Abstract
Clustered regularly interspaced short palindromic repeats and associated Cas proteins (CRISPR-Cas) are the only known adaptive immune system in prokaryotes. CRISPR-Cas system provides sequence-specific immunity against invasion by foreign genetic elements. It carries out its functions by incorporating a small part of the invading DNA sequence, termed as spacer into the CRISPR array. Although the CRISPR-Cas systems are mainly responsible for adaptive immune functions, their alternative role in the gene regulation, bacterial pathophysiology, virulence, and evolution has started to unravel. In several species, these systems are revealed to regulate the processes beyond adaptive immunity by employing various components of CRISPR-Cas machinery, independently or in combination. The molecular mechanisms entailing the regulatory processes are not clear in most of the instances. In this review, we have discussed some well-known and some recently established noncanonical functions of CRISPR-Cas system and its fast-extending applications in other biological processes.
Similar content being viewed by others
Introduction
CRISPR-Cas system defends the prokaryotes from invasion by mobile genetic elements (MGEs) including phages, plasmids, and transposons. This system was first observed in the various archaeal genomes in the late 1980s. Its apparent prevalence in the broad range of bacterial and archaeal lineages suggested the common function of CRISPR-Cas systems in prokaryotes (Jansen et al. 2002; Mojica et al. 2000). After initial discovery, owing to the unique sequence and structure of a repeat-spacer array, inference of CRISPR-Cas function in prokaryotes took almost two decades. Detailed sequence analysis and bioinformatics studies revealed that the spacers in the CRISPR array target the phage and plasmid sequences (Bolotin et al. 2005; Pourcel et al. 2005). Thereafter, experimental evidence showed that CRISPR-Cas prevented phage infection in Streptococcus thermophilous as well as averted the invasion by plasmid in Staphylococcus epidermidis, in a sequence-specific manner (Barrangou et al. 2007; Marraffini and Sontheimer 2010).
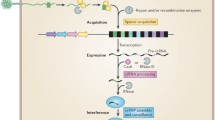
The brief outline of the CRISPR-Cas mechanism of action is shown in the following Fig. 1. The defining feature of the CRISPR-Cas system is the presence of a repeat-spacer array preceded by an AT-rich promoter sequence and adjoining Cas operon lying nearby in the genome (Charpentier et al. 2015). The CRISPR-Cas mediates its function by integrating a small nucleic acid sequence known as a spacer, acquired from invading foreign genetic element into the CRISPR array and thus confers the cell a unique and heritable form of adaptive immunity. This embodies the first adaptation step of the CRISPR-Cas mechanism. This is followed by the expression step, whereby transcription of CRISPR loci takes place which generates pre-crRNA (pre-CRISPR RNA). Pre-crRNA is recognized by ribonuclease and cleaved into smaller repeat-spacer units to generate crRNAs. This crRNA in combination with Cas proteins forms monomeric/multimeric complexes, which are part of the interference complex. In the last interference step, crRNA hybridizes to the complementary target sequence from the invading genetic element and thus recruits the interference complex onto the target sequence leading to its degradation (Charpentier et al. 2015).
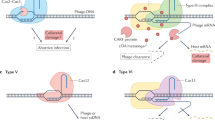
CRISPR-Cas system mechanism of action (the pictorial presentation over here depicts the mechanism of action of type I-E CRISPR-Cas system characterized from E. coli). CRISPR-Cas system protects the organism from incoming phage or plasmid infection by incorporating a small stretch of DNA into the CRISPR array with the help of Cas1 and Cas2 protein dimmers. This process is known as adaptation or acquisition. Cas1 and Cas2 are the universal protein as are found in most CRISPR types. During the next infection from the same phage or plasmid, the CRISPR array is transcribed to produce the pre-crRNA in the expression step, which is then cleaved by Cas6 protein to produce a repeat-spacer unit called crRNA. The palindromic repeat sequences lead to the formation of hairpin-loop structure which in association with spacer sequence functions as a guide for the Cas proteins. After recognizing and binding the protospacer sequence, it loads the cascade complex. In the final interference step, Cas3 protein, a part of cascade complex, comprising the nuclease activity cleaves the non-target strands in 3’ to 5’ direction leaving 200–300 bp nick. The degradation of the nicked DNA is further completed by cascade-independent activity of ssDNA nuclease activity of Cas3
Over the past 15 years, impeccable progress has been achieved in understanding the molecular function of the CRISPR-Cas system. Such developments helped in unraveling the alternative functions of CRISPR-Cas system. The alternate functions of CRISPR-Cas systems are described in detail in various published articles (Newsome et al. 2021; Ratner et al. 2015; Sampson and Weiss 2013; Sampson and Weiss 2014; Westra et al. 2014). CRISPR-Cas systems are incredibly versatile owing to the unique spacers of diverse origin, diversity in adaptation and effector complex, and a large repertoire of cas family genes (Makarova et al. 2002). The diversity in CRISPR-Cas function is mainly attributed to the versatility found in cas gene encoded diverse proteins with different enzymatic domains (Makarova et al. 2011). CRISPR-Cas is a multifaceted system and targets not only dsDNA (double-stranded DNA) but ssDNA (single-stranded DNA) and ssRNA (single-stranded RNA) as well. CRISPR type with RNA-targeting potential has been instrumental in the realization that the CRISPR-Cas system can play a role in endogenous gene regulation, likewise the RNAi system in eukaryotes (Westra et al. 2014). In addition, the observation that CRISPR-Cas can occasionally incorporate spacer from the chromosomal region has also given rise to the perspective that CRISPR-Cas systems might have gene regulatory function (Stern et al. 2010).
Pathogenicity and virulence
Almost three years later after the initial discovery of CRISPR-Cas adaptive immune function, in Pseudomonas aeruginosa, it was observed that type I-F CRISPR system was involved in the regulation of biofilm-forming capacity and swarming motility of this pathogen (Palmer and Whiteley 2011; Zegans et al. 2009). The first well-documented, direct evidence of regulation of pathogenicity and virulence was discovered in Francisella novicida. It has developed a mechanism to subvert the host immune response by modulating the gene expression with the help of CRISPR-Cas system. Once inside the macrophage, Francisella novicida enters the phagosomes containing abundant receptors for the recognition of pathogen (Jones et al. 2012). Toll-like receptor 2 (TLR2) is one such receptor that recognizes bacterial lipoprotein (BLP) and further elicits the immune response of the host (Aliprantis et al. 1999; Brightbill et al. 1999). Francisella novicida, instead, employs Cas9, tracer RNA (trans-activating), and scaRNA (small CRISPR-associated RNA) to lower the expression levels of BLP protein on the outer envelope. This results in the reduced TLR2 activation and milder immune response, thus allowing the organism to thrive in the host (Jones et al. 2012). Neisseria meningitis in the human lung epithelial model was also shown to utilize Cas9 for the infection of the host. Cas9 was shown to be essential for attachment to the cell wall, invasion, and replication inside the host (Heidrich et al. 2019; Sampson et al. 2013). Likewise in Streptococcus pyogenes, Cas9 was observed to impact the epithelial cell binding and growth in human blood cells. In addition, it was observed that it attenuated the virulence of the pathogen in murine necrotizing skin infection model. In addition, Cas9 was also shown to help Campylobacter jejuni to attach and invade the host in the colorectal epithelial cell model (Louwen et al. 2013). Transcriptome analysis of the wild-type and cas9 mutant strains of Campylobacter jejuni further revealed that Cas9 was involved in the upregulation of virulence factors. It was shown experimentally that the cas9 mutant strain was unable to form biofilm. Further, it lacked the ability of intracellular invasion and adhesion, survival inside the host and motility, suggesting the wide-ranging impact of cas9 on the survival and pathogenicity of the organism (Shabbir et al. 2018). The cas9 gene was found to affect the expression level of virulence determinants in several other microorganisms as well, suggesting its involvement in alternate functions other than the adaptive immunity (Gao et al. 2019).
In Legionella pneumophila, type II-B CRISPR-Cas expression was observed to be induced during intracellular growth in macrophages of amoebas, including Acanthamoeba castellanii and Hartmannella vermiformis. The cas2 gene was found to be important for the growth of organism inside the macrophages. It was reported that the Cas2 protein from this organism comprised of DNase as well as RNase activity. Further, Gunderson and colleagues argued that, as of now, it is not clear that whether the RNase or DNase or both catalytic activities are required for infection; however, the RNase activity appeared to be the dominant in their study. It was established that the nuclease activity was essential for the initiation of infection in the host. It was also established experimentally that the introduction of cas2 in Legionella pneumophila strain lacking CRISPR-Cas led to on an average 60-fold more infectivity (Gunderson and Cianciotto 2013). In Acinetobacter baumannii, harboring type II system, the CRISPR-Cas was shown to be associated with high biofilm-producing ability. These strains in addition to containing genes exclusive for biofilm formation also lacked plasmids, which suggested that the CRISPR-Cas has dual role of adaptive immune function as well as a modulator of group behavior (Mangas et al. 2019). Moreover, Cas3 protein from the type I system, in addition to type II system, was found to modulate the pathogenicity and virulence of several microorganisms. For instance, in Salmonella enteric, cas3 deletion was shown to decrease the biofilm-forming capacity (Cui et al. 2020). In addition, loss of cas3 also affected the intracellular invasiveness inside the host. Transcriptome analysis revealed that cas3 deletion affected quorum-sensing genes, type three secretion systems (T3SS), salmonella pathogenicity island-1 (SPI-1), and genes related to flagella formation (Cui et al. 2020). Cas3 was also reported to regulate the biofilm formation in addition to fluoride resistance in Streptococcus mutans (Tang et al. 2019). In contrast to other studies, cas3 deletion in Porphyromonas gingivalis increased the virulence inside the THP-1 cells. These results also concurred with the earlier studies on Galleria mellonella infection model, whereby infection with cas3 deletion mutant led to high mortality compared to when infected with wild-type strain (Solbiati et al. 2020). The table (Table 1) given below includes some of the other examples of noncanonical functions performed by CRISPR-Cas system in the prokaryotes.
Bacterial physiology
In Myxococcus xanthus, three cas genes including cas8, cas7, and cas5 known as devT, devR, and devS, respectively, were found to be essential for sporulation and fruiting body development (Thony-Meyer and Kaiser 1993). Further, it was observed that devT mutation affected the aggregation, sporulation, and chemotaxis. It was found that the levels of devT affected the transcript levels of the fruA gene, responsible for fruiting body formation (Boysen et al. 2002). It was suggested that the FruA protein has an affinity for the regulatory site in the cas locus, which induces the devT expression and subsequently leads to the positive feedback loop. The role of the CRISPR array in the regulation of fruiting body formation is not clear, though two self-targeting spacers were observed in the CRISPR array in Myxococcus xanthus. Among the two, one was targeting prophage gene encoding integrase, and the other was targeting the cas gene. From the observation, it was inferred that the both cas gene and CRISPR array might be functioning together to alter the physiology of the organism (Viswanathan et al. 2007). In Salmonella typhi, deletion of cas genes belonging to type I-E system hampered the expression levels of outer membrane proteins such as OmpC and OmpF to various extents. It was found that the Cas proteins were regulating the upstream ompR gene (a porin regulator), henceforth altering the profile of outer membrane porin molecules. Microarray studies by the same group revealed that the CRISPR-Cas system is capable of regulating several other Omp molecules in Salmonella typhi. The modulation in the expression level of outer membrane porins is reported to affect several biological processes including response to oxidative stress, bile salt resistance, osmotic balance, and chemotaxis as well virulence (Medina-Aparicio et al. 2021). In addition, in Pseudomonas aeruginosa, a spacer with a partial match to the DMS3 prophage gene altered the pathophysiology of the organism (Cady and O'Toole 2011). In Aggregatibacter actinomycetemcomitans, the spacer was found to target metabolic glycogen phosphorylase enzyme encoding gene (glgp). The exact outcome of this self-targeting is not known as yet; however, looking at the nature of the targeted gene, it seems that it might be affecting bacterial metabolism (Jorth and Whiteley 2012). In Listeria monocytogenes, the CRISPR locus (also termed as rliB locus) comprises four spacer-long CRISPR array without encompassing any associated cas genes (Mandin et al. 2007; Sesto et al. 2014). The spacers in the CRISPR array have been found to be significantly matching with multiple targets including two-component system, a transcriptional regulator, and iron transport (feoAB) gene. Though this CRISPR-Cas system lacks cas genes, it is still capable of utilizing Cas proteins produced exogenously and hamper the plasmid uptake. Nevertheless, it was revealed that the rliB locus is also capable of repressing the expression of gene by hybridizing with the transcript (Mandin et al. 2007).
Response to stress
In many bacterial species, it has been observed that in response to envelop stress, the expression of the CRISPR-Cas system is induced. This phenomenon was first observed in Escherichia coli, where the overexpression of membrane targeting protein activated the expression levels of the CRISPR-Cas system (Perez-Rodriguez et al. 2011). It was also observed that the levels of reporter green fluorescent protein (GFP), fused to the excretory molecule (ssTorA), were reduced in cells lacking chaperone DnaK. Deletion of type I-E cas operon, as well as the BaeSR system, restored the expression of ssTor-GFP. BaeSR, a two-component system, is well-known to upregulate the cas genes expression in response to membrane stress (Baranova and Nikaido 2002; MacRitchie et al. 2008). It was suggested that in E. coli, the combined effect of induced Cas levels and occurrence of ssTorA-targeting spacer (partially matching) in CRISPR array led to reduction in the levels of membrane protein, thereby impacting the transport channel across the cell (Perez-Rodriguez et al. 2011). In addition, in Streptococcus thermophilus and Archaean Sulfolobus islandicus, the CRISPR-Cas system was found to be induced in the aftermath of phage induced envelope stress (Quax et al. 2013; Young et al. 2012). In Myxococcus xanthus, dev operon corresponding to type I-B CRISPR cassette is also found to be activated during stress (Viswanathan et al. 2007). In Streptococcus mutans, it was observed that type II-A and type I-C CRISPR systems are involved in temperature stress tolerance. At higher temperature, the double mutants lacking both CRISPR types showed reduced capabilities to survive in comparison to single CRISPR mutant. Moreover, type II-A CRISPR deletion mutant showed lower growth rate during membrane stress and oxidative stress as compared to type I-C mutants. Intriguingly, it was observed that two-component stress response regulator VicK/R regulated these two CRISPR systems differently. This suggested that various stress factors modulate the expression levels of CRISPR systems in the organism through the channel of signaling molecules with one such being the VicK/R tw-component system. VicK/R was observed to negatively regulate the type II-A system and positively regulate the type I-C system (Serbanescu et al. 2015). In another study, in Streptococcus mutans, deletion of cdaA gene, coding for diadenylate cyclase required for the synthesis of c-di-AMP molecule, led to upregulation of CRISPR 1 locus and downregulation of CRISPR 2 locus. Moreover, deletion of cdaA affected the sensitivity to hydrogen peroxide and production of extracellular polymeric substances. The authors suggested that the CRISPR-Cas systems are indirectly linked to the stress response mechanisms in this organism (Cheng et al. 2016). In Bacillus cereus, the introduction of the CRISPR-Cas in CRISPR-Cas negative strains resulted in decreased tolerance to various stresses as well as decreased pathogenicity (Zheng et al. 2020). In conclusion, it can be remarked that indeed these CRISPR-Cas systems are involved in functions other than the adaptive immunity, though most of the molecular mechanisms pertaining to these processes are still obscure and under investigation. The graphic description of the various environmental stress factors that activate the CRISPR-Cas system is depicted in the following Fig. 2.
CRISPR-Cas in response to stress; in response to different environmental stress, the CRISPR-Cas system is activated by different mechanisms. These stress factors include cell membrane damage by phage infection; envelop stress, heat shock/high temperatures, oxidative stress, and other cellular signaling molecules or DNA damaging factors. The exact downstream mechanism after the activation of the CRISPR-Cas system by which the cell tackles the environmental stress is not identified yet
Endogenous gene regulation
Various researchers speculated that spacers targeting the chromosome, which are being tolerated in the CRISPR array by the organism, might be involved in the gene regulation process (Sorek et al. 2008). Computational analysis of Escherichia coli type I-E CRISPR-Cas system, to investigate the spacer targets, revealed that CRISPR spacers have a high propensity to target chromosomes as compared to phage genomes. In addition, it was revealed that spacers targeted transcriptionally active regions at a higher rate. The study suggested that the type I-E CRISPR-Cas system in Escherichia coli plays a role in endogenous gene regulation (Bozic et al. 2019). In addition to computational analysis, several experimental evidences showed that the CRISPR arrays are involved in gene regulation. In Francisella novicida, the type II CRISPR-Cas system was shown to modulate the expression of a protein-coding gene associated with bacterial virulence. The tracrRNA along with Cas9 and small CRISPR-associated RNA repressed the BLP expression inside the host. It was demonstrated that the bacterium avoided cell death by targeting the BLP mRNA instead of encoding gene (Jones et al. 2012; Sampson et al. 2013). The BLP protein triggers the host T cell-mediated immune response; thus lowering the expression levels of this protein. It enables the bacterium to escape the immune system and thrive inside the host (Sampson et al. 2013). Similarly, in Pelobacter carbinolicus, a self-targeting spacer was identified to target the histidyl-tRNA synthetase (hisS) and lower its expression. The crRNA having the spacer identical to hisS gene was found to compete with the RNA polymerase, thus hindering the transcription of hisS gene. The hisS gene unlike most of the self-targeting spacers was found at the trailer side of the 111 spacer-long CRISPR loci, indicating that the spacer is being tolerated for ages by the bacterium. This self-targeting of hisS gene also resulted in lower frequency of histidine amino acid in the protein molecules of Pelobacter carbinolicus in comparison to the other closely related strains (Aklujkar and Lovley 2010). The type III system which targets mRNA instead of DNA is also observed to be involved in gene regulatory functions. For instance, in Pyrococcus furiosus, Cmr (Cas module RAMP termed as RNA associated mysterious proteins) complex was revealed to recognize and cleave the endogenous complementary RNAs, providing direct evidence that CRISPR-Cas can regulate the transcription and gene expression (Hale et al. 2012). Likewise, Porphyromonas gingivalis which contains the type III system also targets the mRNA and might lead to gene regulation (Endo et al. 2015). Nonetheless, Cas2 protein, part of the adaptation machinery in most CRISPR-Cas systems, can cleave the ssRNA as well as dsDNA, suggesting the potential role of most CRISPR types in gene regulation (Makarova et al. 2015). The various alternative functions regulated by CRISPR-Cas system have been depicted in the following Fig. 3.
CRISPR-Cas mediated physiological processes; CRISPR-Cas is activated due to extracellular signals such as phage infection, DNA invasion, and environmental stress. Upon activation, the CRISPR array and Cas proteins either function in combination or alone to carry out the various biological processes. CRISPR array usually mediates the pathophysiological changes or gene regulation processes by directly targeting the DNA or mRNA by a self-targeting spacer. Cas proteins modulate the cell physiology, virulence, and bacterial behavior either by activating the downstream pathways or by direct interaction with the other proteins or molecules. These processes are multifaceted, and the mechanism entailing various pathways awaits experimental elucidation
Bacterial genome remodeling
In order to avoid cell death due to self-targeting spacers, organisms are shown to remodel the targeted region either by removing the protospacer or deleting the entire targeted region. Several studies have observed that self-targeting forced the organism to evolve. In most of the circumstances, self-targeting of gene located in pathogenicity island (PI) resulted in deletion of entire PI regions. In Pectobacterium atrosepticum, spacer targeting of horizontally acquired island 2 (HAI2) led to deletion of the entire 100 kb HAI2 region to avoid the self-targeting of the genome (Vercoe et al. 2013). Spacer targeting erythromycin (ermB) gene, present in the plasmid and integrated between the flanking IS elements on both sides, also resulted in generation of deletion mutants either lacking the ermB gene or 75% of 66 kb plasmid (Hullahalli et al. 2018). Targeting of lacZ region both in gram-negative Escherichia coli carrying type I-E system and gram-positive Streptococcus thermophilus harboring type II-A systems also led to large genome deletions of up to 35 kb and 37.4 kb size, respectively (Canez et al. 2019; Cui and Bikard 2016). In line with these studies, in other gram positives including Staphylococcus aureus and Streptococcus thermophilus, self-targeting led to loss of large fragments of targeted regions (Guan et al. 2017; Selle et al. 2015). Thus, it can be concluded that the consequence of self-targeting event, as suggested by various studies in diverse range of organisms comprising different CRISPR-Cas types, is the remodeling of the bacterial genome.
Association with DNA repair system
There exists a close association between the CRISPR-Cas systems and DNA repair machinery at different levels, and this has a diverse impact on the microorganisms. These two separate systems usually coincide directly at the adaptation step. In general, during the incorporation process, a nick generated in the first direct repeat by nucleases in order to incorporate a new spacer is again sealed by a DNA repair system (Killelea and Bolt 2017). It has also been shown in the type I-E system in Escherichia coli which utilizes the DNA polymerase from the DNA repair pathway during the spacer acquisition process (Ivancic-Bace et al. 2015). Besides, the common transcription regulator, Cas3a was seen to activate the process of CRISPR-spacer incorporation as well as DNA repair process, suggesting co-operation between the two at several stages (Liu et al. 2015). In addition, it has been shown that proteins involved in the DNA repair pathway and many of the Cas proteins share homologous regions such as helicase domain from Cas3 protein (type I system), nuclease from Cas4 protein (type I and type II), and RuvC like nuclease (type II and type V) (Hudaiberdiev et al. 2017; Shmakov et al. 2017; Sinkunas et al. 2013; Westra et al. 2012; Xiao et al. 2018; Zhang et al. 2012). Furthermore, experimental evidence in Escherichia coli revealed that cas1 deletion resulted in mutants defective in DNA repair pathway (Smith 2012).
Cas1 protein was also found to interact with several cellular repair proteins like RecB, RecC, and RuvB (Babu et al. 2011). The RecBCD system recognizes breaks in dsDNA during the replication process. This repair machinery moves along the replication fork and degrades damaged DNA till it detects the Chi site. An earlier study speculated that degraded DNA products of repair machinery are incorporated as a template for new spacer adaptation processes. Thus, RecBCD can be regarded as part of the anti-phage defense system as it degrades the linear phage DNA and facilitates the incorporation of the spacer. This system avoids the incorporation of the host genome into the CRISPR array by recognizing the Chi site which is present in the chromosomal DNA frequently at a very high rate (Levy et al. 2015). It was observed that RecB deletion hampered naïve spacer adaptation by type I-E systems in E. coli (Ivancic-Bace et al. 2015). According to the hypothesis, in the naïve acquisition process, RecBCD is recruited to repair the DNA damage induced by Cas1 during spacer incorporation (Ivancic-Bace et al. 2015; Levy et al. 2015). Recently, in a study, it was observed that RecG primes the adaptation of spacer from the MGEs by using the pre-existing spacer in type I systems (Heussler et al. 2016; Ivancic-Bace et al. 2015).
Despite sharing similarities at many levels and enabling each other’s successful function, there exists some antagonizing relationship between the two. The CRISPR-Cas adaptive immune function relies on the inability of the DNA repair pathway to reverse the DNA damage induced by Cas nucleases. In a comparative study, CRISPR-Cas machinery and DNA repair systems were analyzed in more than 5000 sequenced bacterial genomes. The study revealed that the type I CRISPR-Cas system has an affinity towards the RecBCD DNA repair system (Levy et al. 2015). This was found to be in concurrence with the observations made in E. coli, where both the systems were having synergistic interaction (Radovcic et al. 2018). However, it was observed that the type II system negatively correlated with the non-homologous end joining (NHEJ) recombination pathway (Faure et al. 2019). To explain the complex relationship between the two, Bernheim and colleagues proposed a hypothesis. Accordingly, CRISPR-Cas systems are frequently subjected to transfer between the species by horizontal gene transfer (HGT). Upon introduction into some new species, CRISPR-Cas continuation in the organism is influenced by several factors, including their production cost, benefit incurred to the host, prevalent phage predation menaces in the population, and inhabitance of other defense systems. In addition, genetic components especially the DNA repair system was also proposed to impact the sustenance of the CRISPR-Cas system in the organism. Proteins involved in the DNA repair system are reported to be under intense selection pressure, and Bernheim et al. proposed that in case of incompatibility, CRISPR-Cas system will be be weeded out from the genome in order to maintain the repair systems. Owing to the strong selection process, organisms will wind up with different combinations of DNA repair systems and CRISPR-Cas systems. The results from this study might help to explain the checkered distribution and diversity of the CRISPR-Cas system in bacteria (Bernheim et al. 2019).
CRISPR-Cas applications
Inhibition of horizontal gene transfer
The rise in multidrug resistance in bacteria has forced the researcher to look for newer tools to tackle the problem of antibiotic resistance. Recently, CRISPR-Cas has also been used as a tool to inhibit the transfer of antibiotic resistance genes from one organism to another. The presence of the CRISPR-Cas system in a genome negatively impacts the concentration of antibiotic resistance determinants in multidrug-resistant (MDR) strains. In one of the study, it has been reported that the CRISPR system is certainly responsible for mitigating the dissemination of mobile genetic elements (MGEs) including prophages and plasmids in Enterococcus faecalis (Palmer and Gilmore 2010). Similarly, a higher number of spacers in the CRISPR array in Streptococcus pyogenes resulted in lower frequency of integrated prophages in the genome (Nozawa et al. 2011). Thus, there is a potential to harness CRISPR systems to curb the dissemination of resistant determinants and virulence in MDR/extreme drug-resistant (XDR) bacteria (Marraffini and Sontheimer 2008). Progress and development in genetic engineering tools made it feasible to introduce the resistance gene targeting spacer into any CRISPR array in a wide range of hosts (Sapranauskas et al. 2011). CRISPR arrays are being designed to target the pathogenicity islands containing antibiotic or virulence genes. This technique allows targeting of only those strains which carry such pathogenicity islands and spares the non-pathogenic populations (Bikard et al. 2014; Citorik et al. 2014; Gomaa et al. 2014). CRISPR-Cas as an antimicrobial resistance tool has been explored only recently. In a study, antibiotic resistance genes were targeted by incorporating the CRISPR-Cas system in lysogenic phage in Escherichia coli. In addition, a spacer targeting the lytic phage was also incorporated in the CRISPR array, and it was postulated that it will further protect the antibiotic-sensitized cells from lytic infection by phage (Yosef et al. 2015).
Typing tool
Way before, their role could be explored in bacterial physiology; CRISPR-Cas systems were utilized for typing the diversity in bacterial species (Grissa et al. 2008). For instance, in Mycobacterium tuberculosis, comprising type III CRISPR system, the variability in the spacer content had routinely been used for typing purposes and study Mycobacterium tuberculosis epidemiology (Abadia et al. 2010; Comas et al. 2009; Gomgnimbou et al. 2012; Kamerbeek et al. 1997). CRISPR typing was also explored in Campylobacter jejuni to reveal the phylogenetic relationship between strains (Schouls et al. 2003). However, unlike Mycobacterium tuberculosis, in Campylobacter jejuni (type II CRISPR system), CRISPR typing was proved to be insufficient to accomplish the understanding of evolutionary relationship. Therefore, CRISPR typing was used in combination with amplified fragment length polymorphism (AFLP) and multilocus sequence typing tool (MLST) (Schouls et al. 2003). A study revealed that polymorphism in the cas gene was linked to a gene found in Campylobacter jejuni strains isolated from patients with Guillain–Barre syndrome (Louwen et al. 2013; vanBelkum et al. 2001). Later on, it was established that single-nucleotide polymorphism along with CRISPR array variations is useful for typing process in Campylobacter jejuni (Louwen et al. 2013). This process of using sequence variation among the spacers in the CRISPR array for studying the diversity is known as spoligotyping. This technique was also used for typing in Corynebacterium diphtheriae and found to be more useful than random amplification of polymorphic DNA (RAPD) and AFLP (Mokrousov et al. 2009). In Legionella pneumophila as well, this technique enabled typing among the environmental strains (Ginevra et al. 2012). Likewise the CRISPR type III and type II, type I CRISPR system was used for typing of Yersinia pestis and Salmonella enterica (Fabre et al. 2012; Vergnaud et al. 2007). CRISPR-Cas-based typing in Yersinia pestis, helped to identify the origin of the ancestral strain which caused the black plague (Vergnaud et al. 2007). In Salmonella, as well, CRISPR typing in combination with other techniques has significantly enhanced the means to distinguish strains causing outbreaks (Cao et al. 2013; Liu et al. 2011). Furthermore, in several other organisms, CRISPR typing has facilitated the delineation of subgroups. For instance, in Erwinia amylovora (type I-E), CRISPR typing helped to distinguish three major groups based on geographic origin (Rezzonico et al. 2011). CRISPR typing in Propionibacterium acnes confirmed three lineages of this organism that have also been identified using other typing techniques (Bruggemann et al. 2012). In Porphyromonas gingivalis, typing based on the CRISPR system, it was observed that with this approach it was possible to regroup the strains beyond the fimbrial gene clusters (Watanabe et al. 2013).
Genome editing
Although all the CRISPR systems have the common feature of recognizing and targeting DNA, most of them are not suited for genetic engineering technologies. For instance, the DNA degradation process of the Ist and IIIrd type CRISPR is not well characterized, and crRNA generation also requires additional Cas6 protein. In contrast, type II CRISPR systems require only a few CRISPR-Cas components for carrying out all the functions. Various studies concluded that the CRISPR-Cas9 complex is ideal for genome editing purposes. Type II targeting of phage and plasmid genetic elements was observed to introduce double-stranded breaks. It was considered to be an important feature from the genome editing point of view in mammalian cells (Garneau et al. 2010). Experimental evidence also revealed that the three elements of type II CRISPR-Cas machinery are more than sufficient to induce double-stranded breaks in mammalian DNA and facilitate the synthesis of single-guide crRNA (sgRNA). Cas9-mediated type II CRISPR systems have been used to target several DNA sequences using small targeting RNA. Small targeting RNA is also known as single-guide RNA (sgRNA) and is synthesized by fusing naturally existing tracrRNA and crRNA having sequence complementarity to the target sequence (Jiang et al. 2015). This genome editing tool has outshined all the other popular tools including transcription activator-like effector nucleases (TALENs) and zinc finger nuclease (ZFNs) (Wang et al. 2013). Genetic engineering involving CRISPR-Cas9 technology involves introduction of the Cas9-sgRNA construct into the cell using transformation. The co-transformation of synthesized recombination DNA template along with the construct increases the chances of incorporating mutations into the target site (Cong et al. 2013). However, to circumvent continual fragmentation of mended homologous recombinant, the recombination DNA template must have mutations to hamper the Cas9 nuclease activity (Wang et al. 2013). Nevertheless, genome editing using Cas9 is a simple and efficient method for editing the genome of several eukaryotic as well as prokaryotic model organisms (Sander and Joung 2014).
Impeccable finding that CRISPR-Cas system can be used for genome editing technologies has opened up the prospects for the use of this system for therapeutic purposes especially for the treatment of cardiovascular diseases, genetic disorders, autoimmune, inflammatory, and immunological disorders, bacterial and viral diseases, metabolic disorders, neurodegenerative diseases, oncological disorders, and others ( Bao et al. 2015; Ebina et al. 2013; Hai et al. 2014; Hu et al. 2014; Wu et al. 2016; Yan et al. 2018; Zhan et al. 2018). Furthermore, recently CRISPR-Cas technology has been used to develop SARS-CoV-2 vaccine candidates (Zhu et al. 2021). Post 2014, tremendous amount of interest had arisen among the scientist and pharmaceutical companies alike to explore and utilize this technology for gene therapy. Over the period, considerable volume of evidence has been gathered in the subject matter, substantiating the therapeutic applications, thus leading to the establishment of numerous strategic partnerships among leading pharmaceutical companies, centered around therapy development and clinical trials. Consequently, several CRISPR-Cas candidates designed for treatment of neurodegenerative and infectious diseases are currently under investigation. Numerous other clinical trials focusing oncological and hematological trials are also underway. Since 2015, close to 2000 patent applications concerning CRISPR-Cas technology has been filed or granted. Currently, based on the recent trends, the CRISPR-Cas abilities appear to be limitless, and the CRISPR-Cas revolution does not seem to end in the near future. However, despite the extraordinary potential that the CRISPR-Cas technology holds, further investigation, probing its safety and therapeutic efficacy in large diverse populations, is required (https://www.researchandmarkets.com/r/o5e94f/ 09/17/2021).
Conclusion
The multiple studies discussed above reasonably support the perspective that CRISPR-Cas systems have a role beyond immunity. Furthermore, it is believed that CRISPR-Cas system, though initially evolved to perform the adaptive immune function in microorganisms, picked up the noncanonical functions later as an adaptation mechanism to various cues from surrounding. However, whether the system evolved as whole, i.e., CRISPR and Cas evolved simultaneously or underwent the adaptation process separately without the influence of the other component, is still uncertain. Owing to the limited amount of information, that too from select few species, it will be too early to reach at any conclusion. It can be said that CRISPR-Cas system is still evolving and its components are picking up noncanonical functions while adjusting in the newer species under novel circumstances. The flexibility and uniqueness of this system to repurpose its function as adaptive defense system of prokaryotes to the modulator of bacterial pathophysiology and several other functions suggest that these systems can be exploited to develop powerful diagnostic, therapeutic, and experimental tools.
Availability of data and material
As no datasets and material were generated in this study, therefore, this section is not applicable.
Code availability
Not applicable.
References
Abadia E, Zhang J, dos Vultos T et al (2010) Resolving lineage assignation on Mycobacterium tuberculosis clinical isolates classified by spoligotyping with a new high-throughput 3R SNPs based method. Infect Genet Evol 10:1066–1074
Aklujkar M, Lovley DR (2010) Interference with histidyl-tRNA synthetase by a CRISPR spacer sequence as a factor in the evolution of Pelobacter carbinolicus. BMC Evol Biol 10:230
Aliprantis AO, Yang RB, Mark MR et al (1999) Cell activation and apoptosis by bacterial lipoproteins through toll-like receptor-2. Science 285:736–739
Babu M, Beloglazova N, Flick R et al (2011) A dual function of the CRISPR-Cas system in bacterial antivirus immunity and DNA repair. Mol Microbiol 79:484–502
Bao D, Ma Y, Zhang X et al (2015) Preliminary characterization of a leptin receptor knockout rat created by CRISPR/Cas9 system. Sci Rep 5:15942
Baranova N, Nikaido H (2002) The baeSR two-component regulatory system activates transcription of the yegMNOB (mdtABCD) transporter gene cluster in Escherichia coli and increases its resistance to novobiocin and deoxycholate. J Bacteriol 184:4168–4176
Barrangou R, Fremaux C, Deveau H et al (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712
Bernheim A, Bikard D, Touchon M et al (2019) A matter of background: DNA repair pathways as a possible cause for the sparse distribution of CRISPR-Cas systems in bacteria. Philos Trans R Soc Lond B Biol Sci 374:20180088
Bikard D, Euler CW, Jiang W et al (2014) Exploiting CRISPR-Cas nucleases to produce sequence-specific antimicrobials. Nat Biotechnol 32:1146–1150
Bolotin A, Quinquis B, Sorokin A et al (2005) Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology (reading) 151:2551–2561
Boysen A, Ellehauge E, Julien B et al (2002) The DevT protein stimulates synthesis of FruA, a signal transduction protein required for fruiting body morphogenesis in Myxococcus xanthus. J Bacteriol 184:1540–1546
Bozic B, Repac J, Djordjevic M et al (2019) Endogenous gene regulation as a predicted main function of type I-E CRISPR/Cas system in E. coli. Molecules 24(4):784
Brightbill HD, Libraty DH, Krutzik SR et al (1999) Host defense mechanisms triggered by microbial lipoproteins through toll-like receptors. Science 285:732–736
Bruggemann H, Lomholt HB, Tettelin H et al (2012) CRISPR/cas loci of type II Propionibacterium acnes confer immunity against acquisition of mobile elements present in type I P. acnes. PLoS One 7:e34171
Cady KC, O’Toole GA (2011) Non-identity-mediated CRISPR-bacteriophage interaction mediated via the Csy and Cas3 proteins. J Bacteriol 193:3433–3445
Canez C, Selle K, Goh YJ et al (2019) Outcomes and characterization of chromosomal self-targeting by native CRISPR-Cas systems in Streptococcus thermophilus. FEMS Microbiol Lett 366(9):fnz105
Cao G, Meng J, Strain E et al (2013) Phylogenetics and differentiation of Salmonella Newport lineages by whole genome sequencing. PLoS ONE 8:e55687
Charpentier E, Richter H, van der Oost J et al (2015) Biogenesis pathways of RNA guides in archaeal and bacterial CRISPR-Cas adaptive immunity. FEMS Microbiol Rev 39:428–441
Cheng X, Zheng X, Zhou X et al (2016) Regulation of oxidative response and extracellular polysaccharide synthesis by a diadenylate cyclase in Streptococcus mutans. Environ Microbiol 18:904–922
Citorik RJ, Mimee M, Lu TK (2014) Sequence-specific antimicrobials using efficiently delivered RNA-guided nucleases. Nat Biotechnol 32:1141–1145
Comas I, Homolka S, Niemann S (2009) Genotyping of genetically monomorphic bacteria: DNA sequencing in Mycobacterium tuberculosis highlights the limitations of current methodologies. PLoS ONE 4:e7815
Cong L, Ran FA, Cox D et al (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823
Cui L, Bikard D (2016) Consequences of Cas9 cleavage in the chromosome of Escherichia coli. Nucleic Acids Res 44:4243–4251
Cui L, Wang X, Huang D et al (2020) CRISPR-cas3 of Salmonella upregulates bacterial biofilm formation and virulence to host cells by targeting quorum-sensing systems. Pathogens 9(1):53
Ebina H, Misawa N, Kanemura Y et al (2013) Harnessing the CRISPR/Cas9 system to disrupt latent HIV-1 provirus. Sci Rep 3:2510
Endo A, Watanabe T, Ogata N et al (2015) Comparative genome analysis and identification of competitive and cooperative interactions in a polymicrobial disease. ISME J 9:629–642
Fabre L, Zhang J, Guigon G et al (2012) CRISPR typing and subtyping for improved laboratory surveillance of Salmonella infections. PLoS ONE 7:e36995
Faure G, Makarova KS, Koonin EV (2019) CRISPR-Cas: complex functional networks and multiple roles beyond adaptive immunity. J Mol Biol 431:3–20
Gao NJ, Al-Bassam MM, Poudel S et al (2019) Functional and proteomic analysis of Streptococcus pyogenes virulence upon loss of its native Cas9 nuclease. Front Microbiol 10:1967
Garneau JE, Dupuis ME, Villion M et al (2010) The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468:67–71
Ginevra C, Jacotin N, Diancourt L et al (2012) Legionella pneumophila sequence type 1/Paris pulsotype subtyping by spoligotyping. J Clin Microbiol 50:696–701
Gomaa AA, Klumpe HE, Luo ML (2014) Programmable removal of bacterial strains by use of genome-targeting CRISPR-Cas systems. mBio 5:e00928–13
Gomgnimbou MK, Abadia E, Zhang J et al (2012) “Spoligoriftyping”, a dual-priming-oligonucleotide-based direct-hybridization assay for tuberculosis control with a multianalyte microbead-based hybridization system. J Clin Microbiol 50:3172–3179
Grissa I, Bouchon P, Pourcel C et al (2008) On-line resources for bacterial micro-evolution studies using MLVA or CRISPR typing. Biochimie 90:660–668
Guan J, Wang W, Sun B (2017) Chromosomal targeting by the type III-A CRISPR-Cas system can reshape genomes in Staphylococcus aureus. mSphere 2(6):e00403- e00417
Gunderson FF, Cianciotto NP (2013) The CRISPR-associated gene cas2 of Legionella pneumophila is required for intracellular infection of amoebae. mBio 4:e00074–13
Hai T, Teng F, Guo R (2014) One-step generation of knockout pigs by zygote injection of CRISPR/Cas system. Cell Res 24:372–375
Hale CR, Majumdar S, Elmore J et al (2012) Essential features and rational design of CRISPR RNAs that function with the Cas RAMP module complex to cleave RNAs. Mol Cell 45:292–302
Heidrich N, Hagmann A, Bauriedl S et al (2019) The CRISPR/Cas system in Neisseria meningitidis affects bacterial adhesion to human nasopharyngeal epithelial cells. RNA Biol 16:390–396
Heussler GE, Miller JL, Price CE et al (2016) Requirements for Pseudomonas aeruginosa type I-F CRISPR-Cas adaptation determined using a biofilm enrichment assay. J Bacteriol 198:3080–3090
Hu W, Kaminski R, Yang F et al (2014) RNA-directed gene editing specifically eradicates latent and prevents new HIV-1 infection. Proc Natl Acad Sci U S A 111:11461–11466
Hudaiberdiev S, Shmakov S, Wolf YI et al (2017) Phylogenomics of Cas4 family nucleases. BMC Evol Biol 17(1):232
Hullahalli K, Rodrigues M, Nguyen UT et al (2018) A semi lethal CRISPR-Cas system permits DNA acquisition in Enterococcus faecalis. bioRxiv aCC-BY-NC-ND 4.0.2.
Ivancic-Bace I, Cass SD, Wearne SJ et al (2015) Different genome stability proteins underpin primed and naive adaptation in E. coli CRISPR-Cas immunity. Nucleic Acids Res 43:10821–10830
Jansen R, Embden JD, Gaastra W et al (2002) Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol 43:1565–1575
Jiang Y, Chen B, Duan C et al (2015) Multigene editing in the Escherichia coli genome via the CRISPR-Cas9 system. Appl Environ Microbiol 81(7):2506–2514
Jones CL, Sampson TR, Nakaya HI et al (2012) Repression of bacterial lipoprotein production by Francisella novicida facilitates evasion of innate immune recognition. Cell Microbiol 14:1531–1543
Jorth P, Whiteley M (2012) An evolutionary link between natural transformation and CRISPR adaptive immunity. mBio 3(5):e00309- e00312
Kamerbeek J, Schouls L, Kolk A et al (1997) Simultaneous detection and strain differentiation of Mycobacterium tuberculosis for diagnosis and epidemiology. J Clin Microbiol 35:907–914
Killelea T, Bolt EL (2017) CRISPR-Cas adaptive immunity and the three Rs. Biosci Rep 37(4):BSR20160297
Levy A, Goren MG, Yosef I et al (2015) CRISPR adaptation biases explain preference for acquisition of foreign DNA. Nature 520:505–510
Liu F, Kariyawasam S, Jayarao BM et al (2011) Subtyping Salmonella enterica serovar enteritidis isolates from different sources by using sequence typing based on virulence genes and clustered regularly interspaced short palindromic repeats (CRISPRs). Appl Environ Microbiol 77:4520–4526
Liu T, Li Y, Wang X et al (2015) Transcriptional regulator-mediated activation of adaptation genes triggers CRISPR de novo spacer acquisition. Nucleic Acids Res 43:1044–1055
Louwen R, Horst-Kreft D, de Boer AG et al (2013) A novel link between Campylobacter jejuni bacteriophage defence, virulence and Guillain-Barre syndrome. Eur J Clin Microbiol Infect Dis 32:207–226
MacRitchie DM, Buelow DR, Price NL et al (2008) Two-component signaling and gram negative envelope stress response systems. Adv Exp Med Biol 631:80–110
Makarova KS, Aravind L, Grishin NV et al (2002) A DNA repair system specific for thermophilic Archaea and bacteria predicted by genomic context analysis. Nucleic Acids Res 30:482–496
Makarova KS, Haft DH, Barrangou R et al (2011) Evolution and classification of the CRISPR-Cas systems. Nat Rev Microbiol 9:467–477
Makarova KS, Wolf YI, Alkhnbashi OS et al (2015) An updated evolutionary classification of CRISPR-Cas systems. Nat Rev Microbiol 13:722–736
Mandin P, Repoila F, Vergassola M et al (2007) Identification of new noncoding RNAs in Listeria monocytogenes and prediction of mRNA targets. Nucleic Acids Res 35:962–974
Mangas EL, Rubio A, Alvarez-Marin R et al (2019) Pangenome of Acinetobacter baumannii uncovers two groups of genomes, one of them with genes involved in CRISPR/Cas defence systems associated with the absence of plasmids and exclusive genes for biofilm formation. Microb Genom 5(11):e000309
Marraffini LA, Sontheimer EJ (2008) CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322:1843–1845
Marraffini LA, Sontheimer EJ (2010) CRISPR interference: RNA-directed adaptive immunity in bacteria and archaea. Nat Rev Genet 11:181–190
Medina-Aparicio L, Rodriguez-Gutierrez S, Rebollar-Flores JE et al (2021) The CRISPR-Cas system is involved in Ompr genetic regulation for outer membrane protein synthesis in Salmonella Typhi. Front Microbiol 12:657404
Mojica FJ, Diez-Villasenor C, Soria E et al (2000) Biological significance of a family of regularly spaced repeats in the genomes of Archaea, bacteria and mitochondria. Mol Microbiol 36:244–256
Mokrousov I, Vyazovaya A, Kolodkina V et al (2009) Novel macroarray-based method of Corynebacterium diphtheriae genotyping: evaluation in a field study in Belarus. Eur J Clin Microbiol Infect Dis 28:701–703
Newsom S, Parameshwaran HP, Martin L et al (2021) The CRISPR-Cas mechanism for adaptive immunity and alternate bacterial functions fuels diverse biotechnologies. Front Cell Infect Microbiol 10:619763
Nozawa T, Furukawa N, Aikawa C et al (2011) CRISPR inhibition of prophage acquisition in Streptococcus pyogenes. PLoS ONE 6:e19543
Palmer KL, Gilmore MS (2010) Multidrug-Resistant Enterococci Lack CRISPR-Cas Mbio 1(4):e00227-e310
Palmer KL, Whiteley M (2011) DMS3-42: the secret to CRISPR-dependent biofilm inhibition in Pseudomonas aeruginosa. J Bacteriol 193:3431–3442
Perez-Rodriguez R, Haitjema C, Huang Q et al (2011) Envelope stress is a trigger of CRISPR RNA-mediated DNA silencing in Escherichia coli. Mol Microbiol 79:584–599
Pourcel C, Salvignol G, Vergnaud G (2005) CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additional tools for evolutionary studies. Microbiology (reading) 151:653–663
Quax TE, Voet M, Sismeiro O et al (2013) Massive activation of archaeal defense genes during viral infection. J Virol 87:8419–8428
Radovcic M, Killelea T, Savitskaya E et al (2018) CRISPR-Cas adaptation in Escherichia coli requires RecBCD helicase but not nuclease activity, is independent of homologous recombination, and is antagonized by 5’ ssDNA exonucleases. Nucleic Acids Res 46:10173–10183
Ratner HK, Sampson TR, Weiss DS (2015) I can see CRISPR now, even when phage are gone: a view on alternative CRISPR-Cas functions from the prokaryotic envelope. Curr Opin Infect Dis 28(3):267–274
Rezzonico F, Smits TH, Duffy B (2011) Diversity, evolution, and functionality of clustered regularly interspaced short palindromic repeat (CRISPR) regions in the fire blight pathogen Erwinia amylovora. Appl Environ Microbiol 77(11):3819–3829
Sampson TR, Saroj SD, Llewellyn AC et al (2013) A CRISPR/Cas system mediates bacterial innate immune evasion and virulence. Nature 497:254–257
Sampson TR, Weiss DS (2013) Alternative roles for CRISPR/Cas systems in bacterial pathogenesis. PLoS Pathog 9:e1003621
Sampson TR, Weiss DS (2014) CRISPR-Cas systems: new players in gene regulation and bacterial physiology. Front Cell Infect Microbiol 4:37
Sander JD, Joung JK (2014) CRISPR-Cas systems for editing, regulating and targeting genomes. Nat Biotechnol 32:347–355
Sapranauskas R, Gasiunas G, Fremaux C et al (2011) The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coli. Nucleic Acids Res 39:9275–9282
Schouls LM, Reulen S, Duim B et al (2003) Comparative genotyping of Campylobacter jejuni by amplified fragment length polymorphism, multilocus sequence typing, and short repeat sequencing: strain diversity, host range, and recombination. J Clin Microbiol 41:15–26
Selle K, Klaenhammer TR, Barrangou R (2015) CRISPR-based screening of genomic island excision events in bacteria. Proc Natl Acad Sci USA 112:8076–8081
Serbanescu MA, Cordova M, Krastel K et al (2015) Role of the Streptococcus mutans CRISPR-Cas systems in immunity and cell physiology. J Bacteriol 197:749–761
Sesto N, Touchon M, Andrade JM et al (2014) A PNPase dependent CRISPR System in Listeria. PLoS Genet 10:e1004065
Shabbir MAB, Tang Y, Xu Z et al (2018) The involvement of the Cas9 gene in virulence of Campylobacter jejuni. Front Cell Infect Microbiol 8:285
Shmakov S, Smargon A, Scott D et al (2017) Diversity and evolution of class 2 CRISPR-Cas systems. Nat Rev Microbiol 15:169–182
Sinkunas T, Gasiunas G, Waghmare SP et al (2013) In vitro reconstitution of cascade-mediated CRISPR immunity in Streptococcus thermophilus. EMBO J 32:385–394
Smith GR (2012) How RecBCD enzyme and Chi promote DNA break repair and recombination: a molecular biologist’s view. Microbiol Mol Biol Rev 76:217–228
Solbiati J, Duran-Pinedo A, Godoy Rocha F et al (2020) Virulence of the pathogen Porphyromonas gingivalis is controlled by the CRISPR-Cas protein Cas3. mSystems 5(5):e00852–20
Sorek R, Kunin V, Hugenholtz P (2008) CRISPR–a widespread system that provides acquired resistance against phages in bacteria and archaea. Nat Rev Microbiol 6:181–206
Stern A, Keren L, Wurtzel O et al (2010) Self-targeting by CRISPR: gene regulation or autoimmunity? Trends Genet 26:335–340
Tang B, Gong T, Zhou X et al (2019) Deletion of cas3 gene in Streptococcus mutans affects biofilm formation and increases fluoride sensitivity. Arch Oral Biol 99:190–197
Thony-Meyer L, Kaiser D (1993) devRS, an autoregulated and essential genetic locus for fruiting body development in Myxococcus xanthus. J Bacteriol 175:7450–7462
van Belkum A, van den Braak N, Godschalk P (2001) A Campylobacter jejuni gene associated with immune-mediated neuropathy. Nat Med 7:752–753
Vercoe RB, Chang JT, Dy RL et al (2013) Cytotoxic chromosomal targeting by CRISPR/Cas systems can reshape bacterial genomes and expel or remodel pathogenicity islands. PLoS Genet 9:e1003454
Vergnaud G, Li Y, Gorge O et al (2007) Analysis of the three Yersinia pestis CRISPR loci provides new tools for phylogenetic studies and possibly for the investigation of ancient DNA. Adv Exp Med Biol 603:327–338
Viswanathan P, Murphy K, Julien B et al (2007) Regulation of dev, an operon that includes genes essential for Myxococcus xanthus development and CRISPR-associated genes and repeats. J Bacteriol 189:3738–3750
Wang H, Yang H, Shivalila CS et al (2013) One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153:910–918
Watanabe T, Nozawa T, Aikawa C et al (2013) CRISPR regulation of intraspecies diversification by limiting IS transposition and intercellular recombination. Genome Biol Evol 5:1099–1114
Westra ER, Buckling A, Fineran PC (2014) CRISPR-Cas systems: beyond adaptive immunity. Nat Rev Microbiol 12:317–326
Westra ER, van Erp PB, Kunne T et al (2012) CRISPR immunity relies on the consecutive binding and degradation of negatively supercoiled invader DNA by cascade and Cas3. Mol Cell 46:595–605
Wu WH, Tsai YT, Justus S et al (2016) CRISPR repair reveals causative mutation in a preclinical model of retinitis pigmentosa. Mol Ther 24:1388–1394
Xiao Y, Luo M, Dolan AE (2018) Structure basis for RNA-guided DNA degradation by cascade and Cas3. Science 361(6397):eaat0839
Yan S, Tu Z, Liu Z et al (2018) A huntingtin knockin pig model recapitulates features of selective neurodegeneration in Huntington’s disease. Cell 173:989–1002e13.
Yosef I, Manor M, Kiro R et al (2015) Temperate and lytic bacteriophages programmed to sensitize and kill antibiotic-resistant bacteria. Proc Natl Acad Sci USA 112:7267–7272
Young JC, Dill BD, Pan C et al (2012) Phage-induced expression of CRISPR-associated proteins is revealed by shotgun proteomics in Streptococcus thermophilus. PLoS ONE 7:e38077
Zegans ME, Wagner JC, Cady KC et al (2009) Interaction between bacteriophage DMS3 and host CRISPR region inhibits group behaviors of Pseudomonas aeruginosa. J Bacteriol 191:210–219
Zhan Y, Sun X, Li B et al (2018) Establishment of a PRKAG2 cardiac syndrome disease model and mechanism study using human induced pluripotent stem cells. J Mol Cell Cardiol 117:49–61
Zhang J, Kasciukovic T, White MF (2012) The CRISPR associated protein Cas4 Is a 5’ to 3’ DNA exonuclease with an iron-sulfur cluster. PLoS ONE 7:e47232
Zheng Z, Zhang Y, Liu Z et al (2020) The CRISPR-Cas systems were selectively inactivated during evolution of Bacillus cereus group for adaptation to diverse environments. ISME J 14:1479–1493
Zhu J, Ananthaswamy N, Jain S et al (2021) A universal bacteriophage T4 nanoparticle platform to design multiplex SARS-CoV-2 vaccine candidates by CRISPR engineering. Sci Adv 7:eabh1547.
Funding
The authors are highly grateful to the Council of Scientific and Industrial Research (CSIR), New Delhi, India, for financial assistance to Ms. Veena Devi in the form of JRF/SRF (vide no. 09/135 (0743) EMR-2016).
Author information
Authors and Affiliations
Contributions
VD, conceptualization and writing—original draft; SC, conceptualization and reviewing; KH, reviewing.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Devi, V., Harjai, K. & Chhibber, S. CRISPR-Cas systems: role in cellular processes beyond adaptive immunity. Folia Microbiol 67, 837–850 (2022). https://doi.org/10.1007/s12223-022-00993-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-022-00993-2