Abstract
In many epidemiologic models, a disease is assumed to spread along a contact network. We aim to infer this network, in addition to the epidemiologic model parameters, from the binary status of individuals observed throughout time. We perform an exact evaluation of the probability for each edge to be part of the network by using the matrix-tree theorem on the set of vertices made of the individual status at all times. This leads to a computational complexity of order \({\mathcal {O}}(mn^2)\), where n is the number of individuals and m the length of the time series. Simulations are provided to demonstrate the efficiency of the proposed method, and it is applied on data concerning seed choices by farmers in India and on data on a measles outbreak.





Similar content being viewed by others
References
Barabási, A.-L., Albert, R.: Emergence of scaling in random networks. Science 286(5439), 509–512 (1999)
Barbillon, P., Thomas, M., Goldringer, I., Hospital, F., Robin, S.: Network impact on persistence in a finite population dynamic diffusion model: application to an emergent seed exchange network. J. Theor. Biol. 365, 365–376 (2015)
Boyd, R., Richerson, P.J., Henrich, J.: The cultural niche: why social learning is essential for human adaptation. Proc. Natl. Acad. Sci. 108(Supplement 2), 10918–10925 (2011)
Brauer, F., Castillo-Chavez, C., Castillo-Chavez, C.: Mathematical Models in Population Biology and Epidemiology, vol. 40. Springer, Berlin (2012)
Britton, T., O’Neill, P.D.: Bayesian inference for stochastic epidemics in populations with random social structure. Scand. J. Stat. 29(3), 375–390 (2002)
Chaiken, S.: A combinatorial proof of the all minors matrix tree theorem. SIAM J. Algebr. Discrete Methods 3(3), 319–329 (1982)
Chow, C., Liu, C.: Approximating discrete probability distributions with dependence trees. IEEE Trans. Inf. Theory IT–14(3), 462–467 (1968)
David, P.A.: Path dependence: a foundational concept for historical social science. Cliometrica 1(2), 91–114 (2007)
Dempster, A.P., Laird, N.M., Rubin, D.B.: Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. B. 39, 1–38 (1977)
Erdős, P., Rényi, A.: On random graphs. I. Publicationes Mathematicae Debrecen. 6, 290–297 (1959)
Flachs, A., Stone, G.D., Shaffer, C.: Mapping knowledge: gis as a tool for spatial modeling of patterns of warangal cotton seed popularity and farmer decision-making. Hum. Ecol. 45(2), 143–159 (2017)
Galab, S., Revathi, E., Reddy, P.P.: Farmers’ suicides and unfolding agrarian crisis in Andhra Pradesh. Agrar. Crisis India. 164–198 (2009). https://doi.org/10.1093/acprof:oso/9780198069096.001.0001
Gomez-Rodriguez, M., Balduzzi, D. and Schölkopf, B.: Uncovering the temporal dynamics of diffusion networks. (2011) Technical Report, arXiv:1508.00286
Gomez-Rodriguez, M., Leskovec, J., Krause, A.: Inferring networks of diffusion and influence. ACM Trans. Knowl. Discov. Data (TKDD) 5(4), 21 (2012)
Griliches, Z.: Hybrid corn revisited: a reply. Econom. J. Econ. Soc. 48, 1463–1465 (1980)
Groendyke, C., Welch, D., Hunter, D.R.: Bayesian inference for contact networks given epidemic data. Scand. J. Stat. 38(3), 600–616 (2011)
Groendyke, C., Welch, D., Hunter, D.R.: A network-based analysis of the 1861 hagelloch measles data. Biometrics 68(3), 755–765 (2012)
Gutierrez, A.P., Ponti, L., Herren, H.R., Baumgärtner, J., Kenmore, P.E.: Deconstructing Indian cotton: weather, yields, and suicides. Environ. Sci. Eur. 27(1), 12 (2015)
Henrich, J.: Cultural transmission and the diffusion of innovations: adoption dynamics indicate that biased cultural transmission is the predominate force in behavioral change. Am. Anthropol. 103(4), 992–1013 (2001)
Herring, R.J., Rao, N.C.: On the’failure of bt cotton’: analysing a decade of experience. Econ. Polit. Wkly. 47, 45–53 (2012)
Kirshner, S.: Learning With Tree-averaged Densities and Distributions. In: NIPS, pp. 761–768 (2007)
Meilă, M., Jaakkola, T.: Tractable Bayesian learning of tree belief networks. Stat. Comput. 16(1), 77–92 (2006)
Myers, S., Leskovec, J.: On the convexity of latent social network inference. In: Proceedings of the 23rd International Conference on Neural Information Processing System, vol. 2, pp. 1741–1749 (2010). http://papers.nips.cc/paper/4113-on-the-convexity-of-latent-social-network-inference
Neal, P.J., Roberts, G.O.: Statistical inference and model selection for the 1861 hagelloch measles epidemic. Biostatistics 5(2), 249–261 (2004)
Oesterle, H.: Statistische Reanalyse einer Masernepidemie 1861 in Hagelloch. Ph.D. thesis, uitgever niet vastgesteld (1993)
Pfeilsticker, A.: Beiträge zur Pathologie der Masern mit besonderer Berücksichtigung der statistischen Verhältnisse
Ray, J., Marzouk, Y.M.: A Bayesian method for inferring transmission chains in a partially observed epidemic. In: Proceedings of the Joint Statistical Meeting (2008)
Stone, G.D.: Agricultural deskilling and the spread of genetically modified cotton in warangal. Curr. Anthropol. 48(1), 67–103 (2007)
Stone, G.D.: Towards a general theory of agricultural knowledge production: environmental, social, and didactic learning. Cult. Agric. Food Environ. 38(1), 5–17 (2016)
Stone, G.D., Flachs, A., Diepenbrock, C.: Rhythms of the herd: long term dynamics in seed choice by Indian farmers. Technol. Soc. 36, 26–38 (2014)
Wang, Y., Chakrabarti, D., Wang, C. and Faloutsos, C.: Epidemic spreading in real networks: an eigenvalue viewpoint. In: 22nd International Symposium on Reliable Distributed Systems, 2003. Proceedings, pp. 25–34. IEEE (2003)
Welch, D., Bansal, S., Hunter, D.R.: Statistical inference to advance network models in epidemiology. Epidemics 3(1), 38–45 (2011)
Yang, L.-X., Draief, M., Yang, X.: The impact of the network topology on the viral prevalence: a node-based approach. PLoS ONE 10(7), e0134507 (2015)
Acknowledgements
The authors sincerely thank the Associate Editor and the two Reviewers for their careful reading, their comments and their advice, which obviously contributed to improve this manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This work has been partially supported by United States Department of Education Jacob K. Javits Fellowship, the National Geographic Society Young Explorer’s Grant 9304-13, the John Templeton Foundation (Glenn Davis Stone PI), Washington University in St. Louis and MIRES funded by the Applied Mathematics and Informatics department of INRA.
A Appendix
A Appendix
Definition 1
(Cofactor matrix) Consider a \(p \times p\) square matrix A. For any pair \(1 \le u, v \le p\), the cofactor \(C_{u, v}\) is defined as \(C_{u, v} := (-1)^{u+v} |A^{u, v}|\), where \(A^{u, v}\) stands for the matrix A deprived from its u-th row and v-th column. The cofactor matrix of A is the \(p \times p\) square matrix C, with general term \(C_{u, v}\).
1.1 A.1 Bayes inference for the parameters
Assume that parameters e and c have independent Beta prior distributions:
In a Bayesian setting, the likelihood given in (5) corresponds to the conditional distribution P(Y|r) so that the joint distribution of (Y, r) is given by
The marginal likelihood of the data Y is obtained by integrating over c and e to get
Both P(Y, r) and P(Y) factorize with respect to e and c, e and c are therefore conditionally independent given Y. The posterior distribution of e is simply \(\text {Beta}\):
whereas the posterior distribution of c does not have a closed-form expression:
but can be easily sampled via Monte Carlo or importance sampling, or numerically integrated. Observe that, in this setting, the weights \(\beta _{ij}\) act as hyper-parameters.
1.2 A.2 EM inference
If T is considered as a latent variable, the proposed model becomes an incomplete data model, for which maximum likelihood inference can be carried out via the EM algorithm (Dempster et al. 1977). Because the distribution of T is parameterized by \(\beta \), the parameters to be inferred become \(\theta = (\beta , c, e)\). To use the EM algorithm, we first need to write the complete log likelihood:
where \(N_{ij}(T) = \sum _t {\mathbb {I}}\{(i, j) \in T^t\}\). Then, we need the conditional expectation of this complete log likelihood \({\mathbb {E}}_{\theta ^q} \left[ \log P_\theta (Y, T) | Y \right] \), that is
E step The E step requires to compute the conditional edge probability
where \(C_{ij}^t\) is computed in the same way as C in Eq. (6), setting the term \(\psi _{ij}^t\) to 0. As a consequence, we get \(C_{ij}^t / C = \left( \psi _{+j}^t - \psi _{ij}^t \right) / \psi _{+j}^t\), so
This provides us with the conditional expected counts:
M step The parameter estimates are updated by maximizing the conditional expectation \( Q(\theta |\theta ^q) := {\mathbb {E}}_{\theta ^q} \left[ \log P_\theta (Y, T) | Y \right] . \) The terms of \(Q(\theta |\theta ^q)\) depending on e are
because, for all j and t, \(\sum _i P_{\theta ^q}\{(i, j) \in T^t | Y\} = 1\). (Each node has one and only one parent at each time.) Notice that the involved quantities do not depend on h, so we get the same estimate as in Sect. 3.2: \({\widehat{e}} = M_{10} / (M_{10} + M_{11})\).
The terms of \(Q(\theta |\theta ^q)\) depending on c are
where \(E^q_{101} = \sum _{i, j, t} P_{\theta ^q}\{(i, j) \in T^t | Y\} Y_i^{t} (1-Y_j^{t}) Y_j^{t+1}\) and \(E^q_{100} = \sum _{i, j, t} P_{\theta ^q}\{(i, j) \in T^t | Y\} Y_i^{t} (1- Y_j^{t}) (1-Y_j^{t+1})\), so at iteration q, the estimate of c is updated to
Then we have to maximize \(Q(\theta |\theta ^q)\) w.r.t. \(\beta \), which is equivalent to maximizing \({\mathbb {E}}_{\theta ^q}[N_{ij}(T) | Y] \log \beta _{ij} - \log B\), where \( \sum _{i, j} {\mathbb {E}}_{\theta ^q}[N_{ij}(T) | Y] \log \beta _{ij} - m \sum _j \log \beta _{+j} \) is maximal for
which leads to the update formula
1.3 Estimates of the parameters e and c for the simulation study
Figures 6 and 7 display, respectively, the results on the estimates of the parameters e and c. The inference for e is mainly satisfactory except for some configurations where this parameter is underestimated in the multiwave setting. This appears in configurations with the lowest value for c (\(c=0.1\)). In these configurations, a total extinction of the epidemic may happen (the lower c, the more likely the total extinction). Then the underestimation of e seems to be a consequence of our simulation choice which enforces that no total extinction occurs in any of the waves. This also leads to overestimate c for the very same configurations while for other configurations, the estimations of c suffers from a negative bias. This may be due to the fact that the edges were sampled in an underlying network instead among all possible pairs of nodes. Nevertheless, these estimates are still satisfactory since they do not prevent us from recovering the edges which is our main goal.
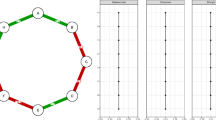
Boxplots of estimates of e. Each boxplot is obtained from 100 simulations. In the simulations, the extinction and infection parameters were set to \(e=0.05\) and \(c\in \{0.1,0.2,0.4\}\). The number of nodes in the network was 20, networks were simulated with Erdős–Rényi (ER) and preferential attachment (PA) topologies, and the density of the network was set to \(d\in \{0.1,0.2,0.4\}\). A simulation was always initialized with a unique node in state 1. A uniwave simulation (uW) consists in a wave of \(m=200\) time steps. A multiwave simulation (mW) consists in \(H=10\) waves of \(m=20\) time steps. For example, PA_uW corresponds to a uniwave simulation over a preferential attachment network
Boxplots of estimates of c. Each boxplot is obtained from 100 simulations. In the simulations, the extinction and infection parameters were set to \(e=0.05\) and \(c\in \{0.1,0.2,0.4\}\). The number of nodes in the network was 20, networks were simulated with Erdős–Rényi (ER) and preferential attachment (PA) topologies, and the density of the network was set to \(d\in \{0.1,0.2,0.4\}\). A simulation was always initialized with a unique node in state 1. A uniwave simulation (uW) consists in a wave of \(m=200\) time steps. A multiwave simulation (mW) consists in \(H=10\) waves of \(m=20\) time steps. For example, PA_uW corresponds to a uniwave simulation over a preferential attachment network
Rights and permissions
About this article
Cite this article
Barbillon, P., Schwaller, L., Robin, S. et al. Epidemiologic network inference. Stat Comput 30, 61–75 (2020). https://doi.org/10.1007/s11222-019-09865-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11222-019-09865-1