Abstract
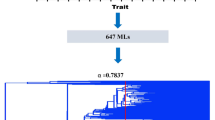
By studying the regulatory mechanism by which rice tillers are controlled, many rice mutants were described as providing important insights into the investigation of tillering control. In this study, we surveyed a new rice mutant produced by ethyl methane sulfonate induction, designated as Oryza sativa low-tiller 1 (oslt1), which had been planted to the fourth generation and proved to stably inherit the low-tillering trait. The statistical analysis of the number of tillers in oslt1 showed that there was a significant lack of tillering in the mutant lines, with the average number being 2.33 as opposed to 8.00 in wild-type plants. The data from a series of crossed populations, including F2 populations produced by selfing of the offspring of oslt1 × ptb and backcross populations of the F1 generation × oslt1, indicated that a single recessive gene controlled the low-tillering trait. To locate the recessive gene, the crossed population in the F2 generation from two parents, oslt1 and Peiai 64, was constructed and the population was used for bulked segregate analysis based on specific locus amplified fragment-sequencing (SLAF-seq) method. The results showed that approximately 100 differential markers were procured by analyzing 7437 polymorphic SLAFs and 1 primary region associated with low-tillering characteristics located on Chr12. In the associated regions, there were 34 candidate genes of 0.24 Mb that might be related to the number of low tillers in oslt1 plants. All of these results are necessary for further fine mapping for this low tillering-related gene. In addition, these identified SLAF-tags will be valuable for marker-assisted selection in rice breeding.






Similar content being viewed by others
References
Abe A, Kosugi S, Yoshida K, Natsume S, Takagi H, Kanzaki H, Matsumura H, Yoshida K, Mitsuoka C, Tamiru M, Innan H, Cano L, Kamoun S, Terauchi R (2012) Genome sequencing reveals agronomically important loci in rice using MutMap. Nat Biotechnol 30:174–178
Altshuler D, Pollara VJ, Cowles CR, Van Etten WJ, Baldwin J, Linton L, Lander ES (2000) An SNP map of the human genome generated by reduced representation shotgun sequencing. Nature 407:513–516
Andolfatto P, Davison D, Erezyilmaz D, Hu TT, Mast J, Sunayama Morita T, Stern DL (2011) Multiplexed shotgun genotyping for rapid and efficient genetic mapping. Genome Res 21:610–617
Arite T, Iwata H, Ohshima K, Maekawa M, Nakajima M, Kojima M, Sakakibar H, Kyozuka J (2007) DWARF10, an RMS1/MAX4/DAD1 ortholog, controls lateral bud outgrowth in rice. Plant J 51:1019–1029
Arite T, Umehara M, Ishikawa S, Hanada A, Maekawa M, Yamaguchi S, Kyozuka J (2009) D14, a strigolactone–insensitive mutant of rice, shows an accelerated outgrowth of tillers. Plant Cell Physiol 50:1416–1424
Booker J, Sieberer T, Wright W, Williamson L, Willett B, Stirnberg P, Turnbull C, Srinivasan M, Goddard P, Leyser O (2005) MAX1 encodes a cytochrome P450 family member that acts downstream of MAX3/4 to produce a carotenoid-derived branch-inhibiting hormone. Dev Cell 8:443–449
Chen SQ, Huang ZF, Dai Y, Qin SW, Gao YY, Zhang LL, Gao Y, Chen JM (2013) The development of 7E chromosome–specific molecular markers for Thinopyrum elongatum based on SLAF-seq technology. PLoS One 8:e65122
Chen W, Yao JB, Chu L, Li Y, Guo XG, Zhang YS (2014) The development of specific SNP markers for chromosome 14 in cotton using next-generation sequencing. Plant Breed 133:256–261
Davey JW, Blaxter ML (2010) RADSeq: next-generation population genetics. Brief Funct Genomics 9:416–423
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Elshire RJ, Glaubitz JC, Sun Q, Poland JA, Kawamoto K, Buckler ES, Mitchell SE (2011) A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS One 6:e19379
Gachomo EW, Jimenez-Lopez JC, Baptiste LJ, Kotchoni SO (2014) GIGANTUS1 (GTS1), a member of Transducin/WD40 protein superfamily, controls seed germination, growth and biomass accumulation through ribosome-biogenesis protein interactions in Arabidopsis thaliana. BMC Plant Biol 14:1–17
Gao ZY, Qian Q, Liu XH, Yan MX, Feng Q, Dong GJ, Liu J, Han B (2009) Dwarf 88, a novel putative esterase gene affecting architecture of rice plant. Plant Mol Biol 71:265–276
Guo SY, Xu YY, Liu HH, Mao ZW, Zhang C, Ma Y, Zhang QR, Meng Z, Chong K (2013) The interaction between OsMADS57 and OsTB1modulates rice tillering via DWARF14. Nat Commun 4:1566–1577
Hamiaux C, Drummond RS, Janssen BJ, Ledger SE, Cooney JM, Newcomb RD, Snowden KC (2012) DAD2 is an alpha/beta hydrolase likely to be involved in the perception of the plant branching hormone, strigolactone. Curr Biol 22:1–5
Huang XH, Feng Q, Qian Q, Zhao Q, Wang L, Wang A, Guan JP, Fan DL, Weng QJ, Huang T, Dong GJ, Sang T, Han B (2009) High-throughput genotyping by whole-genome resequencing. Genome Res 19:1068–1076
Ito S, Kitahata N, Umehara M, Hanada A, Kato A, Ueno K, Mashiguchi K, Kyozuka J, Yoneyama K, Yamaguchi S, Asami T (2010) A new lead chemical for strigolactone biosynthesis inhibitors. Plant Cell Physiol 51:1143–1150
Kent WJ (2002) BLAT—the BLAST-like alignment tool. Genome Res 12:656–664
Li XY, Qian Q, Fu ZM, Wang YH, Xiong GS, Zeng DL, Wang XQ, Liu XF, Teng S, Hiroshi F, Yuan M, Luo D, Han B, Li JY (2003) Control of tillering in rice. Nature 422(6932):618–621
Liang WH, Shang F, Lin QT, Lou C, Zhang J (2014) Tillering and panicle branching genes in rice. Gene 537:1–5
Lin H, Wang RX, Qian Q, Yan MX, Meng XB, Fu ZM, Yan CY, Jiang B, Su Z, Li JY, Wang YH (2009) DWARF27 an iron-containing protein required for the biosynthesis of strigolactones, regulates rice tiller bud outgrowth. Plant Cell 21:1512–1525
Lin QB, Wang D, Dong H, Gu SH, Cheng ZJ, Gong J, Qin RZ, Jiang L, Li G, Wang JL, Wu FQ, Guo XP, Zhang X, Lei CL, Wang HY, Wan JM (2012) Rice APC/C(TE) controls tillering by mediating the degradation of MONOCULM 1. Nat Commun 20:752
Liu W, Kohlen W, Lillo A, Op den Camp R, Ivanov S, Hartog M, Limpens E, Jamil M, Smaczniak C, Kaufmann K, Yang WC, Hooiveld GJ, Charnikhova T, Bouwmeester HJ, Bisseling T, Geurtsa R (2011) Strigolactone biosynthesis in Medicago truncatula and rice requires the symbiotic GRAS-type transcription factors NSP1 and NSP2. Plant Cell 23:3853–3865
Minakuchi K, Kameoka H, Yasuno N, Umehara M, Luo L, Kobayashi K, Hanada A, Ueno K, Asami T, Yamaguchi S, Kyozuka J (2010) FINE CULM1 (FC1) works downstream of strigolactones to inhibit the outgrowth of axillary buds in rice. Plant Cell Physiol 51:1127–1135
Rabiger DS, Drews GN (2013) MYB64 and MYB119 are required for cellularization and differentiation during female gametogenesis in Arabidopsis thaliana. Plos Genetics 9:e1003783–e1003783
Sorefan K, Booker J, Haurogné K, Goussot M, Bainbridge K, Foo E, Chatfield S, Ward S, Beveridge C, Rameau C, Leyser O (2003) MAX4 and RMS1 are orthologous dioxygenase-like genes that regulate shoot branching in Arabidopsis and pea. Genes Dev 17:1469–1474
Souer E, Houwelingen AV, Kloos D, Mol J, Koes R (1996) The no apical meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 85:159–170
Stirnberg P, van de Sande K, Leyser HMO (2002) MAX1 and MAX2 control shoot lateral branching in Arabidopsis. Development 129:1131–1141
Sun FL, Zhang WP, Xiong GS, Yan MX, Qian Q, Li JY, Wang YH (2010) Identification and functional analysis of the MOC1 interacting protein. J Genet Genomics 37:69–77
Sun XW, Liu DY, Zhang XF, Li WB, Liu H, Hong WG, Jiang CB, Guan N, Ma CX, Zeng HP, Xu CH, Song J, Huang L, Wang CM, Shi JJ, Wang R, Zheng XH, Lu CY, Wang XW, Zheng HK (2013) SLAF-seq: an efficient method of large–scale de novo SNP discovery and genotyping using high-throughput sequencing. PLoS ONE 8:e58700
Tabuchi H, Zhang Y, Hattori S, Omae M, Shimizu–Sato S, Oikawa T, Qian Q, Nishimura M, Kitano H, Xie H, Fang XH, Yoshida H, Kyozuka J, Chen F, Sato Y (2011) LAX PANICLE2 of rice encodes a novel nuclear protein and regulates the formation of axillary meristems. Plant Cell 23:3276–3287
Takeda T, Suwa Y, Suzuki M, Kitano H, Ueguchi–Tanaka M, Ashikari M, Matsuoka M, Ueguchi C (2003) The OsTB1 gene negatively regulates lateral branching in rice. Plant J 33:513–520
Wang YH, Li JY (2010) Branching in rice. Plant Biol 14:1–6
Xia C, Chen LL, Rong TZ, Li R, Xiang Y, Wang P, Liu CH, Dong XQ, Liu B, Zhao D (2015) Identification of a new maize inflorescence meristem mutant and association analysis using SLAF–seq method. Euphytica 202:35–44
Xu C, Wang YH, Yu YC, Duan JB, Liao ZG, Xiong GS, Meng XB, Liu GF, Qian Q, Li JY (2012) Degradation of MONOCULM 1 by APC/C(TAD1) regulates rice tillering. Nat Commun 20:750
Yan HF, Saika H, Maekawa M, Takamure I, Tsutsumi N, Kyozuka J, Nakazono M (2007) Rice tillering dwarf mutant dwarf3 has increased leaf longevity during darkness–induced senescence or hydrogen peroxide-induced cell death. Genes Genet Syst 82:361–366
Zhang SY, Li G, Fang J, Chen WQ, Jiang HP, Zou JH, Liu X, Zhao XF, Li XB, Chu CG, Xie Q, Jiang XN, Zhu LH (2010) The interactions among DWARF10, auxin and cytokinin underlie lateral bud outgrowth in rice. Journal of Integrative Plant Biol 52:626–638
Zhang ZY, Li JJ, Yao GX, Zhang HL, Dou HJ, Shi HL, Sun XM, Li ZC (2011) Fine mapping and cloning of the grain number per-panicle gene (Gnp4) on chromosome 4 in rice (Oryza sativa L). J Integr Agr 10:1825–1833
Zou JH, Chen ZX, Zhang SY, Zhang WP, Jiang GH, Zhao XF, Zhai WX, Pan XB, Zhu LH (2005) Characterizations and fine mapping of a mutant gene for high tillering and dwarf in rice (Oryza sativa L). Planta 222:604–612
Zou JH, Zhang SY, Zhang WP, Li G, Chen ZX, Zhai WX, Zhao XF, Pan XB, Xie Q, Zhu LH, The rice (2006) HIGH–TILLERING DWARF1 encoding an ortholog of Arabidopsis MAX3 is required for negative regulation of the outgrowth of axillary buds. Plant J 48:687–698
Acknowledgments
This work was supported by grants from the Genetically Modified Organisms Breeding Major Projects of China (2014ZX08010-003 and 2016ZX08010-003) and the Guizhou Science and Technology Major Project (2012 6005).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig 1
The aa and ab ratio on each chromosome. (DOCX 4953 kb)
Supplementary Fig 2
Segregation of SSR Marker (RM 27601) in oslt1 × Peiai 64 F2 populations. (DOCX 102 kb)
Supplementary Table 1
The information of associated SLAF Markers: SLAF16002, SLAF16015 and SLAF16030. (DOCX 17 kb)
Supplementary Table 2
The Annotation and Regulation of Candidate Genes. (DOCX 28 kb)
Rights and permissions
About this article
Cite this article
Li, Y., Zeng, XF., Zhao, YC. et al. Identification of a New Rice Low-Tiller Mutant and Association Analyses Based on the SLAF-seq Method. Plant Mol Biol Rep 35, 72–82 (2017). https://doi.org/10.1007/s11105-016-1002-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-016-1002-2