Abstract
Several cultural or religious groups claim descent from a common ancestor. The extent to which this claimed ancestry is real or socially constructed can be assessed by means of genetic studies. Syed is a common honorific title given to male Muslims belonging to certain families claiming descent from the Prophet Muhammad through his grandsons Hassan and Hussein, who lived 1,400 years ago and were the sons of the Prophet’s daughter Fatima. If all Syeds really are in direct descent from Hassan and Hussein, we would expect the Y chromosomes of Syeds to be less diverse than those of non-Syeds. Outside the Arab world, we would also expect to find that Syeds share Y chromosomes with Arab populations to a greater extent than they do with their non-Syed geographic neighbours. In this study, we found that the Y chromosomes of self-identified Syeds from India and Pakistan are no less diverse than those non-Syeds from the same regions, suggesting that there is no biological basis to the belief that self-identified Syeds in this part of the world share a recent common ancestry. In addition to Syeds, we also considered members of other hereditary Muslim lineages, which either claim descent from the tribe or family of Muhammad or from the residents of Medinah. Here, we found that these lineages showed greater affinity to geographically distant Arab populations, than to their neighbours from the Indian subcontinent, who do not belong to an Islamic honorific lineage.
Similar content being viewed by others
Introduction
Several studies have been carried out in order to better understand the link between social and biological kinship in human societies. In particular, it is relatively widespread for traditional societies and religious groups to claim descent from a common ancestor, and genetic analyses have been used to test some of these claims. For instance, Y chromosome data have allowed the assessment of the genetic relatedness of individuals belonging to paternally inherited Jewish castes (Behar et al. 2003; Skorecki et al. 1997; Thomas et al. 1998). It has also been used to investigate oral histories concerning the foundation of an ancient ruling dynasty in Cameroon (Veeramah et al. 2008) and to test beliefs of common ancestry which are widespread in traditional tribes and clans from Central Asia (Chaix et al. 2004).
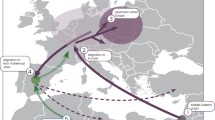
In this study, we were interested in investigating the claim of a recent common origin for a particular religious group: the Syeds. According to Islamic tradition, the Prophet Muhammad’s grandsons, Hassan and Hussein, were given the title ‘Syed’, meaning chief. They were the sons of Muhammad’s youngest daughter, Fatima (Walker 1998). Muhammad himself had no son who survived into adulthood. In the early days of Islamic expansion, male line descendents of Hassan and Hussein gained eminence and collectively became known as Syeds (Walker 1998). Today's Syeds, who are found throughout the Muslim world, still claim patrilineal descent from Hassan or Hussein (Levy 1957; Fig. 1). Despite Syeds having no formal religious authority above non-Syed Muslims, they usually have elevated social status, particularly, although not exclusively, within Shiite communities (Weekes 1984). In some more traditional areas of the Muslim world, Syeds are practically a local aristocracy who command great respect among the common people.
The different levels of relatedness in the lineage studied. Syeds (represented by black squares and circles) claim direct paternal descend of Muhammad (M) through the sons of his daughter Fatima (F), Hassan (Ha) and Hussein (Hu). Hashemites also claim to be related to Muhammad but through his great-grandfather Hashim (Hi). Quraysh are not related to Muhammad but were part of the same tribe (represented by grey squares and circles). Ansari are from the city of Medinah and are meant to have helped Muhammad during times of need. Dotted lines indicate that not all generations are represented
Alongside Syeds there are other patrilineally defined lineages in the Islamic world that are often attributed high social status through claimed descent either from the family or tribe of Muhammad (Hashemites and Quraysh) or from the so-called Ansari (Arabic ‘helpers’) who were the inhabitants of the city of Medinah, a major centre of the early Islamic faith, during the Prophet Muhammad’s life time. The Hashemites originally trace back their ancestry to Hashim ibn Abd al-Manaf, the great-grandfather of the prophet Muhammad (Blackwell 2009), and the Quraysh claim membership of the original tribe of Muhammad (Weekes 1984). Here, we collectively refer to these lineages as ‘Islamic honorific lineages’ (IHL). Arabs had been trading all along the southern coasts of India even before Muslim Arabs began the conquest of Sind in 711 ad. As Islam expanded, honorific families spread across the Muslim world (Israeli 1982).
Based on a survey of gene frequencies at six loci, Aarzoo and Afzal (2005) showed a relative proximity of Syeds with Arab groups. However, studying genetic variation on the Y chromosome could prove more informative in this specific case. It has indeed proved highly valuable in testing claims of common patrilineal ancestry (Behar et al. 2003; Foster et al. 1998; Skorecki et al. 1997; Thomas et al. 1998; Zerjal et al. 2003). In particular, rapidly evolving microsatellites allow Y chromosomes to be classified into detailed haplotypes while slowly evolving markers such as single nucleotide polymorphisms, insertions and deletions, referred to here as unique event polymorphisms (UEPs), allow Y chromosomes to be classified into broader haplogroups. Furthermore, under appropriate circumstances and using the knowledge of mutation rates for Y chromosome microsatellites (Bianchi et al. 1998; Heyer et al. 1997; Kayser et al. 2000), estimates of the time to a common ancestor can be made (Behar et al. 2003; Thomas et al. 1998; Zerjal et al. 2003). If patrilineal descent has indeed been followed since the time of Muhammad, we would expect the Y chromosome haplotypes of living Syeds to be considerably less diverse than those found among their non-Syed neighbours and to be derived from an ancestral haplotype at some point in the last 1,500 years. Furthermore, we would expect to see a higher proportion of Y chromosome haplotypes that are typical of Arab populations among IHL individuals living outside the Arab world, than we would in their non-IHL neighbours.
Here, we examine the Y chromosomes of Syed, Hashemite, Quraysh and Ansari men originating from the Indian subcontinent within at least the last two generations and who now live in the British Isles. We use these data to test the following two hypotheses: (a) that the Y chromosomes of Syed men are significantly less diverse than those of their non-IHL lineage neighbours and (b) that men belonging to the Syed, Hashemite, Quraysh and Ansari lineages show a significantly greater patrilineal genetic affinity to Arab populations than to their non-IHL neighbours.
Material and methods
Study populations and sampling
We obtained buccal swab samples from 56 Syed, 16 Quraysh, 1 Hashmite and 5 Ansari men of Pakistani or Indian origin, currently living in London and Manchester. Donors were initially identified by surnames associated with their respective lineages and later confirmed as belonging to those lineages through self-identification. Buccal swab samples were collected by post; a storage buffer (0.05% SDS, 0.05 M EDTA pH 8.0) was added to stabilise DNA during transport and storage. Anonymous questionnaires were used to gather information about lineage status, place of birth, first and second languages, father’s place of birth, father’s first and second languages, grandfather’s place of birth and recent family migration paths. We studied the first and second generation of individuals whose parents had migrated from the Indian subcontinent to the UK within the last 50 years. For comparison we collected 37 individuals from the same population background who did not claim Syed, Hashemite, Quraysh or Ansari status (non-IHL lineage). DNA was extracted using standard organic phase methods.
Molecular analysis
Y chromosomes were typed in all samples for six microsatellites (DYS19, DYS388, DYS390, DYS391, DYS392, DYS393) and 11 UEP markers (92R7, M9, M13, M17, M20, SRY + 465, SRY4064, SRY10831, sY81, Tat, YAP) as previously described (Thomas et al. 1999). In addition, the UEP marker 12f2 was typed as described by Rosser et al. (2000). Microsatellite repeat numbers were assigned according to the nomenclature of Kayser et al. (1997). Y chromosome haplogroups were defined by the 12 UEP markers according to the nomenclature of the Y Chromosome Consortium (2002; see Supplementary Figure 1 and Supplementary Table 1).
Statistical and population genetic analysis
Unbiased genetic diversity, h, and its standard error were calculated using the formula given by Nei (1987). Tests for the significance of differences in h values were carried out by means of bootstrap resampling and a standard two-tailed z test. As a conservative measure, only the larger of the two P values was used (Thomas et al. 2002).
The Y chromosomes of IHL and non-IHL men from the Indian subcontinent were compared to compatible data from 12 Arab populations using the genetic distance measure F ST (Weir 1996) and Nei’s measure of genetic identity I (Nei 1987), based either on haplogroup or full haplotype frequencies, and using R ST (Michalakis and Excoffier 1996), based upon microsatellite data. The 12 Arab population samples were Algerians (n = 164; unpublished), Arabs from the Highland region between the northern West Bank of the Jordan river and southern Israel/southwest Jordan (n = 24; Nebel et al. 2000), Israeli and Palestinian Arabs (n = 143; Nebel et al. 2000), Arabs from the Israeli village of Kfar Kara (n = 135; unpublished), Israeli Bedouins (n = 25; see Supplementary Table 2), Jordanian Bedouins (n = 24; see Supplementary Table 2), Kordofanian Arabs from Sudan (n = 69; unpublished), Kuwaitis (n = 72; unpublished), North Sudanese (n = 208; unpublished), Syrians (n = 72; Thomas et al. 2002), Yemenis (n = 93; Thomas et al. 2000). To test if IHL men had a greater proportion of Arab ancestry than their non-IHL neighbours from India and Pakistan, we employed two methods. In the first, we compared the IHL sample and the non-IHL sample to each of the 12 Arab populations separately. The number of times that IHL individuals were genetically closer to the Arab population was compared to a binomial expectation with 12 tries and a probability of 0.5. In the second, Y chromosome data from 12 Arab populations was combined and compared to the IHL and the non-IHL samples. The significance of difference in F ST, Nei’s I and R ST was assessed by bootstrap resampling the data 1,000 times for all pairwise population comparisons using the PopA software (http://www.ucl.ac.uk/tcga/software/).
We used an admixture model to quantify the respective contributions (or admixture coefficients) of two hypothetical parental populations—Arab populations pooled together (p1) and non-IHL populations from India and Pakistan (p2)—to the presumably hybrid gene pool of the IHL sample (pH). The method used is based on the estimation of average coalescence times between random pairs of genes sampled both within and between populations and is implemented by the program Admix (Bertorelle and Excoffier 1998). This method allows one either to consider molecular distances between alleles or to assume equal distances between alleles. Here, we chose the latter approach because (a) by considering allele frequencies, the estimated admixture coefficients are less affected by the stochasticity of the mutation process and (b) the time-scale of the admixture process we want to investigate is more comparable to the short times through which genetic drift acts than to the long times through which mutations accumulate.
Finally, we carried out a Principal Coordinate Analysis (classical multidimensional scaling (MDS)) on R ST genetic distances (Fig. 2) to visually represent the genetic affinities of the various populations in two dimensions.
Results
The 12 UEP markers defined seven observed haplogroups (Table 1), and the addition of six microsatellites defined a total of 75 observed haplotypes (see Supplementary Table 1) among the 115 southern samples from the Indian subcontinent. No instances of homoplasy of microsatellite haplotypes across UEP haplogroups were observed. No high-frequency modal haplotype was observed (the two highest frequency haplotypes both equal 0.0893 in Syeds), and most haplotypes occurred only once in our samples. The three most frequent haplogroups were BR*(xDE,J,K), R1a1 and J.
Gene diversity was 0.8045 ± 0.0227 in Syeds and 0.7372 ± 0.0393 in the non-IHL sample. The difference in genetic diversity between Syed and non-IHL individuals is not statistically significant, either by performing a z test (P = 0.1388) or a bootstrapping method (P = 0.1312). It is worth noting that, contrary to expectation, gene diversity is actually higher in Syeds than in the non-IHL sample, although the difference is not statistically significant.
We then tested whether the IHL sample (or Syeds only) showed greater affinity to Arab populations than their non-IHL neighbours from the Indian subcontinent. The haplogroup frequencies of the Arab populations are presented in Table 2. We calculated pairwise genetic distances (F ST, R ST) and Nei’s measure of genetic identity between the IHL sample (and Syeds only) and each of the 12 Arab populations. We found that the IHL sample was significantly closer to the Arab populations than were their non-IHL neighbours (Table 3) in all comparisons, except when considering Syeds only for F ST based on UEP + STR-based haplotype frequencies. This is probably due to the smaller sample size considered (56 individuals instead of 115 when considering all IHL individuals) and the large number of singletons observed when UEP and STR data are considered.
We also compared various genetic distances (Nei’s I, F ST and R ST) between the 12 Arab populations pooled together and either the IHL (or Syeds only) or the non-IHL sample from India and Pakistan (Table 4). By means of a bootstrap procedure, we found that when considering both UEP + STR-based haplotypes, Nei’s index of genetic identity between Arab populations and the Syeds was significantly greater than the one between the Arab populations and the non-IHL sample. Considering R ST values, the genetic distance between the Arab populations and the non-IHL sample was significantly greater than the one between the Arab populations and the Syeds. No significant difference in the two comparisons was found when considering F ST values based on haplotype frequencies. This is likely to be due to the reduced power of F ST in this case because of the large number of singleton haplotypes.
To obtain an estimate of the proportion of Arab ancestry among the IHL sample, we performed an admixture analysis based on STR allele frequencies. We found a contribution of 84.6% of the Arabic populations and 15.4% of the non-IHL populations to the IHL sample. Bootstrapping the data 1,000,000 times resulted in an admixture coefficient estimate for the Arabic populations of 94.1 ± 7.1%.
Finally, the MDS analysis based on R ST values (Fig. 2) illustrates the genetic diversity of the Arab populations and the clear association of the Syeds with other Arab populations, as well as their lack of genetic affinity with the neighbouring populations of India and Pakistan.
Discussion
This study shows that the Y chromosomes of a sample of self-identified Syed men exhibit the same level of genetic diversity as their non-IHL neighbours from the Indian subcontinent. However, self-identified men belonging to the IHL (Syeds, Hashemites, Quraysh and Ansari) show a greater genetic affinity to Arab populations—despite the geographic distance—than do their neighbouring populations from India and Pakistan.
In some rare cases, high frequency modal haplotypes may be representative of particular communities. For instance, the Cohanim are Jewish priests who also have a patrilineal mode of inheritance and present a single haplotype (the Cohen Modal Haplotype) at a very high frequency (Thomas et al. 1998). One possible contributing factor to the presence of modal haplotypes in some Jewish communities is that they remained in relative isolation for the last 500 years (Thomas et al. 1998). Unlike in Cohanim, the Y chromosomes of Levites present a pattern more comparable to the one found in Syeds (Behar et al. 2003). The Levites constitute another paternally inherited Jewish caste; they claim to be the descendants of Levi, one of the sons of Jacob, and also perform particular religious functions. Their Y chromosomes also show evidence of multiple origins. However, once divided geographically into two groups, specific high-frequency haplotypes can be identified (Behar et al. 2003).
In contrast to these populations, no specific haplotypes were identified among the Indian and Pakistani Syed samples examined here. So what are the factors that could account for the lack of genetic similarity among the IHL sample and more specifically Syeds?
-
1.
When compared to the relative geographic isolation of the Jewish communities, the distribution of the IHL, and Syeds in particular, is likely to have resulted from a rapid expansion of Islam throughout the world (Levy 1957). Indeed, the spread of Arabic language and culture and Arab identity in the Middle East and North Africa began shortly after the advent of Islam in the seventh century and was followed by Arab Muslim expansion on other continents. From the end of the caliphs’ power in the tenth century to the beginning of the sixteenth century, the geographic extent of the Muslim world almost doubled (Wuthnow 1998). It is therefore likely that during these expansions, gene flow from neighbouring or newly Islamised communities occurred regularly.
-
2.
Biased sampling and errors due to small sample size may have occurred, and this may have influence our results. However, we believe it is unlikely that our sampling strategy significantly skewed our results. Despite our sample sizes being relatively small (78 IHL individuals, including 56 self-identified Syeds), our data were sufficient to test the two hypotheses under investigation. Indeed, the possibility of a recent common ancestor was rejected, whereas the presence of an elevated Arab ancestry was accepted with strong statistical support. If we had insufficient data, we would not have been able to find statistical support for this hypothesis. In addition, the number of Cohanim Jews previously studied (Skorecki et al. 1997) was not much higher (68 individuals in total). We therefore believe that increasing the number of Syed Y chromosomes analysed would probably identify even more haplotypes and only serve to support our results.
Furthermore, our samples were collected from individuals originally from India and Pakistan, therefore possibly producing a bias in regional representation. It would certainly be worth studying the Y chromosomes of IHL individuals from other parts of the world to assess whether this lack of genetic uniformity is an exception or the rule among Syeds. In South Asia, there have been many different points of entry and routes of expansion for Muslims over several centuries (Israeli 1982). It is a possibility that more gene flow occurred in this region than in other parts of the world. Nonetheless, for the time being, this first genetic study shows that Syeds from the Indian subcontinent do not present any sign of recent common ancestry.
Finally, there is the question of the extent to which a Diaspora sample, in this case London and Manchester Pakistanis and Indians, adequately reflects the Y chromosomes of their source populations. The importance of an appropriate Y chromosome sampling scheme has already been stressed in other studies. For instance, it has been shown that in Armenians, the whole population is not always appropriately represented by its displaced subset (Weale et al. 2001). We are not aware of any population genetics study of immigrants from the Indian subcontinent in the UK. However, the individuals sampled were unrelated and initially chosen based on their surnames and later self-identified as Syeds; at present we see no reason why they would represent a distinct subset.
-
3.
Last but not least, the transmission of honorific titles and in particular the Syed status may not have been as strictly patrilineal as traditionally thought. Instead, it is possible that the Syed status could have been passed on following other routes. For instance, it is known that this status has occasionally been transmitted to individuals whose mother was the daughter of a Syed but whose father was a non-Syed and instances in which high caste converts to Islam took on the Syed honorific title are also known (Kilic 2007). Even if this kind of transmission occurred irregularly, it could be enough to explain why the Y chromosomes of Syeds fail to show a recent common ancestry. However, it is believed that marriages between a Syed women and a man of a lower social rank was very rare in the past (Kilic 2007).
It is interesting to note that even before the advent of genetics, Islamic genealogists already rejected many requests of Syed status based on insufficient evidence or fake documents. This suggests that self-identification as a Syed is probably not the best way of sampling this specific group. Moreover, here we only studied Sunni Muslims from India and Pakistan. It would be interesting to study Shiite Muslims (mostly present in Iran and Iraq) as they might have different authentication criteria of the Syed status, particularly given that this title is claimed through descent from at least one of the Shiite Imams. We could also investigate whether the Y chromosomes of Syeds which have been officially identified as such by a genealogical committee show signs of common ancestry. The advantages conferred by the title of Syed are sufficiently great that even the strictest precautions are still unable to stop the presentation of false claims (Kilic 2007). The common expression reported by Sir Denzil Ibbetson (Ibbetson 1916) illustrates well this fact: ‘Last year I was a weaver, this year I am a Shekh, and next year if prices rise I shall be a Saiyad [Syed]’. It is also known that, at least in the case of the Ottoman society at the end of the sixteenth century, the number of claims was so high that strict measures had to be taken in order to restrict the approval of Syed status and to expel those who had joined illegally (Kilic 2007). This might have been the case in other regions as well.
The other important finding of this study is that Syeds, and individuals belonging to IHL in general, appear to be more closely related to Arab populations than to the neighbouring populations. Regardless of the method used, either comparing the IHL sample to each of the Arab population individually or performing bootstrap resampling tests pooling those populations together, we found that the IHL sample was significantly closer to other Arab populations.
In addition, the admixture analysis revealed a contribution of as much as 85% from geographically distant Arab populations to the IHL sample. The overall pattern is certainly more complex than modelled in this study. For instance, it is not clear how genes from a modern population can represent well the genes of a population of the past. Extensive gene flow from other sources into the hybrid populations could have had an influence on our estimates, as well as genetic drift since admixture (see for instance Belle et al. 2006 and Guimaraes et al. 2009). However, our result indicates that a very large proportion of genes from the IHL sample can be traced back to Arab ancestors.
These results are remarkable when one considers that in most instances human populations are primarily related on the basis of geography rather than cultural traits, such as languages (see for instance Ramachandran et al. 2005; Belle and Barbujani 2007). Regarding the Indian subcontinent in particular, it has been suggested that Y chromosomal heritage in India was more influenced by geographical proximity than by religious practices (Gutala et al. 2006). Here, we show that for the IHL, this does not appear to be the case. Our results rather confirm the study of Aarzoo and Afzal (2005) who have shown, based on autosomal allele frequency data, that Syeds and other IHL from Northern India are closer to Arab populations than to Hindus.
To conclude, this study opens the door to further genetic investigations of the Syed lineage. For instance, it would be interesting to investigate additional populations of both self-identified and officially recognised Syeds from different parts of the world, including those from other Sunni and Shiite communities. At present, our study shows that Syeds from the Indian subcontinent have a greater affinity to Arab populations than to their geographic neighbours but do not show any evidence of a recent common patrineal ancestry.
References
Aarzoo SS, Afzal M (2005) Gene diversity in some Muslim populations of North India. Hum Biol 77:343–353
Behar DM, Thomas MG, Skorecki K et al (2003) Multiple origins of Ashkenazi Levites: Y chromosome evidence for both Near Eastern and European ancestries. Am J Hum Genet 73:768–779
Belle EM, Barbujani G (2007) Worldwide analysis of multiple microsatellites: language diversity has a detectable influence on DNA diversity. Am J Phys Anthropol 133:1137–1146
Belle EM, Landry PA, Barbujani G (2006) Origins and evolution of the Europeans' genome: evidence from multiple microsatellite loci. Proc Biol Sci 273:1595–1602
Bertorelle G, Excoffier L (1998) Inferring admixture proportions from molecular data. Mol Biol Evol 15:1298–1311
Bianchi NO, Catanesi CI, Bailliet G et al (1998) Characterization of ancestral and derived Y-chromosome haplotypes of New World native populations. Am J Hum Genet 63:1862–1871
Blackwell S (2009) British military intervention and the struggle for Jordan, King Hussein, Nasser and the Middle East crisis 1955–1958. Routledge, USA
Chaix R, Austerlitz F, Khegay T, Jacquesson S, Hammer MF, Heyer E, Quintana-Murci L (2004) The genetic or mythical ancestry of descent groups: lessons from the Y chromosome. Am J Hum Genet 75:1113–1116
Foster EA, Jobling MA, Taylor PG et al (1998) Jefferson fathered slave's last child. Nature 396:27–28
Guimaraes S, Ghirotto S, Benazzo A et al (2009) Genealogical discontinuities among Etruscan, Medieval, and contemporary Tuscans. Mol Biol Evol 26:2157–2166
Gutala R, Carvalho-Silva DR, Jin L et al (2006) A shared Y-chromosomal heritage between Muslims and Hindus in India. Hum Genet 120:543–551
Heyer E, Puymirat J, Dieltjes P et al (1997) Estimating Y chromosome specific microsatellite mutation frequencies using deep rooting pedigrees. Hum Mol Genet 6:799–803
Ibbetson DCJ (1916) Punjab castes. B.R. Publishing Corporation, Delhi, reprint 1974
Israeli R (1982) The crescent in the East: Islam in Asia Minor. Riverdale Company, Md, London Curzon
Kayser M, Caglia A, Corach D et al (1997) Evaluation of Y-chromosomal STRs: a multicenter study. Int J Legal Med 110:125–133
Kayser M, Roewer L, Hedman M et al (2000) Characteristics and frequency of germ line mutations at microsatellite loci from the human Y chromosome, as revealed by direct observation in father/son pairs. Am J Hum Genet 66:1580–1588
Kilic R (2007) Sayyids and Sharifs in the Ottoman State: on the borders of the true and the false. The Muslim World 96:21–35
Levy R (1957) The social structure of Islam. Second edition of the sociology of Islam. Cambridge University Press, Cambridge
Michalakis Y, Excoffier L (1996) A generic estimation of population subdivision using distances between alleles with special reference for microsatellite loci. Genetics 142:1061–1064
Nebel A, Filon D, Weiss DA et al (2000) High-resolution Y chromosome haplotypes of Israeli and Palestinian Arabs reveal geographic substructure and substantial overlap with haplotypes of Jews. Hum Genet 107:630–641
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Ramachandran S, Deshpande O, Roseman CC et al (2005) Support from the relationship of genetic and geographic distance in human populations for a serial founder effect originating in Africa. Proc Natl Acad Sci USA 102:15942–15947
Rosser ZH, Zerjal T, Hurles ME et al (2000) Y-chromosomal diversity in Europe is clinal and influenced primarily by geography, rather than by language. Am J Hum Genet 67:1526–1543
Skorecki K, Selig S, Blazer S et al (1997) Y chromosomes of Jewish priests. Nature 385:32–35
Thomas MG, Skorecki K, Ben Ami H et al (1998) Origins of old testament priests. Nature 394:138–140
Thomas MG, Bradman N, Flinn HM (1999) High throughput analysis of 10 microsatellite and 11 diallelic polymorphisms on the human Y-chromosome. Hum Genet 105:577–581
Thomas MG, Parfitt T, Weiss DA et al (2000) Y chromosomes traveling south: the cohen modal haplotype and the origins of the Lemba: the "Black Jews of Southern Africa". Am J Hum Genet 66:674–686
Thomas MG, Weale ME, Jones AL et al (2002) Founding mothers of Jewish communities: geographically separated Jewish groups were independently founded by very few female ancestors. Am J Hum Genet 70:1411–1420
Veeramah KR, Zeitlyn D, Fanso VG, Mendell NR, Connell BA, Weale ME, Bradman N, Thomas MG (2008) Sex-specific genetic data support one of two alternative versions of the foundation of the ruling dynasty of the Nso′ in Cameroon. Current Anthropology 49:707–714
Walker B (1998) Foundations of Islam: the making of a world faith. Peter Owen Publishers, London & Chester Springs, London
Weale ME, Yepiskoposyan L, Jager RF et al (2001) Armenian Y chromosome haplotypes reveal strong regional structure within a single ethno-national group. Hum Genet 109:659–674
Weekes RV (1984) Muslim peoples: a world ethnographic survey. Greenwood Press, Westport, CT
Weir BS (1996) Genetic data analysis II. Sinauer Associates Inc., Sunderland, MA
Wuthnow R (1998) The encyclopedia of politics and religion. Congressional Quarterly Books, Washington DC
Zerjal T, Xue Y, Bertorelle G et al (2003) The genetic legacy of the Mongols. Am J Hum Genet 72:717–721
Acknowledgments
For the collection of comparison data, we would like to thank Neil Bradman, Steve Jones, Leila Nadjiba Laredj, Erik Jelinek, Mohamed Farouk Elamin and Pat Smith. We would also like to thank Zahid Parvez and the Muslim Council of Britain for their comments and advice on the manuscript, and the AHRC Centre for the Evolution of Cultural Diversity (CECD) for supporting this research.
Author information
Authors and Affiliations
Corresponding author
Additional information
Elise M. S. Belle and Saima Shah contributed equally to the work.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Belle, E.M.S., Shah, S., Parfitt, T. et al. Y chromosomes of self-identified Syeds from the Indian subcontinent show evidence of elevated Arab ancestry but not of a recent common patrilineal origin. Archaeol Anthropol Sci 2, 217–224 (2010). https://doi.org/10.1007/s12520-010-0040-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12520-010-0040-1