Abstract
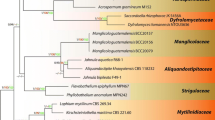
Aquatic hyphomycetes are a polyphyletic group of fungi that play a crucial role in organic matter turnover in streams. They have been traditionally identified based on the morphology of conidia collected in stream water or obtained from leaves colonized in nature upon aeration in the laboratory. Therefore, species identification is limited by our ability to induce conidium production and to establish pure cultures. Conidial shapes are believed to be the result of convergent evolution, so similar conidia may be produced by different conidiogenesis processes, which may prevent unambiguous identification. Currently, a great effort in fungal taxonomy is being made at introducing a set of criteria based on comparisons of selected nucleotide sequences instead of or in addition to phenotypic characters. We examined the suitability of ITS1-5.8S-ITS2 rRNA gene region or its subregions (ITS1 and ITS2) to identify aquatic hyphomycetes, by sequencing and comparing these regions in 94 fungal isolates belonging to 19 species collected in Portuguese streams with different environmental conditions during 8 years. Sequences of ITS1, ITS2 and ITS1-5.8S-ITS2 rRNA genes of the Portuguese isolates of aquatic hyphomycetes and those from the GenBank exhibited taxonomic cohesiveness, the isolates grouped with their respective species but all Tricladium species did not group within the Tricladium genus. Cohesiveness was not observed between isolates with respect to location, condition of stream or date of collection. Evolutionary divergences (ITS1-5.8S-ITS2 sequences; Kimura 2-parameter distance) between con-specific isolates were shallow and a deep divergence between species was generally observed. The NJ trees based on ITS1 or ITS2 rRNA gene sequences had lower statistical support for some internal nodes, we therefore propose ITS1-5.8S-ITS2 rRNA gene as barcode for identifying species of aquatic hyphomycetes.



Similar content being viewed by others
References
Alexopoulos CJ, Mims CW, Blackwell M (1996) Introductory mycology, IVth edn. Wiley, New York
Baral HO, Krieglsteiner GJ (1985) Bausteine zu einer Askomyzeten-Flora der BR Deutschland: In Süddeutschland gefundene inoperculate Discomyzeten mit taxonomischen, ökologischen und chorologischen Hinweisen. Beih Z Mykol 6:1–160
Bärlocher F (2005) Freshwater fungal communities. In: Dighton J, White JF, Oudemans P (eds) The fungal community: its organization and role in the ecosystem. CRC, Boca Raton, pp 39–59
Bärlocher F (2007) Molecular approaches applied to aquatic hyphomycetes. Fungal Biol Rev 21:19–24
Bärlocher F (2009) Reproduction and dispersal in aquatic hyphomycetes. Mycoscience 50:3–8
Bärlocher F, Kendrick B (1974) Dynamics of the fungal population on leaves in a stream. J Ecol 63:761–791
Bärlocher F, Charette N, Letourneau A, Nikolcheva LG, Sridhar KR (2010) Sequencing DNA extracted from single conidia of aquatic hyphomycetes. Fungal Ecol 3:115–121
Baschien C, Marvanová L, Szewzyk U (2006) Phylogeny of selected aquatic hyphomycetes based on morphological and molecular data. Nova Hedwig 83:311–352
Belliveau M, Bärlocher F (2005) Molecular evidence confirms multiple origin of aquatic hyphomycetes. Mycol Res 109:1407–1417
Campbell J, Marvanová L, Gulis V (2009) Evolutionary relationships between aquatic anamorphs and teleomorphs: Tricladium and Varicosporium. Mycol Res 113:1322–1334
Descals E (1997) Ingoldian fungi: some field and laboratory techniques. Boll SocHist Nat Balears 40:169–221
Descals E (2005) Techniques for handling Ingoldian fungi. In: Graça MAS, Bärlocher F, Gessner MO (eds) Methods to study litter decomposition: a practical guide. Springer, Netherlands, pp 129–142
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Gessner MO, Gulis V, Kuehn KA, Chauvet E, Suberkropp K (2007) Fungal decomposers of plant litter in aquatic ecosystems. In: Kubicek CP, Druzhinina IS (eds) The Mycota, vol IV, Environmental and microbial relationships. Springer, Berlin, pp 301–324
Godeas AM (1985) Hifomicetes (Deuteromycotina) acuáticos de Tierra del Fuego I. Physis (Buenos Aires) Secc B 43:7–9
Gulis V, Marvanová L, Descals E (2005) An illustrated key to the common temperate species of aquatic hyphomycetes. In: Graça MAS, Bärlocher F, Gessner MO (eds) Methods for studying litter decomposition. Kluwer Academic, Dordrecht, pp 130–153
Hajibabaei M, Smith MA, Janzen DH, Rodriguez JJ, Whitfield JB, Hebert PDN (2006) A minimalist barcode can identify a specimen whose DNA is degraded. Mol Ecol Notes 6:959–964
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hall BG (2008) Phylogenetic trees made easy: a how-to manual, 3rd edn. Sinauer Associates, Sunderland
Hawksworth DL (2001) The magnitude of fungal diversity: the 1.5 million species estimate revisited. Mycol Res 105:1422–1432
Hebert PDN, Gregory TR (2005) The promise of DNA barcoding for taxonomy. Syst Biol 54:852–859
Hebert PDN, Cywinska A, Ball SL, deWaard JR (2003) Biological identifications through DNA barcodes. Proc R Soc Lond 270:313–321
Kimura M (1980) A simple method of estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Letourneau A, Seena S, Marvanová L, Bärlocher F (2010) Potential use of barcoding to identify aquatic hyphomycetes. Fungal Divers 40:51–64
Lieckfeldt E, Seifert KA (2000) An evaluation of the use of ITS sequences in the taxonomy of the Hypocreales. Stud Mycol 45:35–44
Marvanová L (1997) Freshwater hyphomycetes: a survey with remarks on tropical taxa. In: Janardhanan KK, Rajendran C, Natarajan K, Hawksworth L (eds) Tropical mycology. Science Publishers Inc, Enfield, pp 169–226
Marvanová L (2007) Aquatic hyphomycetes and their meiosporic relatives: slow and laborious solving of a jig-saw puzzle. In: Ganguli BN, Deshmukh SK (eds) Fungi multifaceted microbes. Anamaya Publishers, New Delhi, pp 128–152
Marvanová L, Pascoal C, Cássio F (2003) New and rare hyphomycetes from streams of northwest Portugal. Part I. Cryptog Mycolog 243:39–358
Nilsson RH, Ryberg M, Kristiansson E, Abarenkov K, Larsson KH, Kõljalg U (2006) Taxonomic reliability of DNA sequences in public sequence databases: a fungal perspective. PLoS ONE 1:e59
Nilsson RH, Kristiansson E, Ryberg M, Hallenberg N, Larsson KH (2008) Intraspecific ITS variability in the kingdom fungi as expressed in the international sequence databases and its implications for molecular species identification. Evol Bioinformatics 4:193–201
Pascoal C, Marvanová L, Cássio F (2005) Aquatic hyphomycete diversity in streams of Northwest Portugal. Fungal Divers 19:109–128
Rossman A (2007) Report of the planning workshop for all fungi DNA barcoding. Inoculum 58:1–5
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sati SC, Tiwari N (1992) A new species of Tricladium from Kumaun Himalaya, India. Mycol Res 96:229–232
Seifert KA, Samson RA, deWaard JR, Houbraken J, Lévesque CA, Moncalvo J-M, Gerry Louis-Seize, Hebert PDN (2007) Prospects for fungus identification using CO1 DNA barcodes, with Penicillium as a test case PNAS USA 104:3901–3906
Shearer CA, Descals E, Kohlmeyer B, Marvanová L, Padgett D, Porter D, Raja HA, Schmit JP, Thorton HA, Voglymayer H (2007) Fungal biodiversity in aquatic habitats. Biodivers Conserv 16:49–67
Smith VS (2005) DNA barcoding: perspectives from a “Partnerships for Enhancing Expertise in Taxonomy” (PEET) debate. Syst Biol 54:841–844
Suberkropp K, Klug MJ (1976) Fungi and bacteria associated with leaves during processing in a woodland stream. Ecology 57:707–719
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Ward RD (2009) DNA barcode divergences among species and genera of birds and fishes. Mol Ecol Resour 9:1077–1085
Webster J, Weber RWS (2007) Introduction to fungi, 3rd edn. Cambridge University Press, New York
White TJ, Bruns TD, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic, San Diego, pp 315–322
Acknowledgement
L. Marvanová was supported by the project MSM0021622416.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Seena, S., Pascoal, C., Marvanová, L. et al. DNA barcoding of fungi: a case study using ITS sequences for identifying aquatic hyphomycete species. Fungal Diversity 44, 77–87 (2010). https://doi.org/10.1007/s13225-010-0056-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-010-0056-y