Abstract
Viral-encoded ATPase can act as a part of molecular motor in genome packaging of DNA viruses, such as vaccinia virus and adenovirus, by ATP hydrolysis and interaction with DNA. Poxviral ATPase (also called A32) is involved in genomic double-stranded DNA (dsDNA) encapsidation, and inhibition of the expression of A32 causes formation of immature virions lacking viral DNA. However, the role of A32 in goatpoxvirus genome packaging and its dsDNA binding property are not known. In this study, purified recombinant goatpoxvirus A32 protein (rA32) was examined for its dsDNA binding property as well as the effect of dsDNA on ATP hydrolysis. We found that rA32 could bind dsDNA, and its ATPase activity was significant increased with dsDNA binding. Effects of magnesium and calcium ions on ATP hydrolysis were investigated also. The ATPase activity was dramatically enhanced by dsDNA in the presence of Mg2+; in contrast, ATPase function was not altered by Ca2+. Furthermore, the enzyme activity of rA32 was completely blocked by Zn2+. Regarding DNA–protein interaction, the rA32-ATP-Mg2+ showed lower dsDNA binding affinity than that of rA32-ATP-Ca2+. The DNA–protein binding was stronger in the presence of zinc ion. Our results implied that A32 may play a role in viral genome encapsidation and DNA condensation.
Similar content being viewed by others
Introduction
Goatpoxvirus (GTPV) belongs to the Capripoxvirus genus, has a linear double-stranded DNA (dsDNA) genome of ~154 kbps, and is bigger than other members of Poxviridae with a virus particle size of 300 nm × 270 nm × 200 nm [1–3]. The viral life cycle of GTPV involves DNA replication, transcription, translation, and virion assembly in the cytoplasm [2, 4].
Poxvirus maturation involves multiple steps during virion assembly. One of the key steps is the forming of an immature virion with nucleoid (IVN) by inserting the viral genomic DNA into immature virion (IV). Four viral proteins (A22, I6, A32, and A13) are critical in poxvirus DNA maturation and packaging [5], as the poxvirus remains in the IV state without progressing into the IVN state if any of these four proteins is absent [5–7].
A32, a late protein with ATPase activity that is widely expressed in the Poxviridae [2], shows sequence similarity to the gene I protein of filamentous single-stranded DNA bacteriophages and the IVa2 gene protein of adenovirus [8]. The vaccinia virus A32 protein is indispensable for viral genome packaging, as A32 deficient virus only produces nascent virion that lacks genomic DNA despite normal DNA replication and processing [9]. Furthermore, the A32 protein from orf virus, a member of the Parapoxvirus genus, was also shown to be an ATPase [4]. The A32 proteins are highly conserved among orf virus strains [10, 11] except the heterogeneous C-terminal tail, and this C-terminal tail does not exist in GTPV [12].
A32 is an ATPase which belongs to the FtsK-HerA family, an ASCE (additional strand, conserved E) group of the P-loop NTPase superfamily [13], and has a role in dsDNA packaging. FtsK protein, a member of FtsK-HerA family, forms a hexamer with a central pore that allows dsDNA to pass through, and it has been shown to mediate DNA translocation in gram-negative bacteria [14]. Like other P-loop NTPases, the classic Walker A and Walker B motifs are present in the A32 proteins [8, 15, 16]. In addition, three other A32-specific motifs are also present in the A32 protein [8, 11]. All five of these motifs play various roles in the ATP hydrolysis in orf virus A32 protein. Mutations in those five motifs lead to significant decrease in ATPase activity in orf virus A32 [17].
The enzymatic activity of the A32 protein and its role in virion formation have been reported previously in vaccinia virus and orf virus [9, 17]. In this study, recombinant A32 protein from GTPV was purified and its ATPase activity was examined. The effects of various divalent ions, as well as that of dsDNA, on ATP hydrolysis were investigated. Our results showed that the GTPV A32 protein could bind dsDNA, and its binding affinity was influenced by divalent ions.
Materials and methods
Construction of A32 expression vector
A previously constructed plasmid, pGM-T-A32-Yunlin, was used to amplify the A32 gene for cloning into an expression vector [12]. The full length of the A32 gene was amplified by PCR using two primers (A32L EcoRI primer: AAAGAATTCGATGAATAGGTTTAAG; A32L XhoI primer: AAACTCGAGTTTTTCATACAATG), which contain EcoRI and XhoI restriction sites at their 5′ ends, respectively (Table 1). PCR was performed in 25 μl reaction mixture that contained 2 pmol of each primer, 5 μl of 5× taq master mix (Protech, New Taipei City, Taiwan), and 50 ng of the pGM-T-A32L-Yunlin plasmid. The PCR program had an initial denaturation step of 94 °C for 4 min, followed by 35 cycles of 94 °C for 35 s, 48 °C for 30 s, and 72 °C for 1 min, and ended with a final extension at 72 °C for 7 min. The PCR product was separated by 2 % agarose gel electrophoresis and then purified.
The purified PCR product and the pET-32b vector (Novagen, Billerica, Massachusetts, USA) were digested with EcoRI and XhoI restriction enzymes at 37 °C for 2 h, and then the digested PCR product and the vector were ligated using T4 DNA ligase. E. coli DH5α competent cells (YEASTERN BIOTECH, Taipei, Taiwan) were transformed with the ligation mix. Plasmids containing the correct inserted DNA were identified and confirmed by automated DNA sequencing. The pET-32b vector containing identified full length A32 gene of the GTPV Yunlin strain was then transformed into E. coli BL21 (DE3) for recombinant protein expression.
Protein expression and purification
Protein expression was carried out using the E. coli strain BL21(DE3) (Invitrogen, New York City, USA). Bacterial cells that had been transformed with the desired construct were used to express the recombinant GTPV A32 protein (rA32); this was done by inducing the cells with 0.8 mM isopropyl β-D-1-thiogalactopyranoside (IPTG) at 24 °C for 24 h. Then, the cell pellet was collected by centrifugation and resuspended in binding buffer (50 mM Tris–HCl, 500 mM NaCl, 5 mM imidazole, pH 8.0). Lysozyme and Triton X-100 were then added in the mixture to a final concentration of 1 mg/ml and 1 %, respectively. This was followed by three cycles of freezing and thawing and later sonicated on ice for 6 min. After centrifugation for 30 min at 4 °C, the supernatant, which contained the native form of the soluble recombinant protein, was collected. This supernatant was used to purify recombinant protein by immobilized metal affinity chromatography (IMAC).
In order to carry out protein purification by IMAC, each column, contained 1 mL of Toyopearl AF-Chelate-650 M (Tosoh Biosicence, Tokyo, Japan), was pretreated with 10 mL 0.1 M NiSO4 at 4 °C for 20 min. Supernatant was added to a column after draining out excess NiSO4 and the column was then incubated at 4 °C for 5 h. Endogenous histidine-containing proteins that were non-specifically bound to the matrix were removed by washing the column several times with washing buffer (50 mM Tris–HCl, 500 mM NaCl, 50 mM imidazole, pH 8.0). Finally, the rA32 proteins were eluted using 3 mL of elution buffer (50 mM Tris–HCl, 400 mM NaCl, 200 mM imidazole, pH 8.0).
Next, the purified protein solution was dialyzed against dialysis buffer (20 mM Tris–HCl, 250 mM NaCl, 20 % glycerol, pH 7.3) for 3 h and this procedure was repeated three times to remove residual imidazole. Each protein sample was quantified by the Bradford method (SIGMA, St. Louis, Missouri, USA). In brief, 20 μl protein solution was added into 600 μl Bradford reagent and the OD595 was used to determine protein concentration.
Thioredoxin in pET-32b vector expressed and purified from E. coli BL21 strain was also obtained using the same procedure. This purified thioredoxin was served a control in ATPase assay.
ATPase assay
The ATPase activity of the recombinant proteins was analyzed. The ATPase assay was performed in a 20 μl reaction. Each reaction contained 4 μl of 5× ATPase assay buffer (250 mM HEPES, 250 mM NaCl, pH 7.3), 5 mM of Mg2+, Ca2+ or/and Zn2+, 2 mM of ATP (SIGMA, St. Louis, Missouri, USA), and 1 μg recombinant protein, with or without 100 ng dsDNA. A 123-bp-long dsDNA PCR product from the A32 open reading frame of GTPV Yunlin strain was used as a substrate to study the effect of dsDNA on the ATPase activity of rA32. This dsDNA PCR product was amplified using the GFP and HRMRP primers, which were published in a previous study [12]. The reaction mixtures with or without protein were first preheated to 37 °C and then the reaction was allowed to proceed for 1 h at 37 °C. The reaction was stopped by snap-cooling on ice water and then frozen at −80 °C. The concentration of inorganic phosphate (Pi) was measured by malachite green assay [18]. Briefly, a 20-μl reaction was mixed with 400 μl of malachite dye (mixture of 0.045 % malachite green solution and 4.2 % ammonium molybdate at a ratio of 3:1) and incubated at room temperature (~25 °C) for one minute. 50 μl of 34 % citric acid was later added to stop the reaction. The OD values at 660 nm were measured, and the Pi concentrations were estimated by fitting the OD values to a standard curve using KH2PO4 solution.
Enzyme kinetics of recombinant ATPase
Eight ATPase assay reactions containing different concentrations of substrate (0.5, 1, 2, 3, 4, 6, 8, 10 mM of purified recombinant protein) were carried out to measure the enzyme kinetics of the recombinant A32. Each reaction contained 4 μl of 5× assay buffer, 5 mM of MgCl2, and 1 μg of recombinant A32 protein. All mixtures and ATP solution were preheated to 37 °C for 1 min. The ATP was then added into the preheated reactions and the mixture was incubated at 37 °C for 1 h. The Vmax and Km values for the ATPase activities were calculated by the double reciprocal plot method.
Electrophoresis mobility shift assay (EMSA)
EMSA were performed using 100 ng of dsDNA (123 bp in the A32 gene made by PCR) and 0.5–2 μg protein in 20 μl reaction mixture. Each reaction contains 4 μl of 5× ATPase assay buffer with or without various cofactors (divalent cations or ATP) at 25 °C for 30 min. The reaction product was analyzed by 2 % agarose gel electrophoresis. Different quantities of dsDNA (100, 75, 50, and 25 ng) were used as the standard control group (Fig. 4a). The free DNA bands (not bound by rA32) in each gel were analyzed and quantified by Quantity One software (Bio-Rad, Hercules, California, USA).
Results
rA32 expression and purification
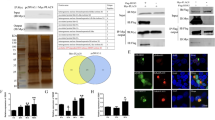
We fused thioredoxin to the N-terminus of the A32 and put his-tags at both the N- and C-terminus of the A32, as illustrated in (Fig. 1a). This recombinant A32 (rA32) was purified by anti-His-tag column. The major protein band had a size of ~45 kDa, which corresponds to the size of thioredoxin at ~17 kDa plus the GTPV A32 protein at ~29 kDa (Fig. 1b). We also confirmed the identity of this 45 kDa protein was the recombinant A32 protein by liquid chromatography–mass spectrometry (data not shown).
a A schematic of the recombinant A32 protein (rA32). The rA32 protein consists of the A32 protein (29 kDa) fused with a thioredoxin molecule (thx, ~17 kDa) at N-terminus. The Hs are histidines that form the hexa-histidine of his-tag. b Purified rA32 protein after dialysis was analyzed by 12 % SDS-PAGE and stained with coomassie blue. A prominent protein band with a size of about 43 kDa is present, which is close to that of the expected rA32. M: protein markers. c The enzyme kinetics of the recombinant A32 protein displayed as a double reciprocal plot. The Vmax and Km are 2.427 × 10−5 mM/sec and 3.59 × 10−3 M, respectively
Effects of divalent cations on the ATPase activity of rA32
A32 proteins from different orf viruses isolated in Taiwan were found to have different ATP hydrolysis activities in the presence of Mg2+ or Ca2+ [17]. These A32 protein sequences from different strains of parapoxvirus are highly conserved except the C-terminal tail, and this C-terminal tail does not exist in the GTPV A32 [12]. We quantified the effect of Mg2+ and Ca2+ on the ATPase activity of the GTPV rA32 by measuring the inorganic phosphate (Pi) releasing rate using the malachite green assay [18]. The ATP hydrolysis activity of the rA32 protein at different ATP concentrations were analyzed and a double reciprocal plot was shown in (Fig. 1c). Our results also showed that the ATP hydrolysis activity of the GPTV A32 was significantly higher in the presence of Mg2+ than Ca2+ by t test analysis (p = 0.035) (Fig. 2).
Effects of metal ions and dsDNA on the ATPase activity of the rA32 protein. Lanes 1–12 indicate the ions and dsDNA composition of each assay condition. The results show that either magnesium ions or calcium ions are essential for enzyme activity; however, a dramatic increase in enzyme activity was found in the presence of magnesium and dsDNA. The error bars are drawn base on the standard deviations. Each value was calibrated by subtracting the negative control (thioredoxin). Double asterisk the value was significantly different between lanes 1 and 2 (p < 0.01). Hash the value was significantly different between lane 1 and 5 (p =0.035). The value was not significantly different between lane 5 and 6
The ATPase activity of A32 influenced by dsDNA
Earlier studies have shown that certain viral and bacterial DNA packaging ATPases can be activated by dsDNA [14, 19–21]. To test whether dsDNA can modulate the ATPase activity of rA32, we used a 123-bps PCR product of the A32 gene as a dsDNA. Interestingly, as shown in Fig. 2, dsDNA augmented this ATPase activity by five-fold in the presence of Mg2+ but not Ca2+ (Fig. 2). As Ca2+ is about 30 % larger than Mg2+, we tested whether Zn2+, another divalent cation with comparable radius to Mg2+, could influence the ATPase activity of rA32. Unexpectedly, we found that Zn2+ completely inhibited ATP hydrolysis, even in the presence of Mg2+ or Ca2+ (Fig. 2). Zn2+ may inhibit ATP hydrolysis by either direct competing with Mg2+/Ca2+ for their binding site(s) on A32 or allosterically inducing a novel conformation that cannot hydrolyze ATP. The possible mechanism of Zn2+ inhibiting ATPase activity of A32 will be discussed later.
The electrophoretic mobility shift assays (EMSAs) was used to analyze binding between rA32 and dsDNA [26]. First, we found that the amount of unbound dsDNA decreased by adding rA32 to the EMSA assay, even in absence of ATP and divalent cations (Fig. 3a). Unbound dsDNA was reduced while the DNA protein complex was retained in the well (Fig. 3a, b). These findings indicated a direct binding between rA32 and dsDNA. To rule out the effect of the thioredoxin that was tagged on the rA32, we performed a control experiment using thioredoxin alone. As expected, thioredoxin failed to bind dsDNA. We further demonstrated that the bound dsDNA can be released by sodium dodecyl sulfate (SDS) (Fig. 3b).Thus, based on these results, we concluded that the dsDNA binds directly onto the GTPV A32 portion of the rA32 (Fig. 3a).
aThe dsDNA binding properties of the rA32 protein. b Effect of detergent on the EMSA [26]. a Lane 1: 2 μg of rA32 was treated with 100 ng dsDNA; the unbound form (f) of the DNA is decreased. Lane 2 is thioredoxin used in binding assay as an internal control. N (lane 3): negative control without protein. b Analysis conditions were the same as (a). In lane 2, SDS was added after treating 2 μg of rA32 with 100 ng dsDNA. F Free DNA unbound, B DNA bounded with protein
Factors influencing the dsDNA binding property of rA32
Previous study has shown that Zn2+ binds directly onto the B204 protein of Sulfolobus-turreted icosahedral virus 2 (STIV2), a member of A32-like ATPase family, and induces a conformational change [20]. To test whether divalent ions or ATP directly interfere the dsDNA binding ability of A32, we performed EMSAs in a variety of conditions. The amount of unbound dsDNA was decreased as the rA32 concentration increased, implying the rA32–dsDNA complex was formed (Fig. 4a, lanes 1–5; and 6–10 were negative control in the absence of protein). We quantified the rA32–dsDNA binding ability under different divalent cations and ATP (Fig. 4). We found that more unbound dsDNA was detected when ATP was included in the EMSA assays, indicating ATP may allosterically induce a conformation that does not favor the binding of dsDNA with A32 (Fig. 4 and group 2 in panel b).
The EMSA results investigating the dsDNA binding properties of rA32 and the effect of various different cofactors. a Lanes 1–5 show the interaction of dsDNA (100 ng) with various amounts of rA32 protein (0–2 μg). Lanes 6–10 act as negative control (in the absence of protein and using dsDNA, 100–0 ng, to allow standard curve construction). No divalent ions were added to the binding reactions. F Free DNA unbound, B DNA bounded with protein. b Effects of divalent ions and ATP on the dsDNA–protein binding as analyzed by EMSA (bands are F forms). c Quantification of panel b results in groups 1–7. The addition of ATP reduced the DNA binding ability of the rA32 protein. A combination of ATP and magnesium ion tremendously decreased DNA–protein binding. d Quantification of panel (b) results of groups 1, 5, 8–10. The results revealed that zinc ions were able to increase DNA–protein binding, but at the same time zinc ions plus ATP decreased binding. Results (c and d) are triple replicates for each experimental condition
rA32, dsDNA, and ATP, in the presence of Mg2+ or Ca2+, exhibited a strong ATPase activity, suggesting that a stable ATP-Mg2+/Ca2+–rA32 complex (an enzyme–substrate complex) with a robust ATP hydrolysis activity was formed. As shown previously, dsDNA potentiated the ATPase activity of ATP-Mg2+-rA32 complex, but not ATP–Ca2+-rA32 complex (Fig. 2), and this result suggested that Mg2+ and Ca2+ may induce two distinct A32 isoforms with different dsDNA binding affinities. In order to explore this phenomenon, dsDNA binding ability of the ATP-Mg2+/Ca2+-rA32 complexes were quantified using EMSA. EMSA reactions containing either Mg2+ or Ca2+ were performed to determine whether the DNA–protein interactions were different when the ATP-Mg2+–rA32 complex and the ATP–Ca2+-A32 complex were compared. We found that Ca2+ slightly lowered dsDNA binding ability in the presence of ATP (Fig. 4) and this effect might be indirectly caused by shielding the negative charges on the dsDNA by Ca2+. Alternatively, Ca2+ did not affect the dsDNA–rA32 interaction and this agrees with the ATPase assay results (Fig. 2). The ATP-Mg2+-rA32 complex, unlike the ATP–Ca2+-rA32 complex, showed a much reduced dsDNA binding ability, suggesting that the interaction between dsDNA and rA32 is affected within this complex by Mg2+ and our previous ATPase assay further supported this hypothesis (Figs. 2, 4).
To confirm the DNA binding ability of the ATP–Ca 2+-rA32 complex and ATP-Mg2+–rA32 complex was not directly influenced by the divalent cations, we carried out EMSAs containing only Mg2+ or Ca2+ and the results showed that the dsDNA binding ability of rA32 was only slightly decreased (Fig. 4 and group 3, 4 in panel b). We speculate that this slight reduction in DNA binding ability by Ca2+or Mg2+ might be caused by shielding the negative charges on the dsDNA and thus interferes with the interaction between dsDNA and rA32. The ATP-Mg2+–rA32 complex, once formed, may participate in dsDNA binding and Mg2+ may affect the stability of this dsDNA–ATP-Mg2+–rA32 complex.
In the presence of zinc ion rA32 can bind dsDNA binding but the ATPase activity is blocked
We found the dsDNA binding ability of rA32 was increased significantly in the presence of Zn2+ (Fig. 4). However, the ATPase activity of rA32 was completely blocked by Zn2+, even in the presence of dsDNA that has been shown to potentiate the Mg2+/Ca2+-dependent ATP hydrolysis ability of the rA32 (Fig. 2). Possible explanations for ATPase activity inhibitory by Zn2+, was either competition for the Mg2+ binding site or the presence of a Zn2+ binding site within rA32, have been proposed earlier in the study. To test via which mechanism the Zn2+ inhibits ATP hydrolysis, we performed EMSA to examine if the interaction between rA32 and dsDNA would be interfered by Zn2+.
The ATP-Mg2+/Ca2+-rA32 complex and Zn2+ were mixed and then EMSAs were performed; reactions containing only Zn2+ and ATP (ATP–rA32(Zn2+)) served as controls (Fig. 4b) groups 8–10. We found both ATP–Ca2+-rA32(Zn2+) (Fig. 4b group 10) and ATP–rA32 (Zn2+) (group 8) exhibited comparable dsDNA binding affinity. This result further supports our previous finding that dsDNA binding affinity is independent of the ATP hydrolyzing ability in the ATP–Ca2+-rA32 complex (Fig. 4). By contrast, the ATP-Mg2+–rA32 (Zn2+) complex exhibited lower dsDNA affinity than that of the ATP–rA32 (Zn2+) complex, indicating the ATP-Mg2+–rA32 complex was still formed in the presence of Zn2+(Fig. 4). Based on these results, we conclude that Zn2+ blocks ATP hydrolysis rather than inhibits ATP binding to the rA32, as Zn2+ inhibited ATPase hydrolysis and dsDNA binding ability of rA32, but it had little effect on Mg2+ binding to the rA32. These findings further suggest that the A32 may contain one or more Zn2+ binding site (s).
Discussion
The poxviral A32 of orf virus has been shown to have ATPase activity, and divalent cations such as Mg2+ and Ca2+ have been shown to modulate the ATPase activity of these proteins [17]. Sequence alignment indicates that the C-terminal tail of A32 proteins of orf viruses are highly variable, but the remainder of the protein is highly conserved [11]. Based on the functional assays, the C-terminal tail of orf virus A32 may play a vital role in regulating ATP hydrolysis.
The GTPV A32 protein consists of 253 amino acids and the C-terminal tail that is commonly found in orf virus A32 is absent [12]. In this study, we found that Mg2+ was much potent than Ca2+ to induce the ATP hydrolysis activity of the rA32 (Fig. 2), but this result contrasts with orf virus A32 proteins which exhibit dramatically different ATP hydrolysis activities with Mg2+ or Ca2+. The heterogeneous C-terminal tail of the poxvirus A32 proteins may play a role in this cation-species dependent ATP hydrolysis.
Poxvirus A32 has been proposed as a dsDNA packaging ATPase, as functional mutations of the A32 may interrupt viral genome encapsidation [9]. However, binding between A32 protein and the viral genomic DNA has yet to be proven. In this study, we have provided evidence showing that the A32 protein directly binds to dsDNA, and dsDNA further potentiates the ATP hydrolysis activity of the A32 protein (Fig. 2). Interestingly, this dsDNA-dependent enhancement of ATPase hydrolysis requires the presence of Mg2+, but not Ca2+ (Fig. 2).
Our data have shown that divalent cations are not required for the dsDNA binding to A32 (Fig. 4). This result further implies that the A32 protein can bind dsDNA in an inactive form. The Gp16 protein of phi29 phage is an ATPase that also mediates genomic DNA packaging and non-hydrolyzable γ-S-ATP further potentiated the DNA binding affinity of the Gp16 protein [22]. By contrast, the dsDNA binding ability of GTPV A32 was suppressed by ATP, and this result implies that ATP may induce conformational change that either directly inhibits the dsDNA binding site(s) or allosterically modulates the dsDNA binding site(s). Furthermore, ATP-Mg2+–A32 complex exhibited a reduced apparent dsDNA binding ability. Because Zn2+ inhibits the ATPase activity, it is unlikely that ATP hydrolysis can generate sufficient energy that facilitates dsDNA sliding through A32 and decreases the apparent dsDNA binding affinity. The active form of the A32 protein has a lower dsDNA binding affinity and is similar with that of the gp17 of T4 phage [21] (Fig. 4).
Zn2+ have been co-crystallized with the STIV2 B204 protein, a member of the A32-like DNA packaging ATPase [20]. The Zn2+ binding sites are located on the inter-subunit surfaces of the B204 hexamer. Binding of Zn2+ leads to conformational changes from one B204 subunit to its neighboring subunits within the hexamer complex [20]. Similarly, our findings suggest that the A32 protein might have one or more Zn2+ binding sites and Zn2+ does not interfere with the enzyme–substrate complex formation; instead, those Zn2+ binding sites may regulate the energy transfer of ATP hydrolysis and subsequently control the dsDNA translocation through the A32 protein.
During the genome packaging, the vaccinia virus genome is condensed from a linear dsDNA (2 nm × 68 mM) to a very condensed nucleoid (200 nm × 0.15 μm) [5]. This nucleoid is wrapped by endoplasmic reticulum (ER) membrane and this membrane-wrapped-nucleoid formed the IVN [23, 24]. However, there are several discrepancies between our results and the existing “DNA packaging role” of A32. Firstly, A32 can directly bind to dsDNA in the absence of other cofactors and the ATPase activity of the enzyme was dramatically enhanced by linear dsDNA (Figs. 2, 3). These results suggest that A32 binds to naked dsDNA rather than condensed nucleoid. Secondly, it is less likely that the oligomeric ring is capable to encapsidate a large-condensed nucleoid; nevertheless, more evidence is required to verify our hypothesis.
Because of the high sequence similarity between A32 proteins and FtsK protein, A32 is classified in the same family with FtsK of FtsK-HerA and ASCE family [13]. The A32 proteins are highly conserved among difference virus species and we speculate that the A32 protein may work similarly as other FtsK-HerA proteins for viral genomic DNA packaging. Earlier study showed that large cytoplasmic DNA crystalloids were observed in rifampin-treated, vaccinia-infected HeLa cells where the vaccinia A32 expression was inhibited [25]. These DNA crystalloids were observed when other DNA packaging proteins were inhibited, which further proved that the A32 protein is involved in viral genomic DNA packaging [5, 9]. We suggest, instead of inserting the condensed nucleoid into the IV of virus, the GPTV A32 translocates a single-uncondensed immature viral genomic DNA from the DNA replicating site to the viral assembly site.
In summary, our results showed that the poxviral A32 can bind directly to the dsDNA, and the ATPase activity of A32 that is crucial for genome packaging is then potentiated upon dsDNA binding. However, as A32 binds single linear dsDNA, rather than condensed nucleoid, A32 may exert its function in the early stage of genome packaging prior to genome condensation. Furthermore, we have also demonstrated that Zn2+ may inhibit viral genome packaging via blocking ATPase hydrolysis and dsDNA translocation, and this phenomenon has not been described previously. Our work indicated that viral A32 protein, via its ATP hydrolysis activity, has a role in dsDNA packaging and translocation.
References
N.J. Maclachlan, E.J. Dubovi, Poxviridae, in Fenner’s veterinary virology, ed. by N.J. Maclachlan, E.J. Dubovi (Elsevier, London, 2011), pp. 151–165
B. Moss, Poxviridae: the viruses and their replication, in Field’s virology, 4th edn., ed. by D.M. Knipe, P.M. Howley (Wolters Kluwer, Philadelphia, 2007), pp. 2905–2930
S. Babuk, T.R. Bowden, G. Parkyn, B. Dalman, D.M. Hoa, N.T. Long, P.P. Vu, do Bieu, X.J. Copps, D.B. Boyle, Yemen and Vietnam capripoxviruses demonstrate a distinct host preference for goats compared with sheep. J. Gen. Virol. 11, 105–114 (2009)
F.Y. Lin, K.W. Chan, H.C. Wang, W.L. Hsu, M.L. Wong, Functional expression of the recombinant ATPase of orf virus. Arch. Virol. 155, 1701–1705 (2010)
R.C. Condit, N. Moussatche, P. Traktman, In a nutshell: structure and assembly of the vaccinia virion. Adv. Virus Res. 66, 31–124 (2006)
J. DeMasi, S. Du, D. Lennon, P. Traktman, Vaccinia virus telomeres: interaction with the viral I1, I6, and K4 proteins. J. Virol. 75, 10090–10105 (2001)
O. Grubisha, P. Traktman, Genetic analysis of the vaccinia virus I6 telomere-binding protein uncovers a key role in genome encapsidation. J. Virol. 77, 10929–10942 (2003)
E.V. Koonin, T.G. Senkevich, V.I. Chernos, Gene A32 product of vaccinia virus may be an ATPase involved in viral DNA packaging as indicated by sequence comparisons with other putative viral ATPase. Virus Genes 7, 89–94 (1993)
M.C. Cassetti, M. Merchlinsky, E.J. Wolffe, A.S. Weisberg, B. Moss, DNA packaging mutant: repression of the vaccinia virus A32 gene results in noninfectious, DNA-deficient, spherical, enveloped particles. J. Virol. 72, 5769–5780 (1998)
R. Yogisharadhya, V. Bhanuprakash, G. Venkatesan, V. Balamurugan, A.B. Pandey, S.B. Shivachandra, Comparative sequence analysis of poxvirus A32 gene encoded ATPase protein and carboxyl terminal heterogeneity of Indian orf viruses. Vet. Microbiol. 156, 72–80 (2012)
K.W. Chan, C.H. Yang, J.W. Lin, H.C. Wang, F.Y. Lin, S.T. Kuo, M.L. Wong, W.L. Hsu, Phylogenetic analysis of parapoxviruses and the C-terminal heterogeneity of viral ATPase proteins. Gene 432, 44–53 (2009)
K.W. Chan, M.L. Lee, W.C. Yang, M.L. Wong, W.L. Hsu, C.F. Ho, Y.C. Hsieh, C.Y. Wang, Differential diagnosis of Goatpox virus in Taiwan by multiplex polymerase chain reaction assay and high-resolution melt analysis. J. Vet. Diagn. Invest. 26, 195–202 (2014)
L.M. Iyer, K.S. Makarova, E.V. Koonin, L. Aravind, Comparative genomics of the FtsK–HerA superfamily of pumping ATPases: implications for the origins of chromosome segregation, cell division and viral capsid packaging. Nucleic Acids Res. 32, 5260–5279 (2004)
T.H. Massey, C.P. Mercogliano, J. Yates, D.J. Sherratt, J. Lowe, Double-stranded DNA translocation: structure and mechanism of hexameric FtsK. Mol. Cell 23, 457–469 (2006)
J. Snider, W.A. Houry, AAA + proteins: diversity in function, similarity in structure. Biochem. Soc. Trans. 36, 72–77 (2008)
D.D. Leipe, Y.I. Wolf, E.V. Koonin, L. Aravind, Classification and evolution of P-loop GTPases and related ATPases. J. Mol. Biol. 317, 41–72 (2002)
F.Y. Lin, K.W. Chan, C.Y. Wang, M.Y. Wong, W.L. Hsu, Purification and functional motifs of the recombinant ATPase of orf virus. Protein Expr. Purif. 79, 210–216 (2011)
P.A. Lanzetta, L.J. Alvarez, P.S. Reinach, O.A. Candia, An improved assay for nanomole amounts of inorganic phosphate. Anal. Biochem. 100, 95–97 (1979)
K.R. Kondabagil, Z. Zhang, V.B. Rao, The DNA translocating ATPase of bacteriophage T4 packaging motor. J. Mol. Biol. 363, 786–799 (2006)
L.J. Happonen, E. Oksanen, L. Liljeroos, A. Goldman, T. Kajander, S.J. Butcher, The structure of the NTPase that powers DNA packaging into Sulfolobus turreted icosahedral virus 2. J. Virol. 87, 8388–8398 (2013)
T.I. Alam, V.B. Rao, The ATPase domain of the large terminase protein, gp17, from bacteriophage T4 binds DNA: implications to the DNA packaging mechanism. J. Mol. Biol. 376, 1272–1281 (2008)
C. Schwartz, H. Fang, L. Huang, P. Guo, Sequential action of ATPase, ATP, ADP, Pi and dsDNA in procapsid-free system to enlighten mechanism in viral dsDNA packaging. Nucleic Acids Res. 40, 2577–2586 (2013)
D. Rodriguez, M. Bárcena, W. Möbius, S. Schleich, M. Esteban, W.J. Geerts, A.J. Koster, G. Griffiths, J.K. Locker, A vaccinia virus lacking A10L: viral core proteins accumulate on structures derived from the endoplasmic reticulum. Cell Microbiol. 8, 427–437 (2006)
G. Griffiths, R. Wepf, T. Wendt, J.K. Locker, M. Cyrklaff, N. Roos, Structure and assembly of intracellular mature vaccinia virus: isolated-particle analysis. J. Virol. 75, 11034–11055 (2001)
P.M. Grimley, E.N. Rosenblum, S.J. Mims, B. Moss, Interruption by rifampin of an early stage in vaccinia virus morphogenesis: accumulation of membranes which are precursors of virus envelopes. J. Virol. 6, 519–533 (1970)
N. Zhao, M. Sun, K. Burns-Huang, X. Jiang, Y. Ling, C. Darby, S. Ehrt, G. Liu, C. Nathan, Identification of Rv3852 as an agrimophol-binding protein in Mycobacterium tuberculosis. PLoS One 10, e0126211 (2015)
Acknowledgments
This study was financially supported by a Grant from the Minister of Science and Technology, Taiwan.
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by Joachim Jakob Bugert.
Rights and permissions
About this article
Cite this article
Lee, ML., Hsu, WL., Wang, CY. et al. Goatpoxvirus ATPase activity is increased by dsDNA and decreased by zinc ion. Virus Genes 52, 625–632 (2016). https://doi.org/10.1007/s11262-016-1349-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-016-1349-3