Abstract
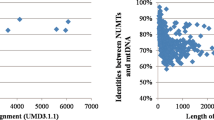
Through bacterial cloning, a non-specific product co-amplified in a previous whole mitochondrial genome study of Canis lupus familiaris was identified as part of a Numt on chromosome 29 of the dog. Even though further analysis confirmed the fidelity of the mitochondrial genome sequencing results, it still highlighted the risk of Numt contamination. A computer-based search of the dog’s nuclear genome for segments homologous to the mtDNA sequence revealed the extent of this risk. Over 150 Numts of various sizes were observed throughout all but two chromosomes, covering all positions of the mtDNA. One of the Numts on chromosome 11 even covered over 95 % of the entire dog mtDNA sequence. This comprehensive list of Numts was provided to assist researchers with the evaluation of dog mtDNA sequencing protocols for Numt co-amplification.

Similar content being viewed by others
References
Bensasson D, Feldman MW, Petrov DA (2003) Rates of DNA duplication and mitochondrial DNA insertion in the human genome. J Mol Evol 57:343–354. doi:10.1007/s00239-004-8301-1
Bravi CM, Parson W, Bandelt HJ (2006) Numts revisited. In: Bandelt HJ, Macaulay V, Richards M (eds) Human mitochondrial DNA and the evolution of Homo sapiens nucleic acids and molecular biology, vol 18. Springer, New York, pp 31–46
Calvignac S, Konecny L, Malard F, Douady CJ (2011) Preventing the pollution of mitochondrial datasets with nuclear mitochondrial paralogs (numts). Mitochondrion 11:246–254. doi:10.1016/j.mito.2010.10.004
Farrelly F, Butow RA (1983) Rearranged mitochondrial genes in the yeast nuclear genome. Nature 301:296–301
Gellissen G, Bradfield JY, White BN, Wyatt GR (1983) Mitochondrial DNA sequences in the nuclear genome of a locust. Nature 301:631–634
Hadler HI, Dimitrijevic B, Mahalingam R (1983) Mitochondrial DNA and nuclear DNA from normal rat liver have a common sequence. Proc Natl Acad Sci USA 80:6495–6499
Hazkani-Covo E, Zeller RM, Martin W (2010) Molecular poltergeists: mitochondrial DNA copies (numts) in sequenced nuclear genomes. PLoS Genet 6:e1000834. doi:10.1371/journal.pgen.1000834
Ishiguro N, Nakajima A, Horiuchi M, Shinagawa M (2002) Multiple nuclear pseudogenes of mitochondrial DNA exist in the canine genome. Mamm Genome 13:365–372. doi:10.1007/s00335-001-2139-2
Kim KS, Lee SE, Jeong HW, Ha JH (1998) The complete nucleotide sequence of the domestic dog (Canis familiaris) mitochondrial genome. Mol Phylogenet Evol 10:210–220. doi:10.1006/mpev.1998.0513
Koressaar T, Remm M (2007) Enhancements and modifications of primer design program Primer3. Bioinformatics 23:1289–1291. doi:10.1093/bioinformatics/btm091
Lin X et al (1999) Sequence and analysis of chromosome 2 of the plant Arabidopsis thaliana. Nature 402:761–768. doi:10.1038/45471
Lindblad-Toh K et al (2005) Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature 438:803–819. doi:10.1038/nature04338
Lopez JV, Yuhki N, Masuda R, Modi W, O’Brien SJ (1994) Numt, a recent transfer and tandem amplification of mitochondrial DNA to the nuclear genome of the domestic cat. J Mol Evol 39:174–190
Mourier T, Hansen AJ, Willerslev E, Arctander P (2001) The Human Genome Project reveals a continuous transfer of large mitochondrial fragments to the nucleus. Mol Biol Evol 18:1833–1837
Parr RL et al (2006) The pseudo-mitochondrial genome influences mistakes in heteroplasmy interpretation. BMC Genom 7:185. doi:10.1186/1471-2164-7-185
Richly E, Leister D (2004) NUMTs in sequenced eukaryotic genomes. Mol Biol Evol 21:1081–1084. doi:10.1093/molbev/msh110
Stupar RM, Lilly JW, Town CD, Cheng Z, Kaul S, Buell CR, Jiang J (2001) Complex mtDNA constitutes an approximate 620-kb insertion on Arabidopsis thaliana chromosome 2: implication of potential sequencing errors caused by large-unit repeats. Proc Natl Acad Sci USA 98:5099–5103. doi:10.1073/pnas.091110398
Tourmen Y, Baris O, Dessen P, Jacques C, Malthiery Y, Reynier P (2002) Structure and chromosomal distribution of human mitochondrial pseudogenes. Genomics 80:71–77
Tsuji J, Frith MC, Tomii K, Horton P (2012) Mammalian NUMT insertion is non-random. Nucleic Acids Res 40:9073–9088. doi:10.1093/nar/gks424
Tsuzuki T, Nomiyama H, Setoyama C, Maeda S, Shimada K (1983) Presence of mitochondrial-DNA-like sequences in the human nuclear DNA. Gene 25:223–229
Untergasser A, Cutcutache I, Koressaar T, Ye J, Faircloth BC, Remm M, Rozen SG (2012) Primer3–new capabilities and interfaces. Nucleic Acids Res 40:e115. doi:10.1093/nar/gks596
Verscheure S, Backeljau T, Desmyter S (2014) Dog mitochondrial genome sequencing to enhance dog mtDNA discrimination power in forensic casework. Forensic Sci Int Genet 12:60–68. doi:10.1016/j.fsigen.2014.05.001
Zhang DX, Hewitt GM (1996) Nuclear integrations: challenges for mitochondrial DNA markers. Trends Ecol Evol 11:247–251
Zhang Z, Schwartz S, Wagner L, Miller W (2000) A greedy algorithm for aligning DNA sequences. J Comput Biol 7:203–214. doi:10.1089/10665270050081478
Acknowledgments
We thank the staff members of the Institute of Legal Medicine at Innsbruck Medical University for scientific help. S. Verscheure is a PhD student at the University of Antwerp supported by a Grant from the Belgian Science Policy Office. This work was conducted in the framework of the FWO research community W0.009.11 N “Belgian Network for DNA Barcoding”.
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
This article does not contain any studies with human participants or animals performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Verscheure, S., Backeljau, T. & Desmyter, S. In silico discovery of a nearly complete mitochondrial genome Numt in the dog (Canis lupus familiaris) nuclear genome. Genetica 143, 453–458 (2015). https://doi.org/10.1007/s10709-015-9844-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-015-9844-3