana T (2003) Purification and kinetics of a raw starch-hydrolyzing, thermostable, and neutral glucoamylase of the thermophilic mold Thermomucor indicae-seudaticae. Biotechnol Progress 19:936–944
Kumar P, Satyanarayana T (2009) Microbial glucoamylases: characteristics and applications. Crit Rev Biotechnol 29:225–255
Li Z, Ji K, Dong W, Ye X, Wu J, Zhou J, Wang F, Chen Q, Fu L, Li S, Huang Y, Cui Z (2017) Cloning, heterologous expression, and enzymatic characterization of a novel glucoamylase GlucaM from Corallococcus sp. strain EGB. Protein Express Purif 129:122–127
Lincoln L, More VS, More SS (2019) Purification and biochemical characterization of extracellular glucoamylase from Paenibacillus amylolyticus strain. J Basic Microbiol 59:375
Michelin M, Ruller R, Ward RJ, Moraes LAB, Jorge JA, Terenzi HF, MdeLTM P (2007) Purification and biochemical characterization of a thermostable extracellular glucoamylase produced by the thermotolerant fungus Paecilomyces variotii. J Ind Microbiol Biotechnol 35:17–25
Ono K, Shigeta S, Oka S (1998) Effective purification of glucoamylase in Koji, a solid culture of Aspergillus oryzae on steamed rice, by affinity chromatography using an immobilized acarbose (BAY g-5421). Agric Biol Chem 52:1707–1714
Pasin TM, Benassi VM, Heinen PR, Damasio ARL, Cereia M, Jorge JA, Polizeli MLTM (2017) Purification and functional properties of a novel glucoamylase activated by manganese and lead produced by Aspergillus japonicus. Int J Biol Macromol 102:779–788
Spinelli LBB, de Polizeli MLTM, Terenzi HF, Jorge JA (1996) Biochemical characterization of glucoamylase from the hyperproducer exo-1 mutant strain of Neurospora erassa. FEMS Microbiol Lett 138:173–177
Tachibana S, Yasuda M (2007) Purification and characterization of heterogeneous glucoamylases from Monascus purpureus. Biosci Biotechnol Biochem 71:2573–2576
Xian L, Feng JX (2018) Purification and biochemical characterization of a novel mesophilic glucoamylase from Aspergillus tritici WZ99. Int J Biol Macromol 107:1122–1130
Xu QS, Yan YS, Feng JX (2016) Efficient hydrolysis of raw starch and ethanol fermentation: a novel raw starch-digesting glucoamylase from Penicillium oxalicum. Biotechnol Biofuels 9:216–220
Yang S, Choi SJ, Kwak J, Kim K, Seo M, Moon TW, Lee Y-W (2013) Aspergillus oryzae strains isolated from traditional Korean Nuruk: fermentation properties and influence on rice wine quality. Food Sci Biotechnol 22:425–432
Zheng Y, Xue Y, Zhang Y, Zhou C, Schwaneberg U, Ma Y (2010) Cloning, expression, and characterization of a thermostable glucoamylase from Thermoanaerobacter tengcongensis MB4. Appl Microbiol Biotechnol 87:225–233
Zheng XW, Tabrizi MR, Nout MJR, Han BZ (2011) Daqu—a traditional Chinese liquor fermentation starter. J I Brewing 117:82–90
Acknowledgements
This research was supported by the Luzhou Lao Jiao Scholarship (15ljsk10) and College student innovation fund (201510622060), College of Biotechnology, Sichuan University of Science and Technology.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, C., Yang, L., Luo, L. et al. Purification and characterization of glucoamylase of Aspergillus oryzae from Luzhou-flavour Daqu. Biotechnol Lett 42, 2345–2355 (2020). https://doi.org/10.1007/s10529-020-02956-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-020-02956-4