Abstract
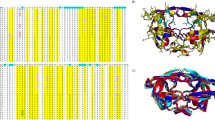
Under drug selection pressure, emerging mutations render HIV-1 protease drug resistant, leading to the therapy failure in anti-HIV treatment. It is known that nine substrate cleavage site peptides bind to wild type (WT) HIV-1 protease in a conserved pattern. However, how the multidrug-resistant (MDR) HIV-1 protease binds to the substrate cleavage site peptides is yet to be determined. MDR769 HIV-1 protease (resistant mutations at residues 10, 36, 46, 54, 62, 63, 71, 82, 84, and 90) was selected for present study to understand the binding to its natural substrates. MDR769 HIV-1 protease was co-crystallized with nine substrate cleavage site hepta-peptides. Crystallographic studies show that MDR769 HIV-1 protease has an expanded substrate envelope with wide open flaps. Furthermore, ligand binding energy calculations indicate weaker binding in MDR769 HIV-1 protease-substrate complexes. These results help in designing the next generation of HIV-1 protease inhibitors by targeting the MDR HIV-1 protease.





Similar content being viewed by others
Abbreviations
- MDR:
-
Multi-drug resistance
- HIV:
-
Human immunodeficiency virus
- WT:
-
Wild type
- HAART:
-
Highly active antiretroviral therapy
- PR:
-
Protease
- MA:
-
Matrix
- CA:
-
Capsid
- NC:
-
Nucleocapsid
- RT:
-
Reverse transcriptase
- IN:
-
Integrase
- RH:
-
RNase H
- RMSD:
-
Root mean square deviation
References
Altman MD, Ali A, Reddy GS, Nalam MN, Anjum SG, Cao H, Chellappan S, Kairys V, Fernandes MX, Gilson MK, Schiffer CA, Rana TM, Tidor B (2008) J Am Chem Soc 130:6099–6113
Bally F, Martinez R, Peters S, Sudre P, Telenti A (2000) AIDS Res Hum Retroviruses 16:1209–1213
Barbaro G, Lucchini A, Barbarini G (2005) Minerva Cardioangiol 53:153–154
Bartlett JA, DeMasi R, Quinn J, Moxham C, Rousseau F (2001) AIDS 15:1369–1377
Chellappan S, Kairys V, Fernandes MX, Schiffer C, Gilson MK (2007) Proteins 68:561–567
Chellappan S, Kiran Kumar Reddy GS, Ali A, Nalam MN, Anjum SG, Cao H, Kairys V, Fernandes MX, Altman MD, Tidor B, Rana TM, Schiffer CA, Gilson MK (2007) Chem Biol Drug Des 69:298–313
Clavel F, Hance AJ (2004) N Engl J Med 350:1023–1035
Condra JH, Schleif WA, Blahy OM, Gabryelski LJ, Graham DJ, Quintero JC, Rhodes A, Robbins HL, Roth E, Shivaprakash M (1995) Nature 374:569–571
Croteau G, Doyon L, Thibeault D, McKercher G, Pilote L, Lamarre D (1997) J Virol 71:1089–1096
Emsley P, Cowtan K (2004) Acta Crystallogr D Biol Crystallogr 60:2126–2132
Grabar S, Pradier C, Le Corfec E, Lancar R, Allavena C, Bentata M, Berlureau P, Dupont C, Fabbro-Peray P, Poizot-Martin I, Costagliola D (2000) AIDS 14:141–149
Gulick RM, Mellors JW, Havlir D, Eron JJ, Meibohm A, Condra JH, Valentine FT, McMahon D, Gonzalez C, Jonas L, Emini EA, Chodakewitz JA, Isaacs R, Richman DD (2000) Ann Intern Med 133:35–39
Holzgrabe U (2004) Pharm Unserer Zeit 33:160
Krissinel E, Henrick K (2004) Acta Crystallogr D Biol Crystallogr 60:2256–2268
Krissinel E, Henrick K (2007) J Mol Biol 372:774–797
Laemmli UK (1970) Nature 227:680–685
Lamzin VS, Wilson KS (1993) Acta Crystallogr D Biol Crystallogr 49:129–147
Li F, Goila-Gaur R, Salzwedel K, Kilgore NR, Reddick M, Matallana C, Castillo A, Zoumplis D, Martin DE, Orenstein JM, Allaway GP, Freed EO, Wild CT (2003) Proc Natl Acad Sci USA 100:13555–13560
Li F, Zoumplis D, Matallana C, Kilgore NR, Reddick M, Yunus AS, Adamson CS, Salzwedel K, Martin DE, Allaway GP, Freed EO, Wild CT (2006) Virology 356:217–224
Logsdon BC, Vickrey JF, Martin P, Proteasa G, Koepke JI, Terlecky SR, Wawrzak Z, Winters MA, Merigan TC, Kovari LC (2004) J Virol 78:3123–3132
Martin DE, Blum R, Wilton J, Doto J, Galbraith H, Burgess GL, Smith PC, Ballow C (2007) Antimicrob Agents Chemother 51:3063–3066
Martin P, Vickrey JF, Proteasa G, Jimenez YL, Wawrzak Z, Winters MA, Merigan TC, Kovari LC (2005) Structure 13:1887–1895
McMichael AJ, Hanke T (2003) Nat Med 9:874–880
Murshudov GN, Vagin AA, Dodson EJ (1997) Acta Crystallogr D Biol Crystallogr 53:240–255
Nalam MN, Ali A, Altman MD, Reddy GS, Chellappan S, Kairys V, Ozen A, Cao H, Gilson MK, Tidor B, Rana TM, Schiffer CA (2010) J Virol 84:5368–5378
Natarajan V, Bosche M, Metcalf JA, Ward DJ, Lane HC, Kovacs JA (1999) Lancet 353:119–120
Palella FJ, Delaney KM, Moorman AC, Loveless MO, Fuhrer J, Satten GA, Aschman DJ, Holmberg SD (1998) N Engl J Med 338:853–860
Pettit SC, Everitt LE, Choudhury S, Dunn BM, Kaplan AH (2004) J Virol 78:8477–8485
Prabu-Jeyabalan M, King NM, Nalivaika EA, Heilek-Snyder G, Cammack N, Schiffer CA (2006) Antimicrob Agents Chemother 50:1518–1521
Prabu-Jeyabalan M, Nalivaika EA, King NM, Schiffer CA (2004) J Virol 78:12446–12454
Prabu-Jeyabalan M, Nalivaika EA, Romano K, Schiffer CA (2006) J Virol 80:3607–3616
Prabu-Jeyabalan M, Nalivaika E, Schiffer CA (2000) J Mol Biol 301:1207–1220
Prabu-Jeyabalan M, Nalivaika E, Schiffer CA (2002) Structure 10:369–381
Roberts JD, Preston BD, Johnston LA, Soni A, Loeb LA, Kunkel TA (1989) Mol Cell Biol 9:469–476
Robinson LH, Myers RE, Snowden BW, Tisdale M, Blair ED (2000) AIDS Res Hum Retroviruses 16:1149–1156
Schooley RT, Mellors JW (2007) J Infect Dis 195:770–772
Sukasem C, Churdboonchart V, Sukeepaisarncharoen W, Piroj W, Inwisai T, Tiensuwan M, Chantratita W (2008) Int J Antimicrob Agents 31:277–281
Takeuchi Y, Nagumo T, Hoshino H (1988) J Virol 62:3900–3902
Tie Y, Boross PI, Wang YF, Gaddis L, Liu F, Chen X, Tozser J, Harrison RW, Weber IT (2005) FEBS J 272:5265–5277
Vagin AA, Steiner RA, Lebedev AA, Potterton L, McNicholas S, Long F, Murshudov GN (2004) Acta Crystallogr D Biol Crystallogr 60:2184–2195
Vaguine AA, Richelle J, Wodak SJ (1999) Acta Crystallogr D Biol Crystallogr 55:191–205
Vickrey JF, Logsdon BC, Proteasa G, Palmer S, Winters MA, Merigan TC, Kovari LC (2003) Protein Expr Purif 28:165–172
Wallace AC, Laskowski RA, Thornton JM (1995) Protein Eng 8:127–134
Weber J, Grosse F (1989) Nucleic Acids Res 17:1379–1393
Zhang YM, Imamichi H, Imamichi T, Lane HC, Falloon J, Vasudevachari MB, Salzman NP (1997) J Virol 71:6662–6670
Zhou J, Chen CH, Aiken C (2004) Retrovirology 1:15
Acknowledgments
This research was supported by the National Institutes of Health grant AI65294 and a grant from the American Foundation for AIDS Research (106457-34-RGGN).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, Z., Wang, Y., Brunzelle, J. et al. Nine Crystal Structures Determine the Substrate Envelope of the MDR HIV-1 Protease. Protein J 30, 173–183 (2011). https://doi.org/10.1007/s10930-011-9316-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-011-9316-2